 Back
BackFundamentals of Protein Structure: Levels, Elements, and Ramachandran Plots
Study Guide - Smart Notes

Fundamentals of Protein Structure
Overview of Protein Structure
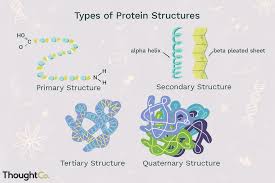
Proteins are complex biological macromolecules essential for life, and their function is determined by their structure. Protein structure is described at four hierarchical levels: primary, secondary, tertiary, and quaternary. Each level of structure contributes to the overall shape and function of the protein.
Primary Structure: The linear sequence of amino acids in a polypeptide chain.
Secondary Structure: Localized, recurring arrangements of amino acids, such as α-helices and β-sheets.
Tertiary Structure: The overall three-dimensional conformation of a single polypeptide chain.
Quaternary Structure: The association of multiple polypeptide chains into a functional protein complex.
Primary Structure
The primary structure of a protein is its unique sequence of amino acids, linked by peptide bonds. This sequence is determined by the gene encoding the protein and dictates all higher levels of structure.
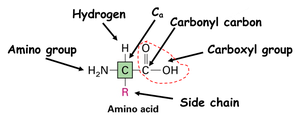
Amino Acid: The building block of proteins, consisting of a central carbon (α-carbon), an amino group, a carboxyl group, a hydrogen atom, and a variable side chain (R group).
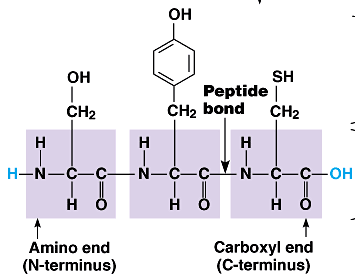
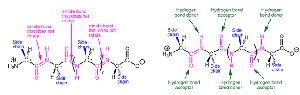
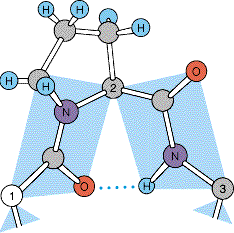
Peptide Bond: A covalent bond formed between the carboxyl group of one amino acid and the amino group of another.
N-terminus: The end of the polypeptide with a free amino group.
C-terminus: The end of the polypeptide with a free carboxyl group.
Secondary Structure
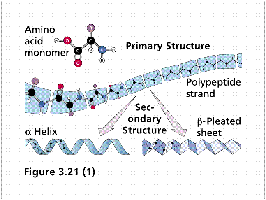
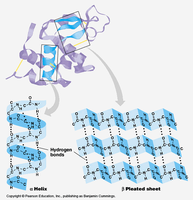
Secondary structure refers to regular, recurring arrangements of adjacent amino acid residues stabilized by hydrogen bonding. The most common elements are α-helices and β-sheets, along with turns and loops.
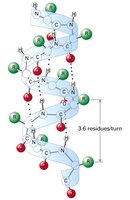
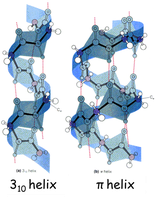
α-Helix: A right-handed coil stabilized by hydrogen bonds between backbone atoms. Each turn contains about 3.6 amino acid residues.
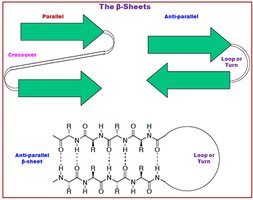
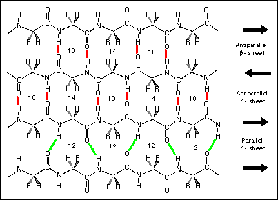
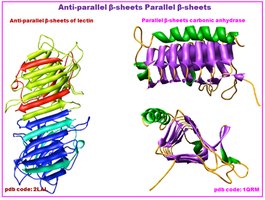
β-Sheet: Extended strands connected laterally by hydrogen bonds, forming parallel or antiparallel sheets.
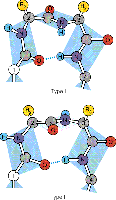
Turns and Loops: Short regions that connect secondary structure elements, often containing proline or glycine residues.
Phi (φ) and Psi (ψ) Angles: Torsional angles around the α-carbon that determine the folding of the polypeptide backbone.
Tertiary Structure
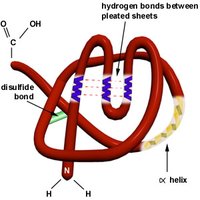
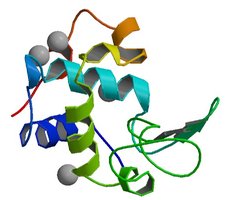
Tertiary structure is the overall three-dimensional shape of a single polypeptide chain, resulting from interactions between side chains (R groups) and secondary structure elements. It is stabilized by various forces, including hydrogen bonds, hydrophobic interactions, disulfide bonds, and electrostatic interactions.
Hydrophobic Interactions: Nonpolar side chains tend to cluster away from water, stabilizing the protein core.
Disulfide Bonds: Covalent bonds formed between cysteine residues, providing extra stability.
Electrostatic Interactions: Attractions between charged side chains.
Hydrogen Bonds: Between polar side chains and backbone atoms.
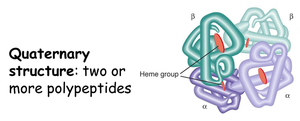
Quaternary Structure
Quaternary structure arises when two or more polypeptide chains (subunits) associate to form a functional protein complex. The arrangement and interaction of these subunits are crucial for the protein's biological activity.
Homodimer/Heterodimer: Complexes of identical or different subunits.
Examples: Hemoglobin (4 chains), insulin (2 chains), DNA polymerase (multiple chains).
Ramachandran Plot
The Ramachandran plot is a graphical representation of the allowed regions of phi (φ) and psi (ψ) angles in a polypeptide backbone. It is used to predict and evaluate protein folding and secondary structure.
Allowed Regions: Most residues fall within three main regions corresponding to α-helices, β-sheets, and left-handed helices.
Steric Hindrance: Not all angle combinations are possible due to atomic clashes.
Applications: Used in protein modeling and structure validation.
Secondary Structure | Typical φ Angle | Typical ψ Angle |
|---|---|---|
Right-handed α-helix | -60° | -45° |
β-sheet | -120° | +120° |
Left-handed α-helix | +60° | +45° |
Special Amino Acids and Disulfide/Diselenide Bridges
Some amino acids, such as cysteine and selenocysteine, play unique roles in protein structure. Cysteine forms disulfide bridges, while selenocysteine can form diselenide bridges, both contributing to protein stability.
Cysteine: Forms disulfide bonds under oxidizing conditions.
Selenocysteine: Similar to cysteine, but contains selenium; forms diselenide bridges.
Bond Type | Residues Involved | Function |
|---|---|---|
Disulfide Bridge | Cysteine | Stabilizes tertiary/quaternary structure |
Diselenide Bridge | Selenocysteine | Redox-sensitive stabilization |
Summary Table: Levels of Protein Structure
Level | Description | Stabilizing Forces |
|---|---|---|
Primary | Amino acid sequence | Peptide bonds |
Secondary | α-helix, β-sheet, turns | Hydrogen bonds |
Tertiary | 3D folding of polypeptide | Hydrophobic interactions, disulfide bonds, electrostatic interactions, hydrogen bonds |
Quaternary | Association of multiple chains | Same as tertiary, plus inter-chain interactions |
Key Equations
Peptide Bond Formation:
Ramachandran Angles:
Additional info:
Some context and details have been inferred to ensure completeness and clarity, such as typical phi/psi angles and the role of selenocysteine in protein structure.
