 Back
BackAmino Acids, Peptides, and Proteins: Structure, Properties, and Function
Study Guide - Smart Notes

AMINO ACIDS, PEPTIDES, AND PROTEINS
Overview of Protein Functions
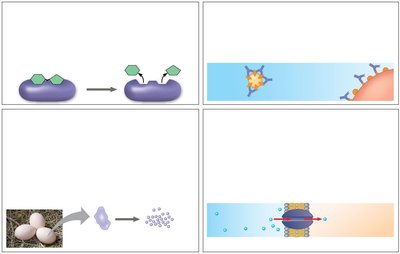
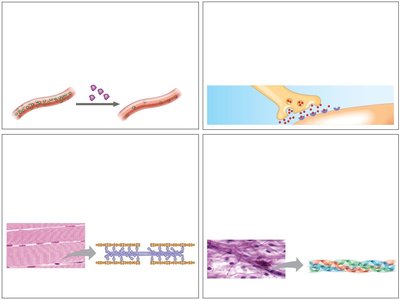
Proteins are essential macromolecules that perform a vast array of functions in biological systems. Their diversity in structure allows them to catalyze reactions, provide structural support, transport molecules, and more.
Enzymatic proteins: Catalyze biochemical reactions, e.g., digestive enzymes hydrolyze food molecules.
Defensive proteins: Protect against disease, e.g., antibodies neutralize pathogens.
Storage proteins: Store amino acids, e.g., casein in milk, ovalbumin in eggs.
Transport proteins: Move substances, e.g., hemoglobin transports oxygen, membrane proteins transport molecules across membranes.
Hormonal proteins: Coordinate activities, e.g., insulin regulates blood sugar.
Receptor proteins: Respond to chemical stimuli, e.g., nerve cell receptors detect signaling molecules.
Contractile and motor proteins: Enable movement, e.g., actin and myosin in muscles.
Structural proteins: Provide support, e.g., keratin in hair, collagen in connective tissue.
AMINO ACIDS: BUILDING BLOCKS OF PROTEINS
General Structure and Properties
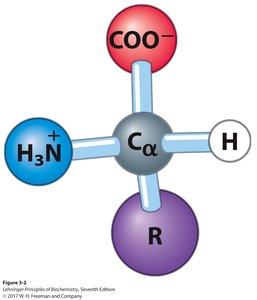
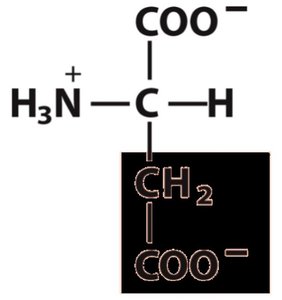
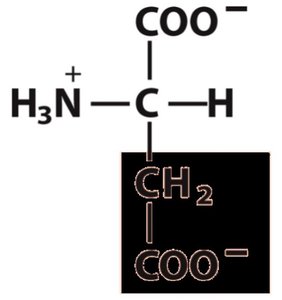
Amino acids are the monomeric units of proteins. Each amino acid contains a central (α) carbon atom bonded to an amino group, a carboxyl group, a hydrogen atom, and a unique side chain (R group).
α-carbon: Tetrahedral geometry with four different substituents (except glycine).
Functional groups: Acidic carboxyl group, basic amino group, hydrogen, and variable R group.
Polymerization: Amino acids link via peptide bonds to form polypeptides.
Classification of Amino Acids
Amino acids are classified based on the properties of their R groups:
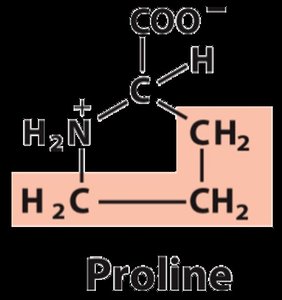
Nonpolar, aliphatic: Glycine, Alanine, Proline, Valine, Leucine, Isoleucine, Methionine
Aromatic: Phenylalanine, Tyrosine, Tryptophan
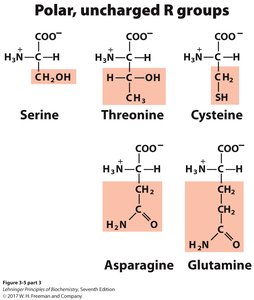
Polar, uncharged: Serine, Threonine, Cysteine, Asparagine, Glutamine
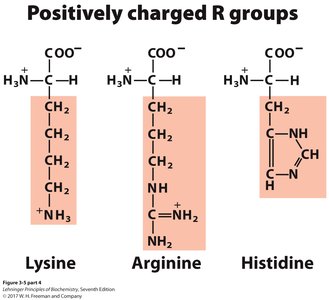
Positively charged (basic): Lysine, Arginine, Histidine
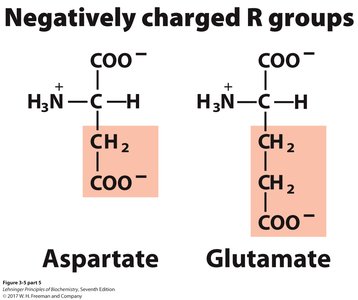
Negatively charged (acidic): Aspartate, Glutamate
v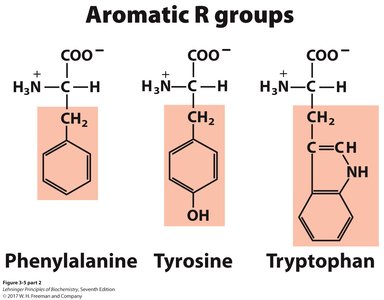
Properties and Notation
Three-letter and one-letter codes: Used for shorthand notation in sequences (e.g., Gly, G).
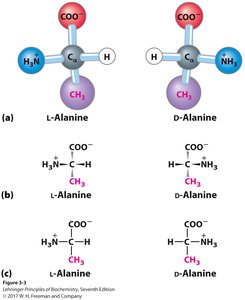
Chirality: All amino acids (except glycine) are chiral; proteins contain only L-amino acids.
Ionization and Acid-Base Properties
Amino acids contain at least two ionizable groups (α-carboxyl and α-amino), each with its own pKa value. The ionization state depends on the pH of the environment.
At low pH: Amino acids are fully protonated (cationic form).
At high pH: Amino acids are fully deprotonated (anionic form).
Zwitterion: At intermediate pH, amino acids exist as zwitterions (net charge = 0).
Buffering: Amino acids can act as buffers near their pKa values.
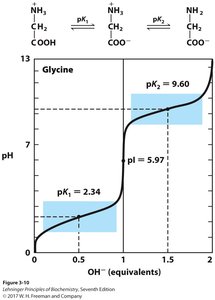
Isoelectric Point (pI)
The isoelectric point is the pH at which the net charge of the amino acid is zero. For amino acids without ionizable side chains:
Formula:
At pI: Amino acid is least soluble and does not migrate in an electric field.
For amino acids with ionizable side chains, the pI is calculated using the two pKa values that surround the zwitterionic form.
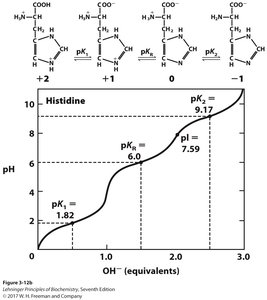
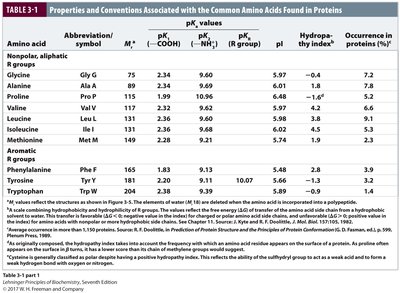
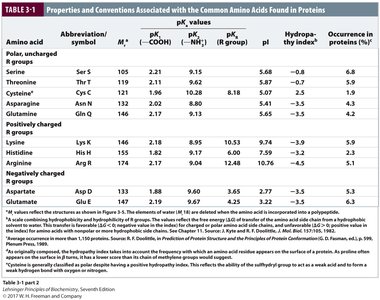
Properties Table of Amino Acids
The following tables summarize the properties, abbreviations, pKa values, and occurrence of the common amino acids found in proteins.
Amino acid | Abbreviation | Mr | pK1 (COOH) | pK2 (NH3+) | pKR | pI | Occurrence (%) |
|---|---|---|---|---|---|---|---|
Glycine | Gly, G | 75 | 2.34 | 9.60 | - | 5.97 | 7.2 |
Alanine | Ala, A | 89 | 2.34 | 9.69 | - | 6.01 | 8.3 |
Phenylalanine | Phe, F | 165 | 1.83 | 9.13 | - | 5.48 | 3.9 |
Serine | Ser, S | 105 | 2.21 | 9.15 | - | 5.68 | 6.8 |
Lysine | Lys, K | 146 | 2.18 | 8.95 | 10.53 | 9.74 | 5.9 |
Aspartate | Asp, D | 133 | 1.88 | 9.60 | 3.65 | 2.77 | 5.3 |
PEPTIDES AND PROTEINS
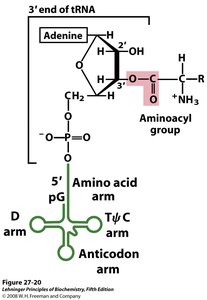
Peptide Bond Formation
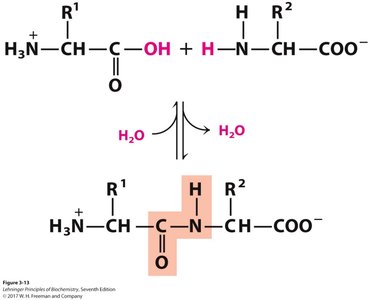
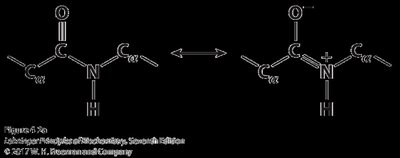
Peptides are formed by the condensation of amino acids, resulting in an amide (peptide) bond. The peptide bond is planar due to resonance, restricting rotation and contributing to protein structure.
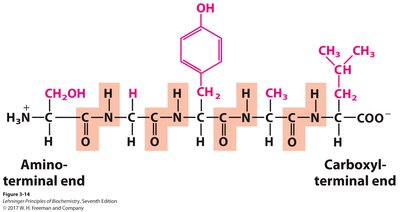
Directionality: Peptide chains are written from the N-terminus (amino end) to the C-terminus (carboxyl end).
Primary structure: The unique sequence of amino acids in a protein.
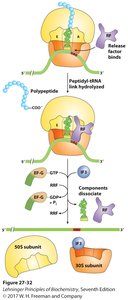
Protein Synthesis
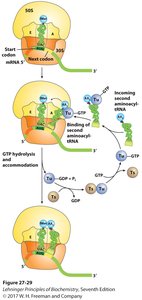
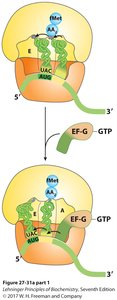
Proteins are synthesized in cells using mRNA as a template. Ribosomes facilitate the assembly of amino acids into polypeptides via tRNA molecules.
Transcription: DNA is transcribed to mRNA.
Translation: mRNA is translated into a polypeptide chain.
PROTEIN STRUCTURE: HIERARCHY AND STABILIZATION
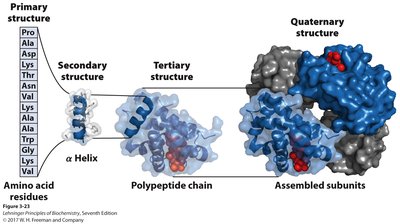
Levels of Protein Structure
Primary structure: Linear sequence of amino acids.
Secondary structure: Local folding into α-helices and β-sheets, stabilized by hydrogen bonds.
Tertiary structure: Overall 3D shape of a single polypeptide, stabilized by hydrophobic interactions, hydrogen bonds, ionic interactions, and disulfide bonds.
Quaternary structure: Assembly of multiple polypeptide subunits into a functional protein complex.
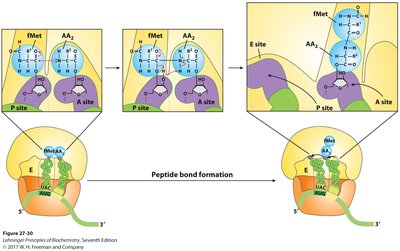
Peptide Bond Properties
The peptide bond is a resonance hybrid, making it planar and rigid. This restricts rotation and influences the folding of the polypeptide chain.
Secondary Structure: α-Helix and β-Sheet
α-Helix: Right-handed coil stabilized by hydrogen bonds between every fourth amino acid. Side chains project outward.
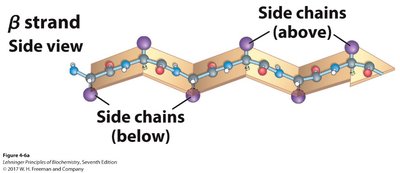
β-Sheet: Extended strands connected laterally by hydrogen bonds. Can be parallel or antiparallel.
Tertiary and Quaternary Structure
Tertiary structure is the overall 3D arrangement of a polypeptide, while quaternary structure refers to the assembly of multiple polypeptides. Stabilizing forces include hydrophobic interactions, hydrogen bonds, ionic interactions, and disulfide bridges.
FIBROUS PROTEINS AND STRUCTURE-FUNCTION RELATIONSHIPS
Fibrous Proteins
Fibrous proteins have structural roles and are characterized by regular secondary structures. Examples include α-keratin (hair, nails), collagen (connective tissue), and silk fibroin (spider silk).
Structure | Characteristics | Examples |
|---|---|---|
α-Helix, cross-linked by disulfide bonds | Tough, insoluble, variable hardness | α-Keratin (hair, feathers, nails) |
β-Conformation | Soft, flexible filaments | Silk fibroin |
Collagen triple helix | High tensile strength, no stretch | Collagen (tendons, bone) |
Protein Analysis Techniques
Purification: Based on charge, size, affinity, solubility, or hydrophobicity.
Sequencing: Edman degradation (classical), mass spectrometry (modern).
SUMMARY
Proteins are polymers of amino acids with diverse structures and functions.
Amino acids are classified by their side chains and have characteristic ionization properties.
Protein structure is hierarchical: primary, secondary, tertiary, and quaternary levels.
Fibrous proteins illustrate the relationship between structure and function.
Modern techniques allow for the purification and sequencing of proteins.
