Back
BackBiochemistry Study Notes: Acid-Base Chemistry, Buffers, Amino Acids, Protein Structure, and Hemoglobin
Study Guide - Smart Notes

Acid-Base Chemistry and Buffers
Acid Dissociation and Equilibrium
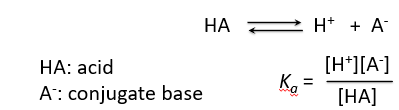
Acids dissociate in water to produce hydrogen ions and their conjugate bases. The equilibrium constant for this dissociation is known as the acid dissociation constant, Ka.
HA: Acid
A-: Conjugate base
Equilibrium:
Derivation of the Henderson-Hasselbalch Equation
The Henderson-Hasselbalch equation relates pH, pKa, and the ratio of conjugate base to acid, providing a practical way to calculate pH in buffer solutions.
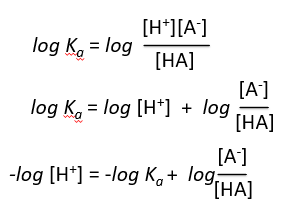
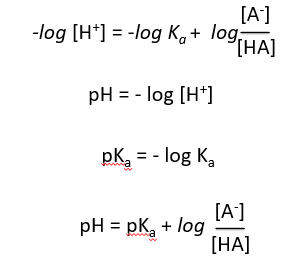
Buffer Systems and Polyprotic Acids
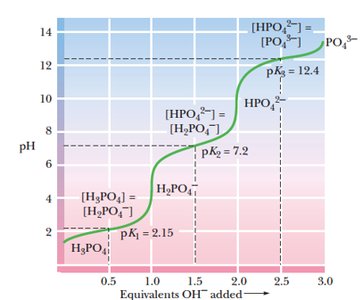
Buffers are solutions that resist changes in pH upon addition of acid or base. Polyprotic acids, such as phosphoric acid, have multiple dissociation steps, each with its own pKa and buffering region.
Buffering occurs near each pKa value.
Phosphoric acid has three pKa values: 2.15, 7.2, and 12.4.
Each region corresponds to a different equilibrium between protonated and deprotonated forms.
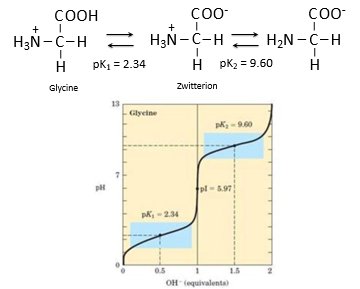
Amino Acid Titration and Zwitterions
Amino acids can act as both acids and bases, and their titration curves show distinct buffering regions. Glycine, for example, has two pKa values and exists as a zwitterion at intermediate pH.
pK1 = 2.34 (carboxyl group)
pK2 = 9.60 (amino group)
Zwitterion: molecule with both positive and negative charges
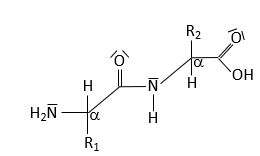
Peptide Bond Structure and Resonance
Peptide Bond Characteristics
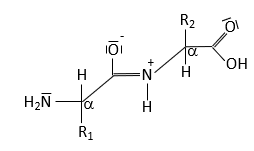
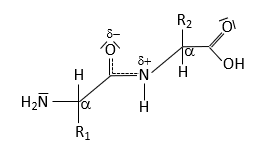
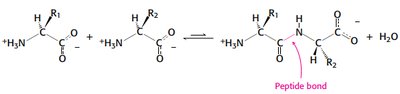
The peptide bond forms between the α-carboxyl group of one amino acid and the α-amino group of another, resulting in partial double-bond character due to resonance. This restricts rotation and influences protein structure.
Resonance leads to partial double-bond character.
No rotation around the peptide bond; rotation occurs around adjacent single bonds.
Torsion angles determine polypeptide chain path.
Amino Acids and pH Effects
Protonation States of Amino Acids
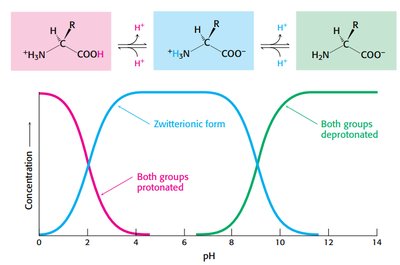
The protonation state of amino acids changes with pH, affecting their charge and structure. At low pH, both groups are protonated; at intermediate pH, the zwitterion dominates; at high pH, both groups are deprotonated.
Isoelectric point: pH at which amino acid exists as a zwitterion.
Charge transitions affect protein folding and function.
Amino Acids as Buffers
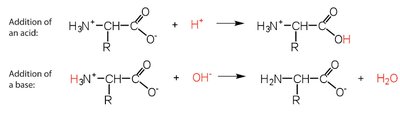
Amino acids can buffer solutions by accepting or donating protons, helping maintain pH stability.
Addition of acid: carboxylate group captures proton.
Addition of base: amino group loses proton.
Blood Buffer System
Bicarbonate Buffer and Physiological Regulation
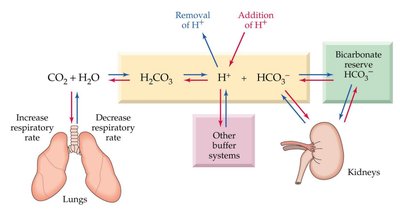
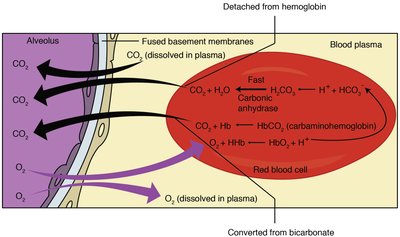
The bicarbonate buffer system maintains blood pH within a narrow range. It involves dynamic regulation by the lungs and kidneys, as well as contributions from proteins and phosphate buffers.
CO2 + H2O → H2CO3 → H+ + HCO3-
Respiratory and renal systems adjust CO2 and HCO3- levels.
Blood pH is tightly regulated (7.35–7.45).
Protein Structure
Primary Structure: Peptide Bond Formation
Proteins are linear polymers of amino acids linked by peptide bonds, formed via condensation reactions.
Peptide bond formation releases water.
Sequence determines protein function and folding.
Secondary Structure: Alpha Helices and Beta Sheets
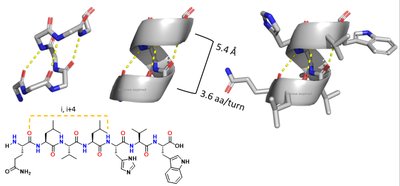
Secondary structure arises from hydrogen bonding between backbone atoms, forming alpha helices and beta sheets.
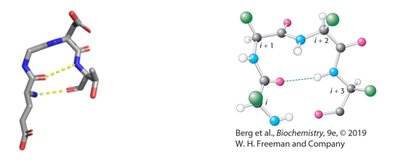
Alpha helix: right-handed, 3.6 residues per turn, stabilized by i to i+4 hydrogen bonds.
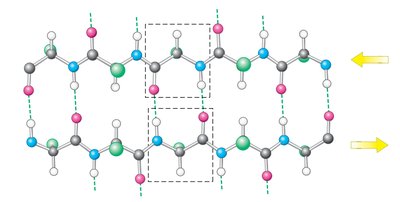
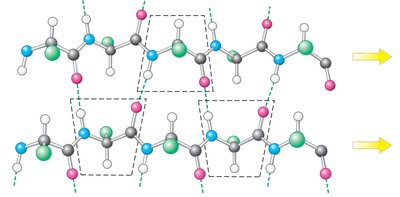
Beta sheet: extended strands, hydrogen bonds between adjacent strands, can be parallel or antiparallel.
Hemoglobin and Myoglobin
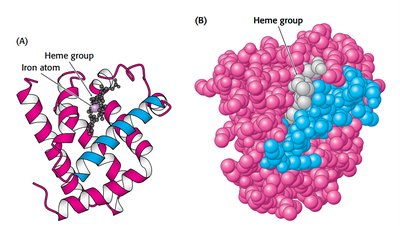
Myoglobin Structure and Function
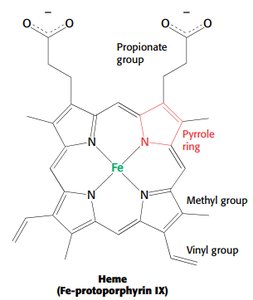
Myoglobin is a single-chain oxygen storage protein in muscle, containing a heme group with a central iron atom. The heme structure is crucial for oxygen binding and release.
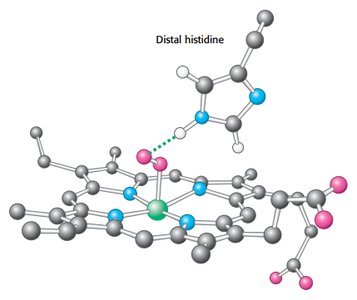
Heme: Fe-protoporphyrin IX, binds oxygen at the iron atom.
Distal histidine stabilizes bound oxygen via hydrogen bonding.
Hemoglobin Structure and Cooperative Binding
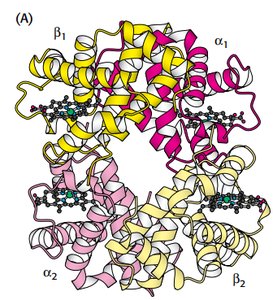
Hemoglobin is a tetrameric protein with two alpha and two beta chains, each containing a heme group. Oxygen binding is cooperative, meaning binding at one site increases affinity at others.
Quaternary structure: α1β1 and α2β2 dimers.
T state (tense): low affinity; R state (relaxed): high affinity.
2,3-BPG stabilizes T state, facilitating oxygen release.
Sickle Cell Disease
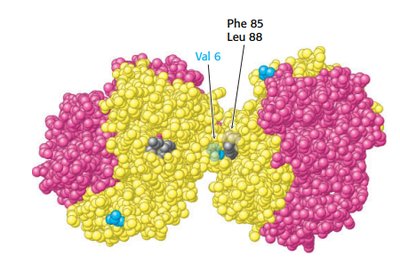
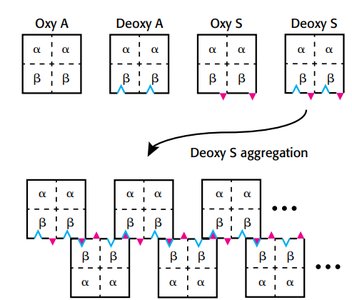
Sickle cell anemia results from a mutation in the hemoglobin beta chain, substituting valine for glutamate at position 6. This creates a hydrophobic patch, leading to aggregation in the deoxy form.
Aggregation occurs only in deoxyhemoglobin S.
Leads to sickling of red blood cells and clinical symptoms.
Bohr Effect and Oxygen Saturation Curves
Bohr Effect
The Bohr effect describes how hemoglobin releases oxygen in response to lower pH and higher CO2 concentrations in tissues.
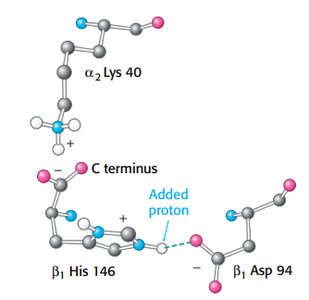
Hydrogen ions and CO2 stabilize T state, promoting oxygen release.
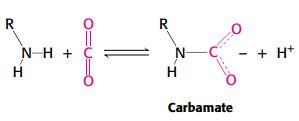
Salt bridges and carbamate formation are key mechanisms.
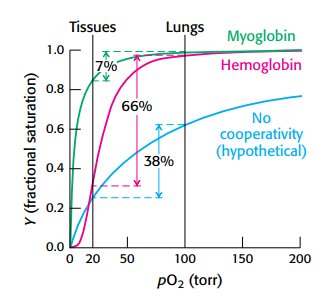
Oxygen Saturation Curves
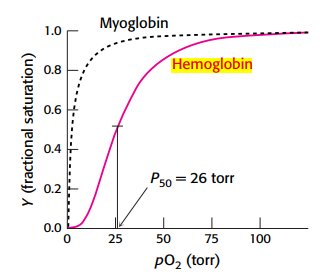
Oxygen binding curves illustrate the differences in affinity and cooperativity between myoglobin and hemoglobin.
Myoglobin: hyperbolic curve, high affinity, no cooperativity.
Hemoglobin: sigmoidal curve, cooperative binding.
P50: partial pressure at which 50% of sites are occupied (26 torr for hemoglobin).
Models of Cooperativity
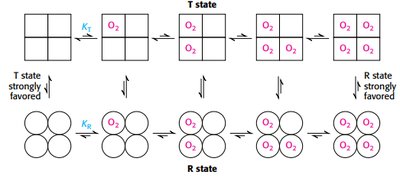
Cooperativity in hemoglobin is explained by the concerted and sequential models, describing transitions between T and R states.
Concerted model: all subunits switch together.
Sequential model: subunits switch individually, increasing affinity stepwise.
Collagen Structure
Collagen Triple Helix
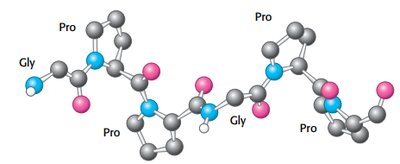
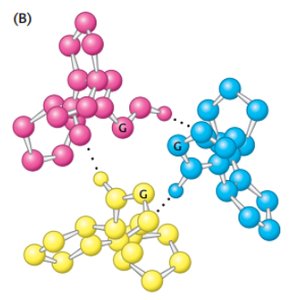
Collagen is a fibrous protein with a unique triple helix structure, stabilized by glycine, proline, and hydroxyproline residues.
Glycine at every third residue allows tight packing.
Hydrogen bonds and steric repulsion stabilize the helix.