 Back
BackBiochemistry Study Guide: Metabolism, Enzymes, Membranes, and Carbohydrates
Study Guide - Smart Notes

Metabolic Pathways and Thermodynamics
Free Energy and Spontaneity
Metabolic reactions are governed by thermodynamic principles, particularly the concept of free energy. The Gibbs free energy change () determines whether a reaction is spontaneous or not.
Exergonic reactions: , spontaneous, release energy.
Endergonic reactions: , non-spontaneous, require energy input.
Equation: (where is enthalpy, is temperature, is entropy)
Standard free energy change:
Example: ATP hydrolysis is exergonic and drives many cellular processes.
Redox Reactions and Coenzymes
Oxidation-Reduction Principles
Redox reactions are central to metabolism, involving electron transfer between molecules. Substances that lose electrons are oxidized and act as reducing agents, while those that gain electrons are reduced and act as oxidizing agents.
UNA SUSTANCIA | O2 | H2 | Valencia | e- | SE COMPORTA COMO |
|---|---|---|---|---|---|
SE OXIDA cuando | gana | pierde | aumenta | pierde | REDUCTOR |
SE REDUCE cuando | pierde | gana | disminuye | gana | OXIDANTE |
Coenzymes: NAD and NADP
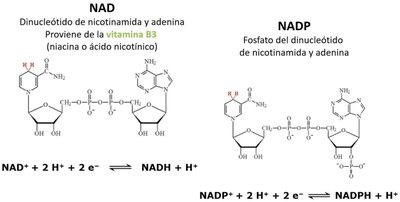
NAD and NADP are essential coenzymes in redox reactions. They accept electrons and protons, becoming reduced forms (NADH, NADPH) that carry energy for biosynthetic and catabolic pathways.
NAD: Derived from vitamin B3 (niacin), involved in catabolic reactions.
NADP: Phosphorylated NAD, involved in anabolic reactions.
Reduction reactions:
Enzyme Catalysis and Regulation
Enzyme Function and Classification
Enzymes are biological catalysts that accelerate reactions by lowering activation energy. Most enzymes are proteins, though some RNA molecules (ribozymes) also have catalytic activity.
Enzyme-substrate complex: Temporary association between enzyme and substrate.
Enzyme classes: Oxidoreductases, Transferases, Hydrolases, Lyases, Isomerases, Ligases.
Cofactors: Metal ions or organic molecules (coenzymes) required for activity.
Enzyme Inhibition
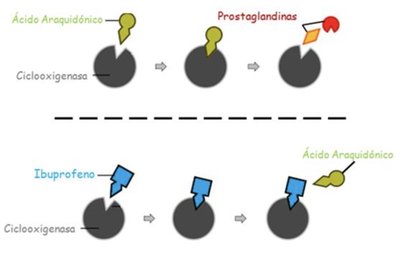
Enzyme activity can be regulated by inhibitors. Competitive inhibitors bind to the active site, preventing substrate binding. Non-competitive inhibitors bind elsewhere, altering enzyme function.
Example: Ibuprofen competitively inhibits cyclooxygenase, blocking prostaglandin synthesis from arachidonic acid.
Biological Membranes
Membrane Structure and Composition
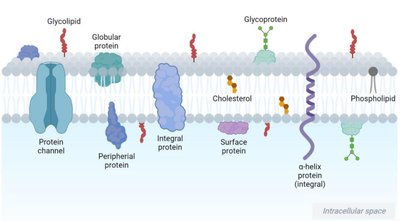
Cell membranes are dynamic, semipermeable barriers composed of lipids, proteins, and carbohydrates. The lipid bilayer provides structural integrity and fluidity, while proteins mediate transport and signaling.
Lipids: Phospholipids, cholesterol, sphingolipids.
Proteins: Integral, peripheral, and surface proteins.
Carbohydrates: Glycoproteins and glycolipids for cell recognition.
Phospholipids and Lipid Classes
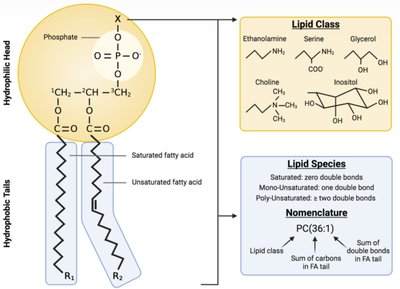
Phospholipids are amphipathic molecules with hydrophilic heads and hydrophobic tails. Variations in head groups and fatty acid saturation define lipid classes and species.
Head groups: Ethanolamine, serine, choline, inositol, glycerol.
Fatty acid tails: Saturated (no double bonds), unsaturated (one or more double bonds).
Nomenclature: PC(36:1) indicates phosphatidylcholine with 36 carbons and 1 double bond.
Membrane Lipid Distribution
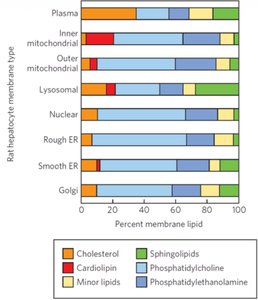
The composition of membrane lipids varies by organelle and cell type, affecting membrane properties and function.
Plasma membrane: High cholesterol, sphingolipids.
Mitochondrial membranes: Cardiolipin, phosphatidylethanolamine.
ER and Golgi: Phosphatidylcholine, minor lipids.
Membrane Transport
Transport Mechanisms
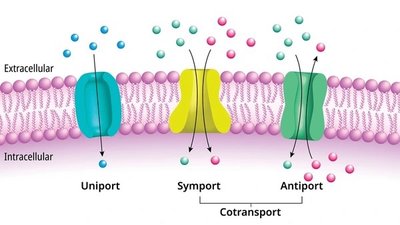
Membranes regulate the movement of molecules via passive and active transport.
Passive transport: Diffusion (simple and facilitated), osmosis.
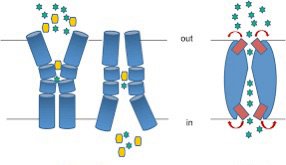
Facilitated diffusion: Requires protein channels or carriers for larger or polar molecules.
Active transport: Requires ATP to move substances against concentration gradients.
Carbohydrates: Structure and Function
Monosaccharides, Disaccharides, and Polysaccharides
Carbohydrates are classified by their functional groups and number of units.
Monosaccharides: Simple sugars (glucose, fructose), classified as aldoses or ketoses.
Disaccharides: Two monosaccharides joined by glycosidic bonds (maltose, lactose, sucrose).
Polysaccharides: Long chains (starch, glycogen, cellulose, chitin).
Artificial vs. Natural Sweeteners
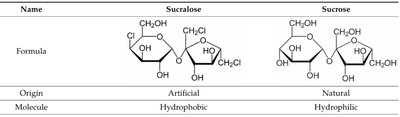
Sucralose is an artificial sweetener structurally similar to sucrose but is not metabolized, making it non-caloric.
Name | Formula | Origin | Molecule |
|---|---|---|---|
Sucralose | See structure | Artificial | Hydrophobic |
Sucrose | See structure | Natural | Hydrophilic |
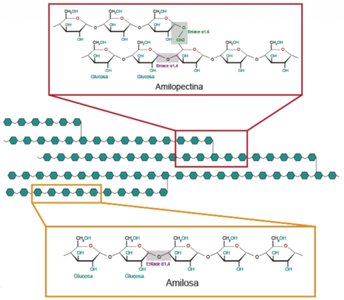
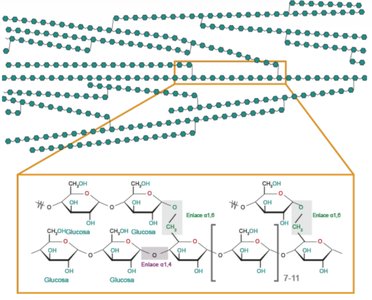
Starch: Amylopectin and Amylose
Starch is a plant polysaccharide composed of amylopectin (branched) and amylose (linear). Glycogen is a similar storage polysaccharide in animals.
Amylopectin: Glucose units with α(1,4) and α(1,6) linkages (branched).
Amylose: Linear glucose units with α(1,4) linkages.
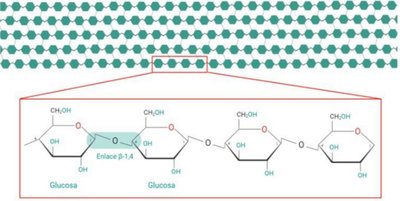
Cellulose Structure
Cellulose is a linear polysaccharide of glucose with β(1,4) linkages, not metabolized by humans but important as dietary fiber.
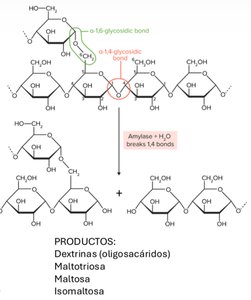
Enzymatic Digestion of Carbohydrates
Amylase hydrolyzes α(1,4) glycosidic bonds in starch, producing dextrins, maltotriose, maltose, and isomaltose.
Glucose Transporters (GLUT)
GLUT Isoforms and Tissue Distribution
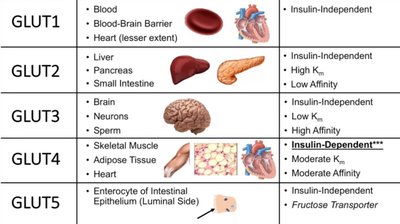
GLUT proteins facilitate glucose transport across membranes. Isoforms differ in tissue expression, affinity (Km), and insulin dependence.
GLUT Isoform | Tissue | Insulin Dependence | Km | Affinity |
|---|---|---|---|---|
GLUT1 | Blood, BBB, Heart | Independent | Low | High |
GLUT2 | Liver, Pancreas, Intestine | Independent | High | Low |
GLUT3 | Brain, Neurons, Sperm | Independent | Low | High |
GLUT4 | Muscle, Adipose, Heart | Dependent | Moderate | Moderate |
GLUT5 | Intestinal Epithelium | Independent | Fructose transporter | - |
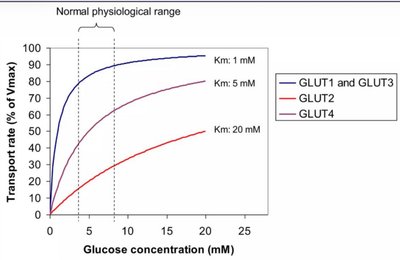
GLUT Kinetics
Transport rate depends on glucose concentration and transporter affinity (Km). Lower Km indicates higher affinity.
Example: GLUT4 is insulin-dependent and mediates glucose uptake in muscle and adipose tissue.
