 Back
BackEnzyme Catalysis and Carbohydrate Structure: Key Concepts in Biochemistry
Study Guide - Smart Notes

Enzyme Catalysis
Introduction to Catalysts and Enzymes
Catalysts are substances that increase the rate of chemical reactions without being consumed in the process. In biological systems, enzymes are highly specific protein catalysts that accelerate biochemical reactions, often by many orders of magnitude, making life possible under physiological conditions.
Catalyst: Increases reaction rate by lowering activation energy, but does not alter the equilibrium position.
Enzyme: A biological catalyst, typically a protein, that facilitates specific biochemical reactions.
Transition State Stabilization: Enzymes lower the activation energy by stabilizing the transition state of the reaction.
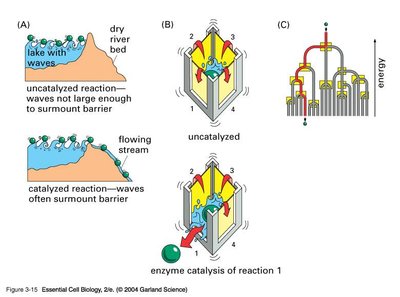
Mechanisms of Enzyme Action
Enzymes accelerate reactions by providing an alternative reaction pathway with a lower activation energy. They achieve this through several mechanisms, including proximity and orientation effects, transition state stabilization, and induced fit.
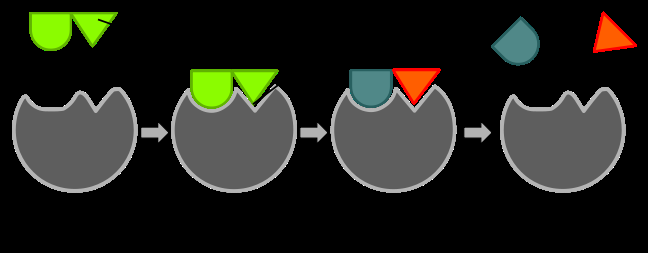
Proximity and Orientation: Enzymes bring substrates close together and orient them optimally for reaction, increasing the likelihood of productive collisions.
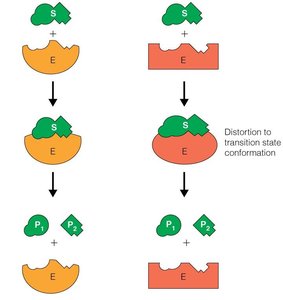
Induced Fit Model: The enzyme changes shape upon substrate binding, enhancing the fit and stabilizing the transition state.
Lock-and-Key Model: The substrate fits into the enzyme's active site without significant conformational change (less favored than induced fit).
Michaelis-Menten Kinetics
The Michaelis-Menten equation describes the rate of enzymatic reactions as a function of substrate concentration:
Vmax: Maximum reaction velocity at saturating substrate concentration.
Km: Substrate concentration at which the reaction rate is half of Vmax; reflects enzyme affinity for substrate.
Kcat: Turnover number; the number of substrate molecules converted to product per enzyme molecule per second.
Michaelis-Menten equation (in LaTeX):
Where is the initial velocity, is substrate concentration, is the maximum velocity, and is the Michaelis constant.
Chemical Mechanisms in Enzyme Catalysis
Enzymes employ several chemical strategies to catalyze reactions:
Nucleophilic and Electrophilic Catalysis: Involvement of electron-rich (nucleophile) and electron-poor (electrophile) groups.
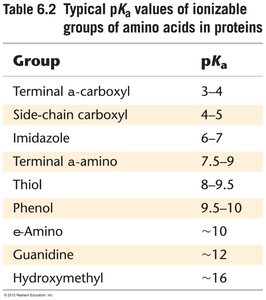
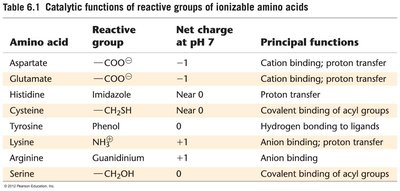
Acid-Base Catalysis: Amino acid side chains act as proton donors or acceptors.
Covalent Catalysis: Formation of a transient covalent bond between enzyme and substrate.
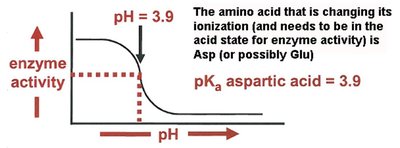
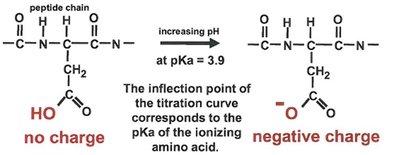
Effect of pH: Enzyme activity is sensitive to pH, which affects the ionization state of active site residues.
Serine Protease Mechanism and the Catalytic Triad
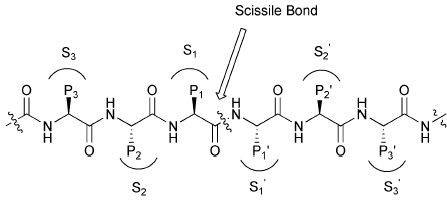
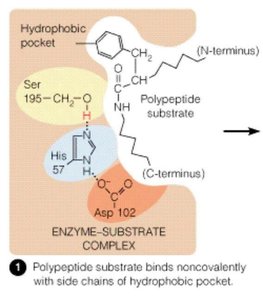
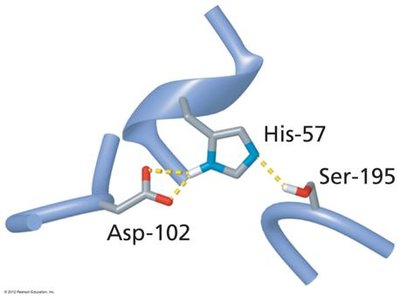
Serine proteases, such as chymotrypsin, use a catalytic triad (Ser195, His57, Asp102) to cleave peptide bonds. The mechanism involves:
Substrate binding in the active site, positioning the scissile bond for attack.
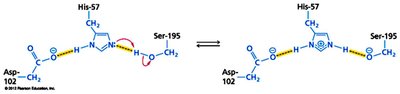
Activation of Ser195 by His57 (which is stabilized by Asp102), enabling nucleophilic attack on the peptide bond.
Formation of a tetrahedral intermediate and subsequent breakdown to release products.
Enzyme Inhibition
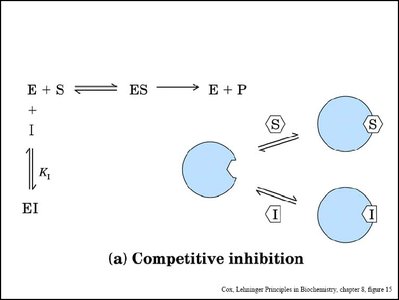
Enzyme inhibitors are molecules that decrease or abolish enzyme activity. They are classified as reversible (competitive, noncompetitive, uncompetitive) or irreversible (mechanism-based/suicide inhibitors).
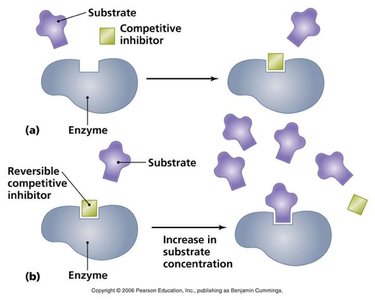
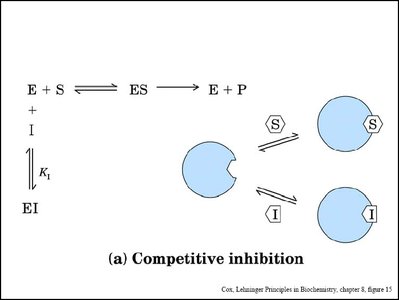
Competitive Inhibition: Inhibitor binds to the active site, competing with the substrate. Increases apparent Km, Vmax unchanged.
Noncompetitive (Mixed) Inhibition: Inhibitor binds to enzyme or enzyme-substrate complex at a site other than the active site. Decreases Vmax, Km may change.
Uncompetitive Inhibition: Inhibitor binds only to the enzyme-substrate complex, decreasing both Km and Vmax.
Regulation of Enzyme Activity
Enzyme activity is regulated to control metabolic pathways. Common regulatory mechanisms include:
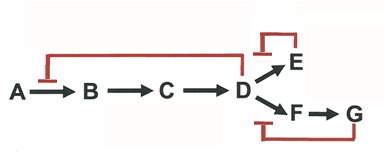
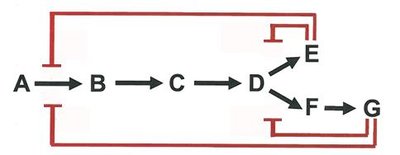
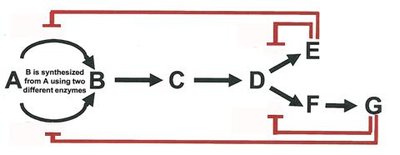
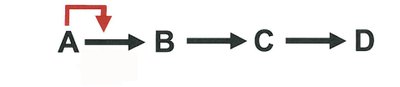
Feedback Inhibition: End product of a pathway inhibits an early enzyme, preventing overproduction.
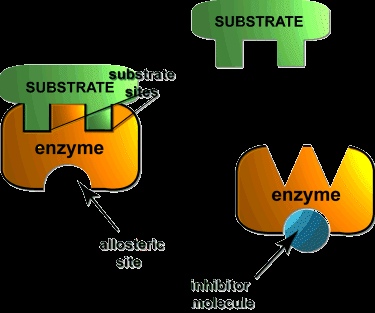
Allosteric Regulation: Effector molecules bind to sites other than the active site, altering enzyme activity.
Activation by Substrate Accumulation: Substrate presence can activate enzymes, often via allosteric mechanisms.
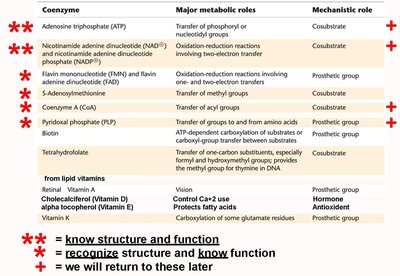
Coenzymes and Essential Ions
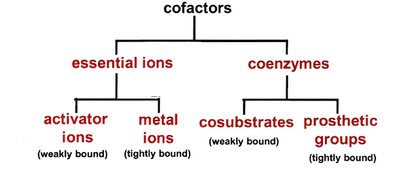
Many enzymes require non-protein cofactors for activity. These include:
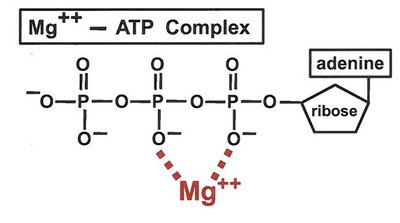
Essential Ions: Metal ions that are either loosely (activator ions) or tightly (metal ions) bound.
Coenzymes: Organic molecules, often derived from vitamins, that participate in catalysis as cosubstrates or prosthetic groups.
Carbohydrate Structure and Stereochemistry
Classification and Structure of Monosaccharides
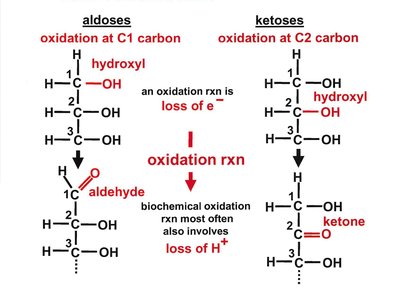
Carbohydrates are classified based on the number of carbon atoms and the type of carbonyl group present:
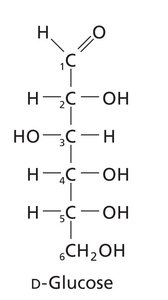
Aldoses: Contain an aldehyde group (e.g., glucose).
Ketoses: Contain a ketone group (e.g., fructose).
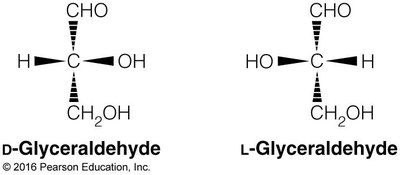
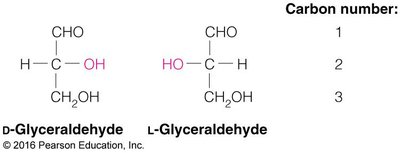
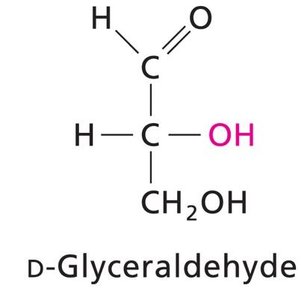
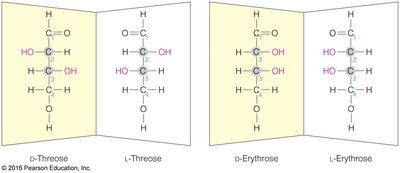
Chirality: Most monosaccharides are chiral, existing as D- or L-enantiomers.
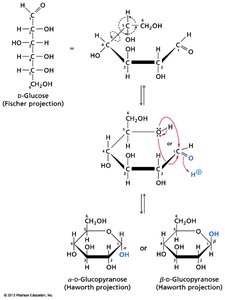
Cyclization and Anomeric Forms
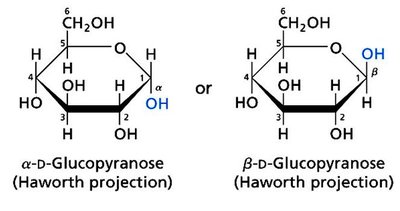
Monosaccharides with five or more carbons can cyclize to form ring structures, generating a new chiral center at the anomeric carbon. The two possible configurations are called anomers (α and β).
Pyranose: Six-membered ring (e.g., glucose).
Furanose: Five-membered ring (e.g., fructose).
Haworth Projection: A common way to represent cyclic sugars.
Stereoisomerism in Carbohydrates
Carbohydrates exhibit several types of stereoisomerism:
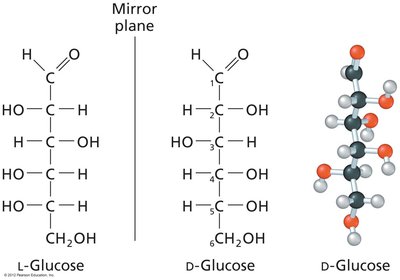
Enantiomers: Non-superimposable mirror images (D- and L-forms).
Diastereomers: Stereoisomers that are not mirror images.
Epimers: Differ at only one chiral center.
Anomers: Differ at the anomeric carbon (α vs. β).
Conformational Isomers: Differ by rotation around single bonds, no bond breaking required.
Example: D-glucose and D-mannose are epimers at C2; D-glucose and D-galactose are epimers at C4.
*Additional info: The notes above are expanded with academic context to ensure clarity and completeness for biochemistry students preparing for exams.*
