 Back
BackChapter 12
Study Guide - Smart Notes

Gene Expression at the Molecular Level
Introduction to Gene Expression
Gene expression is the process by which information encoded in DNA is converted into functional molecules within the cell, primarily proteins. This process is fundamental to understanding how genes determine traits and cellular functions. - DNA serves as the blueprint for cellular activities. - Gene expression involves two levels: the molecular function of the protein product and the organism’s trait conferred by the gene. - The molecular function of a protein affects cell structure and function, ultimately determining the organism’s phenotype.
Historical Perspective: Inborn Errors of Metabolism
The relationship between genes and enzyme production was first proposed by Archbold Garrod in 1908, who studied patients with metabolic defects known as "inborn errors of metabolism." - Alkaptonuria: A disorder where the body accumulates abnormal levels of homogentisic acid due to a missing enzyme. - Garrod hypothesized that diseases could result from missing enzymes, a concept foundational to molecular genetics. 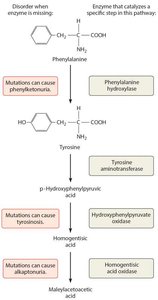
What Do Genes Do? The One-Gene, One-Enzyme Hypothesis
George Beadle and Edward Tatum (1941) expanded Garrod’s work by creating defective genes in bread mold (Neurospora crassa) and observing the resulting phenotypes. - Null or loss-of-function alleles: Mutations that result in nonfunctioning proteins. - Their experiments led to the one-gene, one-enzyme hypothesis: each gene contains the information to make a specific enzyme. - Srb and Horowitz (1944) further tested this hypothesis using a three-step metabolic pathway for arginine synthesis. 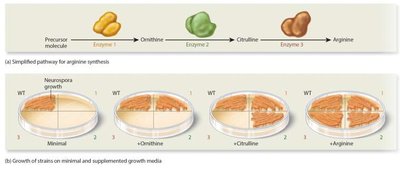
Modern Understanding: One Gene, One Polypeptide
The original hypothesis was modified as scientists discovered that: - Genes encode not only enzymes but also other types of proteins. - Some proteins are composed of multiple polypeptides, each encoded by a separate gene. - Example: Hemoglobin is composed of four polypeptides.
The Central Dogma of Molecular Biology
Flow of Genetic Information
The central dogma summarizes the flow of information in cells: DNA → RNA → Protein. - Transcription: DNA is used as a template to synthesize RNA. - Translation: RNA (specifically mRNA) is used to synthesize proteins. - Genes are stretches of DNA that code for proteins, and the sequence of DNA determines the sequence of RNA, which in turn determines the sequence of amino acids in a protein.
Roles of Transcription and Translation
- Transcription: The process of making a complementary RNA copy from a DNA template. - Translation: The process of synthesizing proteins from mRNA by interpreting nucleotide sequences as amino acids. - In eukaryotes, RNA processing occurs, converting pre-mRNA into mature mRNA. - Some genes encode functional RNAs (structural or regulatory) rather than polypeptides.
Exceptions to the Central Dogma
- Some genes code for RNAs that are not translated into proteins but have important cellular functions. - In certain viruses, information can flow from RNA back to DNA via reverse transcriptase.
Transcription: From DNA to RNA
Structure of a Gene
Genes contain several important regions: - Promoter: Signals the beginning of transcription. - Terminator: Signals the end of transcription. - Transcribed region: Contains the information specifying the amino acid sequence. - Regulatory sequence: Site for binding regulatory proteins that influence transcription rate. 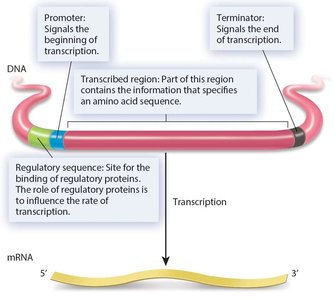
Stages of Transcription
Transcription occurs in three main stages: - Initiation: RNA polymerase recognizes the promoter region (in bacteria, aided by sigma factor) and forms an open complex. - Elongation: RNA polymerase synthesizes RNA using the template strand in the 3’ to 5’ direction, producing RNA in the 5’ to 3’ direction. Uracil (U) replaces thymine (T) in RNA. - Termination: RNA polymerase reaches a termination sequence, causing dissociation of the polymerase and RNA transcript from DNA. 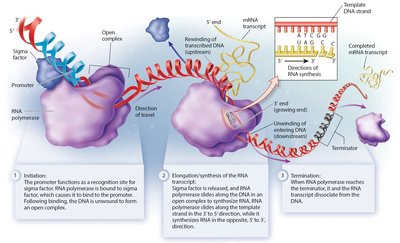
Direction of Transcription
- The direction of transcription and the DNA strand used as a template can vary among genes. - RNA synthesis always occurs in the 5’ to 3’ direction, with the DNA template read 3’ to 5’. 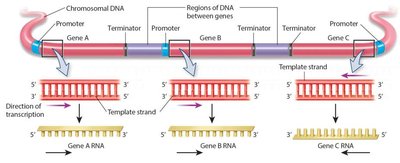
Eukaryotic Transcription
- Eukaryotes have three forms of RNA polymerase: RNA polymerase II (mRNA), I (rRNA), and III (tRNA and other nonstructural RNAs). - RNA polymerase II requires five general transcription factors to initiate transcription. 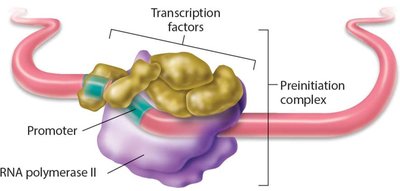
RNA Modification in Eukaryotes
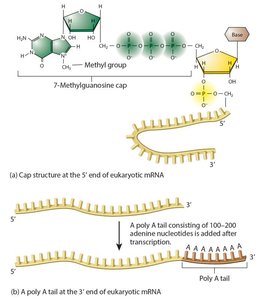
- Eukaryotic mRNAs are initially synthesized as pre-mRNA and require processing to become mature mRNA. - Introns: Non-coding sequences transcribed but not translated. - Exons: Coding sequences found in mature mRNA. - Splicing: Removal of introns. - Capping: Addition of a modified guanosine to the 5’ end. - Poly A tail: Addition of 100–200 adenine nucleotides to the 3’ end, increasing stability. 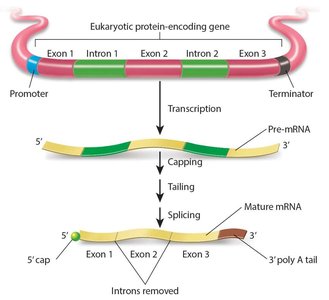
RNA Splicing
- Introns are removed by the spliceosome, a complex of snRNPs (small nuclear RNA + proteins). - Alternative splicing allows a single gene to produce multiple protein products. - Some RNAs (rRNA and tRNA) are self-splicing and act as ribozymes. 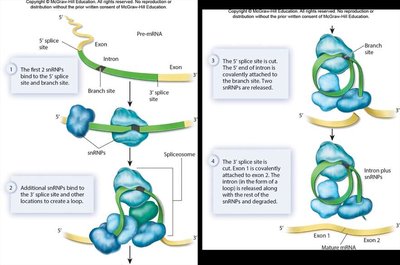
The Genetic Code and Translation
The Genetic Code Hypothesis
Francis Crick proposed that the sequence of bases in DNA acts as a code specifying the 20 amino acids. - The information in DNA is not directly translated into proteins but uses RNA intermediaries (mRNA, tRNA, rRNA).
Cracking the Genetic Code
- Nirenberg and Matthaei used synthetic RNA in cell-free systems to determine which codons specify which amino acids. - Example: PolyU RNA codes for phenylalanine. 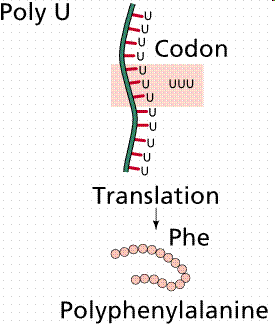
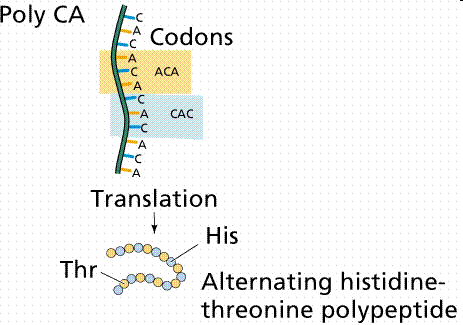
Properties of the Genetic Code
- Redundant: Most amino acids are encoded by more than one codon. - Unambiguous: Each codon specifies only one amino acid. - Non-overlapping: Codons are read one at a time. - Nearly universal: Codons specify the same amino acids in almost all organisms. - Conservative: Codons for the same amino acid often share the first two bases. 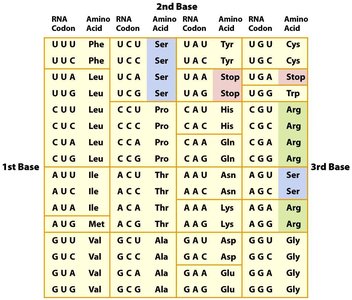
Codons and Anticodons
- Codon: Three-base sequence in mRNA specifying an amino acid. - Anticodon: Three-base sequence in tRNA complementary to the codon. 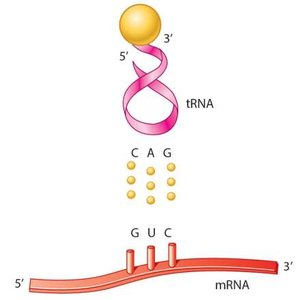
Reading Frame and Frameshift Mutations
- The start codon defines the reading frame for translation. - Insertion or deletion of bases can shift the reading frame, altering the amino acid sequence (frameshift mutation).
Translation: From mRNA to Protein
Components Required for Translation
- mRNA: Carries the genetic code. - tRNA: Brings amino acids to the ribosome. - Ribosomes: Site of protein synthesis. - Translation factors: Assist in the process.
tRNA Structure and Function
- tRNAs have a cloverleaf structure with an anticodon and an acceptor stem for amino acid binding. - Aminoacyl-tRNA synthetase catalyzes the attachment of amino acids to tRNA, ensuring specificity (the "second genetic code"). 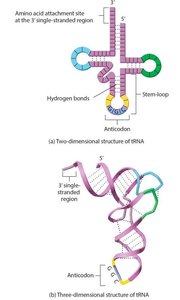
Ribosome Structure
- Prokaryotes have one kind of ribosome; eukaryotes have distinct ribosomes in different compartments. - Ribosomes are composed of large and small subunits and have discrete sites for tRNA binding: P (peptidyl), A (aminoacyl), and E (exit). 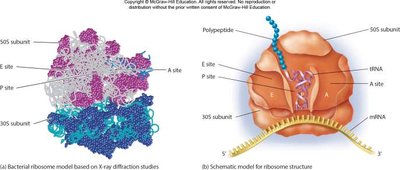
Stages of Translation
Translation occurs in three stages: - Initiation: Assembly of mRNA, the first tRNA, and ribosomal subunits. Requires initiation factors and GTP hydrolysis. 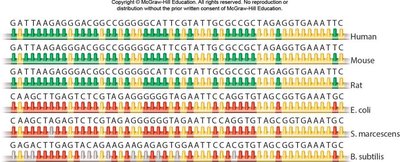
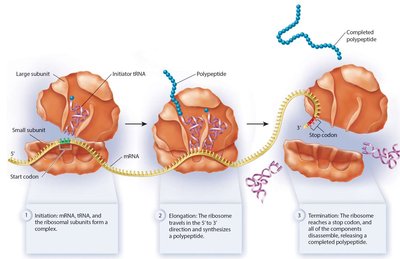
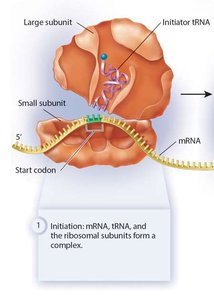 - Elongation: Aminoacyl tRNA brings new amino acids to the A site, peptide bonds are formed, and the ribosome translocates along the mRNA.
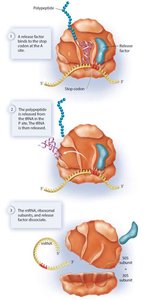
- Elongation: Aminoacyl tRNA brings new amino acids to the A site, peptide bonds are formed, and the ribosome translocates along the mRNA. 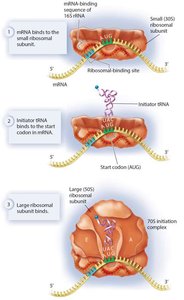 - Termination: When a stop codon is reached, release factors bind, the polypeptide is released, and the ribosomal complex disassembles.
- Termination: When a stop codon is reached, release factors bind, the polypeptide is released, and the ribosomal complex disassembles. 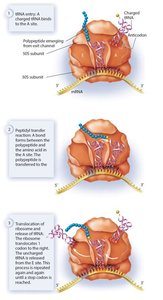
Evolutionary Conservation of Translational Components
- The gene for small subunit rRNA (SSU rRNA) is found in all genomes and is highly conserved. - Similar sequences are found in more closely related species, providing a basis for evolutionary relationships. 
Summary Table: Key Steps in Gene Expression
Step | Process | Key Molecules | Location |
|---|---|---|---|
Transcription | DNA → RNA | RNA polymerase, transcription factors | Nucleus (eukaryotes), cytoplasm (prokaryotes) |
RNA Processing | Pre-mRNA → Mature mRNA | Spliceosome, capping enzymes, poly A polymerase | Nucleus (eukaryotes) |
Translation | mRNA → Protein | Ribosome, tRNA, aminoacyl-tRNA synthetase | Cytoplasm |
Key Equations and Concepts
Central Dogma: Transcription Direction: Translation Direction: Genetic Code: Frameshift Mutation: Aminoacyl-tRNA Synthetase Reaction: Peptidyl Transfer Reaction:
Conclusion
Understanding gene expression at the molecular level is essential for comprehending how genetic information is translated into functional proteins, which in turn determine cellular structure, function, and organismal traits. The central dogma, genetic code, and mechanisms of transcription and translation are foundational concepts in molecular biology. ----------------------------------------
