 Back
BackFundamental Chemical Principles and Biomolecular Structure in Biochemistry
Study Guide - Smart Notes

Chemical Foundations of Biomolecules
Common Functional Groups in Biomolecules
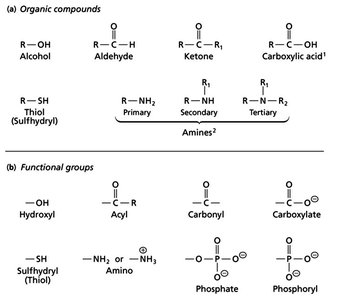
Biomolecules are composed of specific functional groups that determine their chemical properties and biological roles. Understanding these groups is essential for interpreting molecular interactions in biochemistry.
Alcohols: Contain a hydroxyl group (-OH) attached to a carbon atom. Example: ethanol.
Aldehydes: Have a carbonyl group (C=O) bonded to a hydrogen and a carbon. Example: formaldehyde.
Ketones: Feature a carbonyl group bonded to two carbons. Example: acetone.
Carboxylic Acids: Possess a carboxyl group (-COOH), which is ionized at physiological pH.
Thiols (Sulfhydryl): Contain an -SH group, important in protein structure (e.g., cysteine).
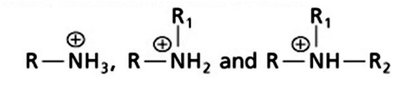
Amines: Classified as primary, secondary, or tertiary based on the number of carbon groups attached to the nitrogen. At physiological pH, amines are typically protonated (ammonium ions).
Phosphate and Phosphoryl Groups: Central to energy transfer and signaling in cells.
Classes of Biological Macromolecules
Biological macromolecules are categorized based on their monomeric units and functions:
Proteins: Polymers of amino acids, perform structural, catalytic, and regulatory roles.
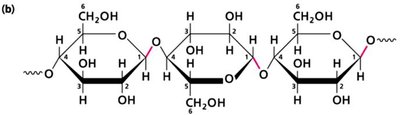
Carbohydrates: Polymers of sugars, serve as energy sources and structural components.
Nucleic Acids: DNA and RNA, polymers of nucleotides, store and transmit genetic information.
Lipids: Fatty acids and derivatives, form membranes and store energy.
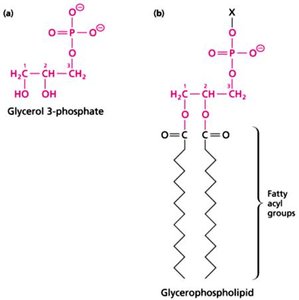
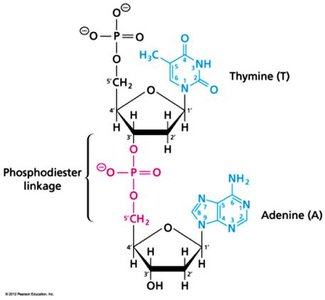
Covalent and Non-Covalent Interactions in Biomolecules
Types of Chemical Bonds
Biomolecular structure and function depend on both covalent and non-covalent interactions:
Covalent Bonds: Strong bonds formed by sharing electron pairs; essential for molecular stability.
Non-Covalent Interactions: Include charge-charge (ionic), hydrogen bonds, van der Waals forces, and hydrophobic interactions. These are weaker but crucial for molecular recognition and structure.
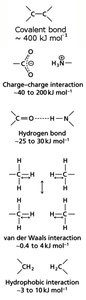
Hydrogen Bonds in Biomolecular Interactions
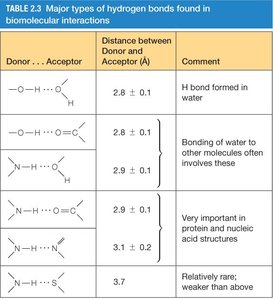
Hydrogen bonds are directional interactions between a hydrogen atom covalently bonded to an electronegative atom (donor) and another electronegative atom (acceptor). They stabilize protein and nucleic acid structures.
Donor...Acceptor | Distance (Å) | Comment |
|---|---|---|
O-H...O | 2.8 ± 0.1 | H bond formed in water |
O-H...N | 2.8 ± 0.1 | Bonding of water to other molecules |
N-H...O | 2.9 ± 0.1 | Important in protein/nucleic acid structures |
N-H...N | 3.1 ± 0.1 | Less common |
S-H...O | 3.7 | Rare; weaker |
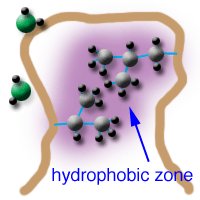
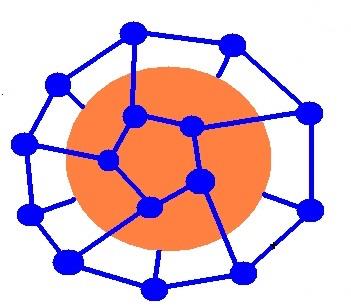
Hydrophobic Effects
Hydrophobic interactions arise when non-polar molecules aggregate to minimize contact with water, increasing system entropy. These effects drive protein folding, membrane formation, and molecular recognition.
Key roles: protein folding, enzyme-substrate binding, lipid bilayer formation.
Water: Structure, Properties, and Ionization
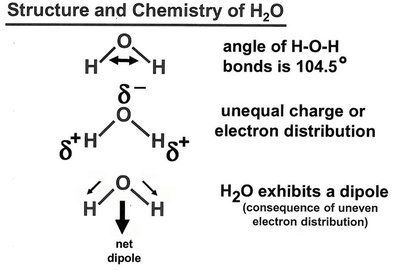
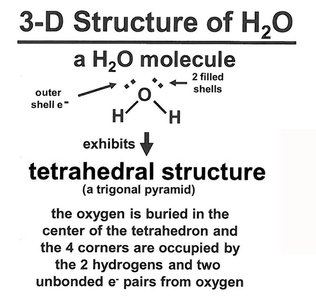
Structure and Chemistry of Water
Water is a polar molecule with a bent geometry (104.5° bond angle), resulting in a net dipole moment. Its tetrahedral structure allows extensive hydrogen bonding, making it an excellent solvent for polar and ionic compounds.
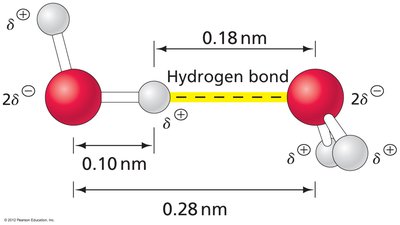
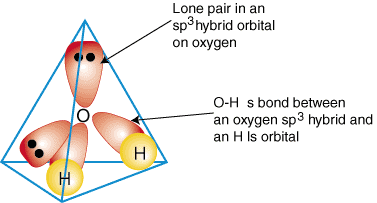
Ionization of Water and pH
Water can ionize to form H+ and OH- ions. The equilibrium constant (Keq) for this reaction is used to define the ion product of water (Kw).
At 25°C, M,
Relationship:
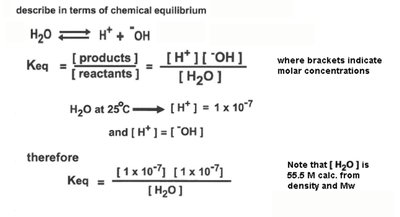
Acids, Bases, and Buffer Systems
Acids are proton donors, bases are proton acceptors (Lowry-Bronsted definition). The strength of an acid is characterized by its dissociation constant (Ka) and pKa value.
Strong acids: Completely dissociate in water (e.g., HCl).
Weak acids: Partially dissociate (e.g., acetic acid).
Henderson-Hasselbalch Equation: Relates pH, pKa, and the ratio of conjugate base to acid.
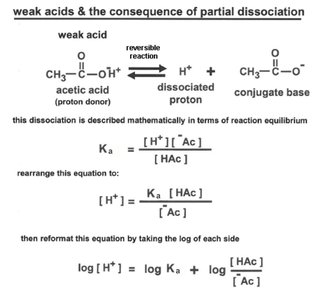
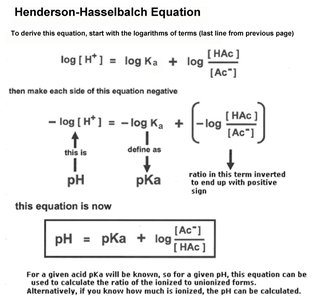
Titration Curves and Buffering
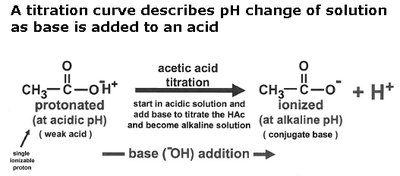
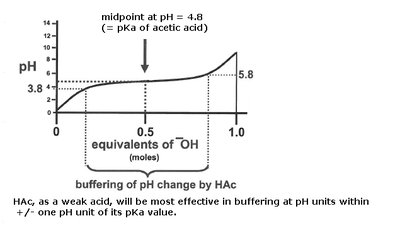
Titration curves illustrate how the pH of a solution changes as base is added to an acid. The midpoint of the curve corresponds to the pKa of the acid, where half of the acid is dissociated.
Buffering is most effective within ±1 pH unit of the pKa.
Amino Acids: Structure, Properties, and Ionization
General Structure of Amino Acids
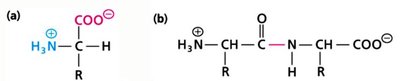
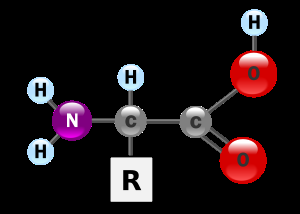
Amino acids are the building blocks of proteins. Each has a central alpha carbon (Cα) bonded to an amino group, a carboxyl group, a hydrogen atom, and a variable side chain (R group).
At physiological pH, amino acids exist as zwitterions (both positive and negative charges).
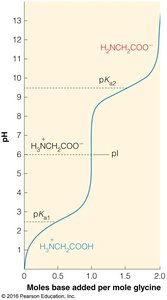
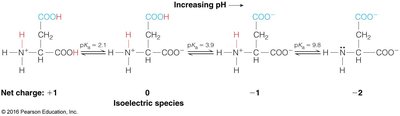
Isoelectric Point (pI) and pKa Values
The isoelectric point (pI) is the pH at which an amino acid carries no net charge. For amino acids with two ionizable groups:
For amino acids with more than two ionizable groups, pI is calculated as the average of the pKa values surrounding the neutral species.
Thermodynamics and Biochemical Reactions
Free Energy and Reaction Direction
The change in free energy (ΔG) determines whether a reaction is spontaneous (exergonic) or non-spontaneous (endergonic):
ΔG < 0: Spontaneous (exergonic)
ΔG > 0: Non-spontaneous (endergonic)
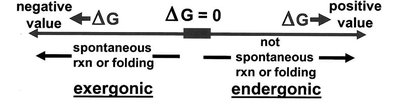
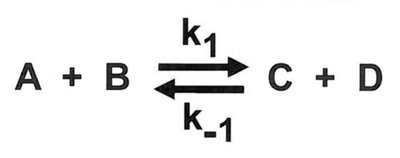
Equilibrium and Coupled Reactions
Biochemical reactions often occur in pathways, where unfavorable reactions are coupled to favorable ones to drive the overall process. The equilibrium constant (Keq) and reaction quotient (Q) are used to predict reaction direction.
At equilibrium: and
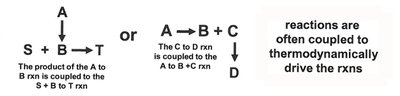
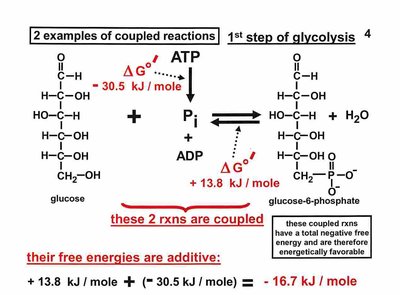
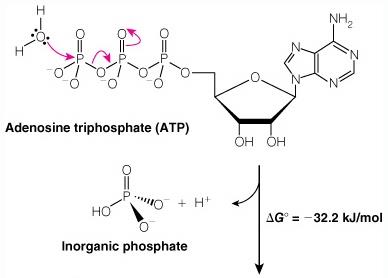
ATP and Phosphoryl Group Transfers
ATP is the universal energy currency in cells. Hydrolysis of ATP releases energy by transferring a phosphoryl group to water or other molecules.
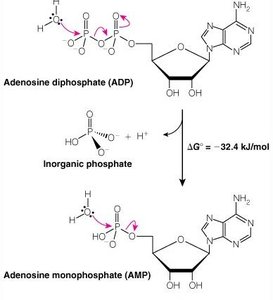
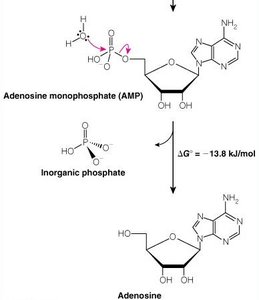
Oxidation-Reduction (Redox) Reactions
Redox Principles and Standard Reduction Potentials
Redox reactions involve electron transfer between molecules. The tendency of a molecule to gain electrons is measured by its standard reduction potential (Eo).
n = number of electrons transferred, F = Faraday constant (96.48 kJ V-1 mol-1)
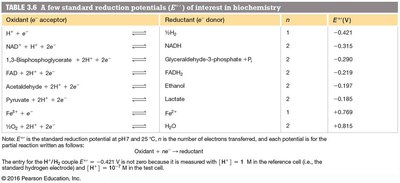
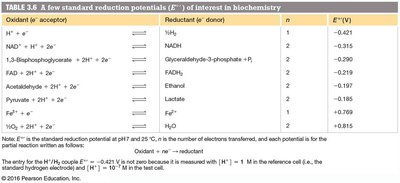
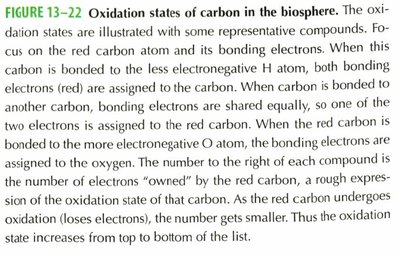
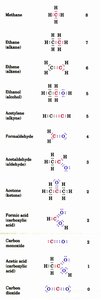
Oxidation States of Carbon
The oxidation state of carbon in organic molecules varies depending on its bonding partners. Carbon bonded to hydrogen is more reduced, while carbon bonded to oxygen is more oxidized.


Amino Acids: Structure and Spectral Properties
General Structure and Classification
All amino acids (except proline) share a common backbone structure. The side chain (R group) determines the chemical properties and classification (aliphatic, aromatic, polar, nonpolar, acidic, basic).
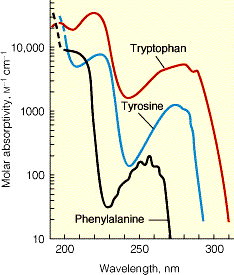
Spectral Properties of Aromatic Amino Acids
Aromatic amino acids (phenylalanine, tyrosine, tryptophan) absorb UV light, especially at 280 nm, which is useful for protein quantification and structural studies.
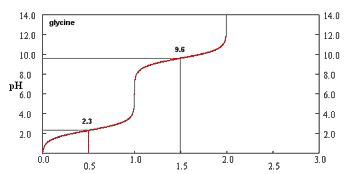
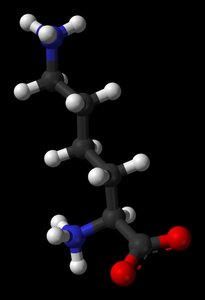
Titration of Amino Acids
Titration curves of amino acids reveal their ionization states at different pH values, with distinct pKa values for the amino and carboxyl groups.
Protein Structure: Primary to Quaternary
Levels of Protein Structure
Proteins exhibit hierarchical structural organization:
Primary Structure: Linear sequence of amino acids.
Secondary Structure: Local folding patterns (α-helix, β-sheet) stabilized by hydrogen bonds.
Tertiary Structure: Overall 3D folding of a single polypeptide chain.
Quaternary Structure: Association of multiple polypeptide chains into a functional complex.
Secondary Structure: α-Helix and β-Sheet
The α-helix is a right-handed coil stabilized by hydrogen bonds between the carbonyl oxygen of residue i and the amide hydrogen of residue i+4. The β-sheet consists of extended strands connected by hydrogen bonds, which can be parallel or antiparallel.
Ramachandran Plot
Ramachandran plots visualize the allowed conformations of φ and ψ dihedral angles in polypeptides, helping to predict secondary structure elements.
Protein Folding and Stability
Protein folding is driven by a balance of favorable internal interactions (hydrophobic effect, hydrogen bonds) and unfavorable entropy changes. The process is highly cooperative and results in a stable, functional structure.
Protein Function: Ligand Binding and Allostery
Ligand Binding and Affinity
Proteins bind ligands with specific affinity, described by the dissociation constant (Kd). Lower Kd values indicate higher affinity.
Oxygen Binding: Myoglobin vs. Hemoglobin
Myoglobin binds oxygen in muscle tissue with a hyperbolic binding curve, while hemoglobin binds oxygen cooperatively in blood, displaying a sigmoidal curve due to allosteric transitions between T (tense) and R (relaxed) states.
Allosteric Regulation
Allostery refers to the regulation of protein activity by binding of effectors at sites other than the active site. Hemoglobin is a classic example, with 2,3-BPG and pH (Bohr effect) modulating oxygen affinity.
Summary Table: Major Functional Groups in Biochemistry
Functional Group | Structure | Example |
|---|---|---|
Hydroxyl | -OH | Alcohols |
Carbonyl | C=O | Aldehydes, Ketones |
Carboxylate | -COO- | Carboxylic acids |
Amino | -NH2, -NH3+ | Amines, amino acids |
Phosphate | -PO42- | Nucleotides, ATP |
Sulfhydryl | -SH | Cysteine |
Additional info: These notes provide foundational knowledge for understanding the chemical basis of life, macromolecular structure, and the energetics of biochemical reactions, as covered in introductory biochemistry courses.
