 Back
BackMolecular Interactions, Cellular Compartmentation, and Calcium Homeostasis: ANP Study Notes
Study Guide - Smart Notes

Molecular Interactions and Cellular Compartmentation
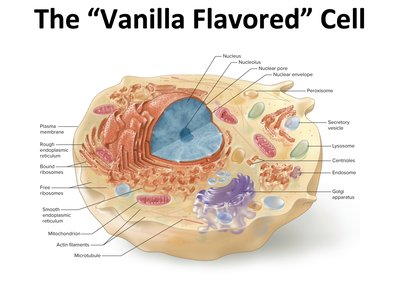
The "Vanilla Flavored" Cell
The basic structure of a eukaryotic cell includes several organelles, each with specialized functions essential for cellular life. Understanding these components is foundational for physiology and molecular biology.
Nucleus: Contains genetic material (DNA) and is the site of transcription.
Endoplasmic Reticulum (ER): Rough ER is studded with ribosomes for protein synthesis; smooth ER is involved in lipid synthesis and detoxification.
Golgi Apparatus: Modifies, sorts, and packages proteins and lipids for secretion or delivery to other organelles.
Mitochondria: The powerhouse of the cell, responsible for ATP production through cellular respiration.
Other organelles: Lysosomes, peroxisomes, cytoskeleton (microtubules, actin filaments), and various vesicles.
Cell Types and Specialization
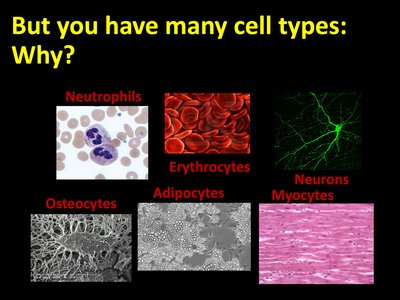
Cells in the human body differentiate into various types, each with unique structures and functions. This diversity allows for the specialization necessary for complex physiological processes.
Neutrophils: White blood cells involved in immune defense.
Erythrocytes: Red blood cells responsible for oxygen transport.
Neurons: Cells specialized for electrical signaling in the nervous system.
Osteocytes: Mature bone cells involved in bone maintenance.
Adipocytes: Fat storage cells.
Myocytes: Muscle cells responsible for contraction.
What Determines Cell Function?
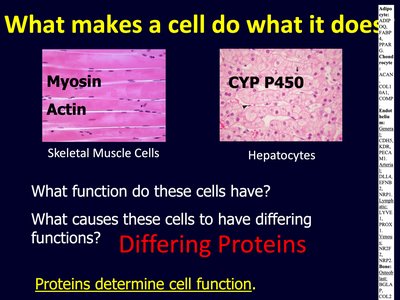
Cell function is determined by the specific proteins it expresses. Different cell types express different sets of proteins, which confer unique properties and functions.
Proteins such as myosin and actin are abundant in muscle cells, enabling contraction.
Enzymes like CYP P450 are found in hepatocytes (liver cells), facilitating detoxification.
Key Point: Proteins determine cell function. The presence or absence of specific proteins can have profound effects on cell behavior and organismal physiology.
Impact of Protein Loss
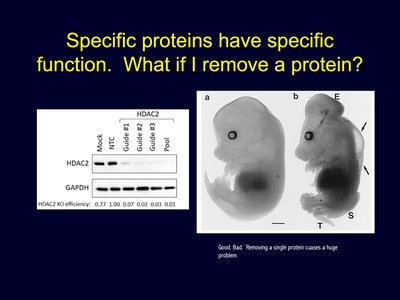
Removing a single protein can disrupt normal cellular and organismal function, highlighting the specificity and importance of protein expression.
Example: Knockout of the HDAC2 protein leads to significant developmental abnormalities.
Molecular Basis of Genetic Information
Nucleotides and DNA Structure
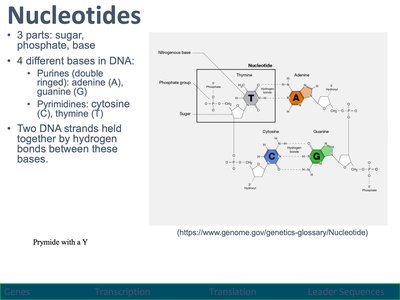
Nucleotides are the building blocks of DNA and RNA, consisting of a sugar, phosphate group, and nitrogenous base. DNA contains four bases: adenine (A), guanine (G), cytosine (C), and thymine (T).
Purines: Adenine and guanine (double-ringed).
Pyrimidines: Cytosine and thymine (single-ringed).
DNA strands are held together by hydrogen bonds between complementary bases.
Base Pairing in DNA
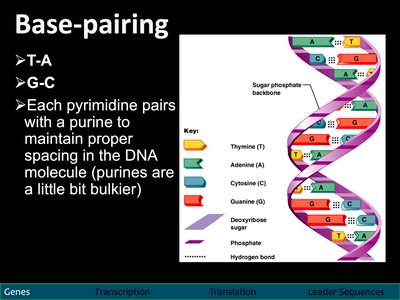
Base pairing ensures the stability and fidelity of the DNA double helix.
A pairs with T (adenine with thymine)
G pairs with C (guanine with cytosine)
Each pyrimidine pairs with a purine to maintain proper spacing.
The Gene
A gene is a stretch of DNA that codes for the synthesis of a specific polypeptide or protein. The sequence of bases determines the amino acid sequence of the protein.
Each set of three bases (codon) codes for one amino acid.
The base sequence is unique for each protein.

From Gene to Protein: The Central Dogma
Overview of the Central Dogma
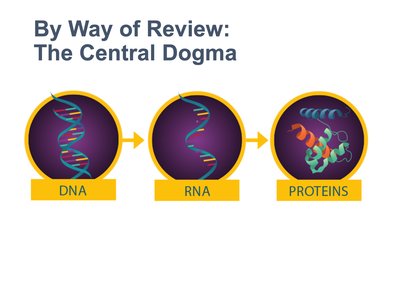
The central dogma of molecular biology describes the flow of genetic information from DNA to RNA to protein.
Transcription: DNA is transcribed into messenger RNA (mRNA).
Translation: mRNA is translated into a protein by ribosomes.
Transcription: From DNA to mRNA
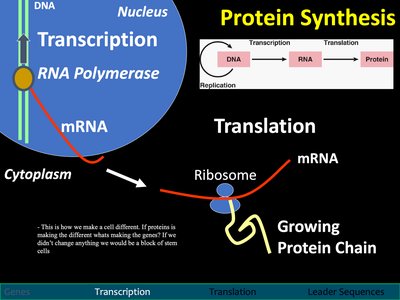
Transcription is the process by which a segment of DNA is copied into mRNA by the enzyme RNA polymerase.
Occurs in the nucleus.
Requires transcription factors to initiate and regulate gene expression.
mRNA carries the genetic code from the nucleus to the cytoplasm for translation.
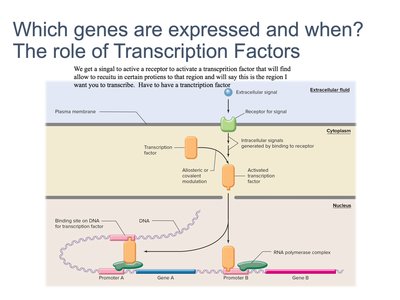
Regulation of Gene Expression: Transcription Factors
Transcription factors are proteins that bind to specific DNA sequences, controlling the rate of transcription of genetic information from DNA to mRNA.
Signals activate transcription factors, which bind to promoter regions of genes.
Transcription factors determine which genes are expressed and when.
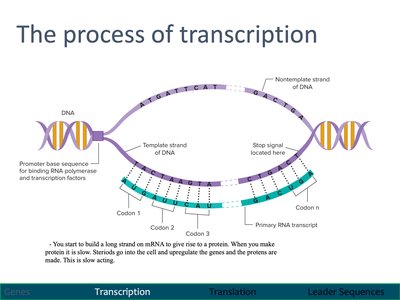
The Process of Transcription
During transcription, RNA polymerase synthesizes a complementary RNA strand from the DNA template.
Promoter regions signal the start of transcription.
Transcription proceeds until a stop signal is reached.
The resulting primary RNA transcript may undergo splicing to remove introns.
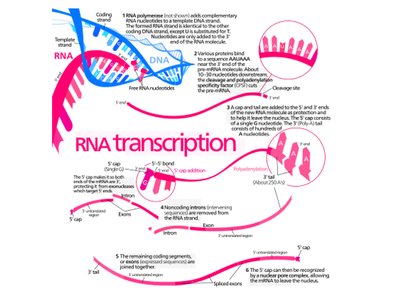
RNA Processing
After transcription, the primary RNA transcript is processed to become mature mRNA.
5' capping, 3' polyadenylation, and splicing are key steps.
Splicing removes non-coding introns, joining coding exons together.

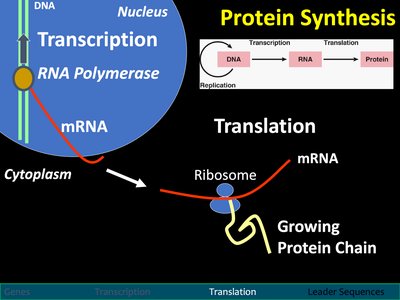
Translation: From mRNA to Protein
Translation is the process by which ribosomes synthesize proteins using the sequence encoded in mRNA.
Occurs in the cytoplasm at ribosomes.
Each codon (three nucleotides) on mRNA specifies an amino acid.
tRNA molecules bring amino acids to the ribosome, matching codons with anticodons.
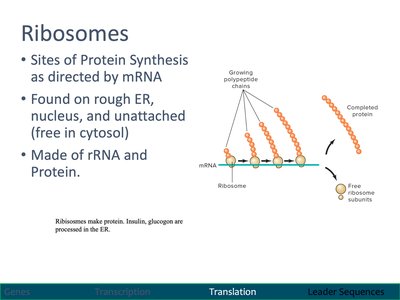
Ribosomes: Sites of Protein Synthesis
Ribosomes are molecular machines made of rRNA and protein, responsible for translating mRNA into polypeptide chains.
Found on rough ER, in the cytosol, and in the nucleus.
Consist of large and small subunits that assemble on mRNA.
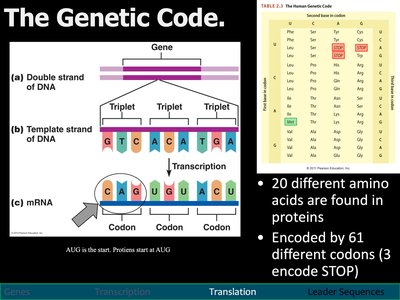
The Genetic Code
The genetic code is the set of rules by which information encoded in mRNA is translated into proteins.
There are 20 amino acids and 61 codons that encode them (3 codons signal stop).
The start codon (AUG) signals the beginning of translation.
Translation: Initiation
Translation initiation involves the assembly of the ribosome on the mRNA and the recruitment of the first tRNA.
The small ribosomal subunit scans for the start codon (AUG).
tRNA carrying methionine binds to the start codon.
The large subunit joins, and elongation begins.
Calcium Homeostasis: Feedback Loops
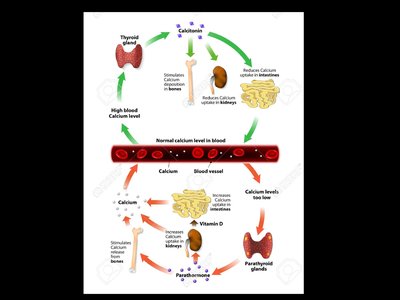
Regulation of Blood Calcium Levels
Blood calcium levels are tightly regulated by negative feedback loops involving the parathyroid and thyroid glands. Disruption of this regulation can lead to muscular and nervous system disorders.
Low blood calcium: Detected by the parathyroid gland, which releases parathyroid hormone (PTH).
PTH actions: Stimulates osteoclasts to release calcium from bone, increases calcium reabsorption in kidneys, and enhances calcium absorption in the intestines (via vitamin D activation).
High blood calcium: Detected by the thyroid gland, which releases calcitonin.
Calcitonin actions: Inhibits osteoclasts, stimulates calcium deposition in bones, and increases calcium excretion in urine.
Feedback Loop Components
Each feedback loop consists of receptors, control centers, and effectors:
Feedback Loop | Receptor | Control Center | Effector |
|---|---|---|---|
Low Calcium (PTH) | Parathyroid gland | Parathyroid gland | Osteoclasts, kidneys, intestines |
High Calcium (Calcitonin) | Thyroid gland | Thyroid gland | Osteoblasts, kidneys |
Example: When calcium is low, the parathyroid gland senses the change (receptor and control center), releases PTH, which acts on osteoclasts, kidneys, and intestines (effectors) to increase blood calcium.

Identifying the Receptor for Low Calcium
In the context of low blood calcium, the receptor is the parathyroid gland, which senses the decrease and initiates the corrective response.
Correct answer: Parathyroid gland

Additional info:
Some slides reference the central dogma and gene expression, which are foundational for understanding how cells produce the proteins that mediate physiological processes, including those involved in calcium homeostasis and cellular compartmentation.
