 Back
BackFundamentals of Protein Structure: Post-Translational Modifications (PTMs)
Study Guide - Smart Notes

Fundamentals of Protein Structure: Post-Translational Modification
Introduction to Post-Translational Modifications (PTMs)
Post-translational modifications (PTMs) are chemical changes that occur to proteins after their synthesis (translation) in the cell. These modifications are essential for regulating protein function, localization, stability, and interactions, and they contribute significantly to the complexity of the proteome.
Definition: PTMs are covalent modifications of proteins, typically mediated by enzymes, that alter protein properties and functions.
Proteome Complexity: PTMs, along with alternative splicing and RNA editing, greatly expand the diversity of proteins derived from a limited number of genes.
Key Functions: PTMs regulate protein activity, localization, interactions, and degradation.
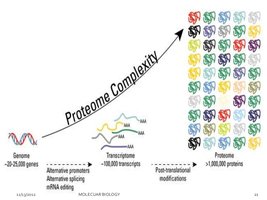
Types of Post-Translational Modifications
There are over 200 types of PTMs, but the most common and biologically significant include glycosylation, phosphorylation, ubiquitination, lipidation (GPI anchor), acetylation, and methylation.
Enzymes Involved: Kinases, phosphatases, transferases, ligases, and proteases.
Reversibility: Many PTMs are reversible, allowing dynamic regulation of protein function.
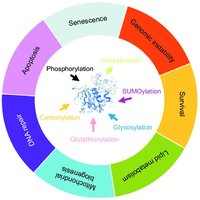
Glycosylation
Overview of Glycosylation
Glycosylation is the enzymatic addition of carbohydrate chains (glycans) to proteins, typically at specific amino acid residues. It is one of the most complex and widespread PTMs, occurring in various cellular compartments and affecting nearly half of plasma proteins.
Key Enzyme: Glycosyltransferases catalyze the addition of sugars.
Locations: Endoplasmic reticulum (ER), Golgi apparatus, cytosol, and membranes.
Biological Roles: Protein folding, cell adhesion, molecular trafficking, receptor activation, solubility, signal transduction, degradation, and secretion.
Disease Relevance: Defects in glycosylation are linked to cancer, diabetes, HIV, Alzheimer’s, and atherosclerosis.
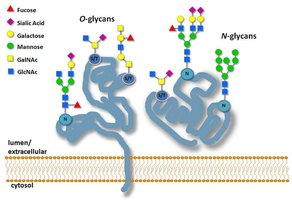
Types of Glycosylation
N-linked Glycosylation: Attachment of glycans to the nitrogen atom of asparagine (Asn) side chains. Consensus sequence: Asn-X-Ser/Thr (X ≠ Pro).
O-linked Glycosylation: Attachment of glycans to the oxygen atom of serine (Ser) or threonine (Thr) side chains. No strict consensus sequence.
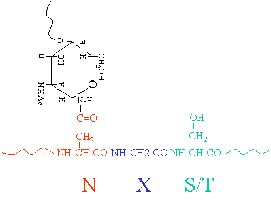
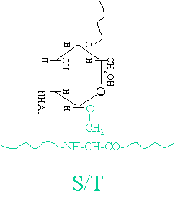
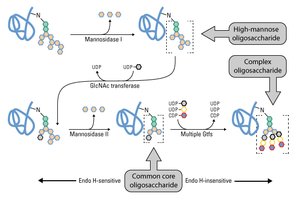
Glycosylation Mechanism and Maturation
Glycan precursors are assembled and then transferred to the polypeptide in the ER. Maturation occurs in the Golgi apparatus, where glycan structures are modified by removal and addition of sugar residues.
Quality Control: Glycosylation in the ER monitors protein folding, ensuring only properly folded proteins proceed to the Golgi.
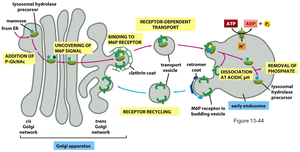
Cellular Trafficking: Glycosylation motifs (e.g., mannose-6-phosphate) act as signals for protein localization, such as targeting lysosomal enzymes.
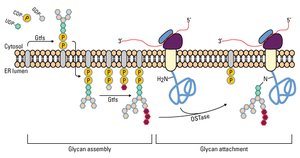
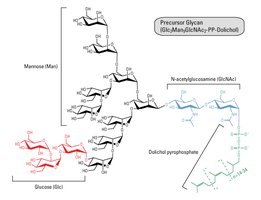
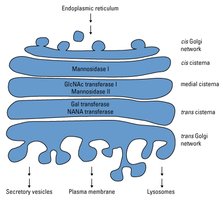
Glycosylation in Lysosomal Enzyme Sorting
Lysosomal proteins are tagged with mannose-6-phosphate in the Golgi, allowing recognition and transport to the lysosome. 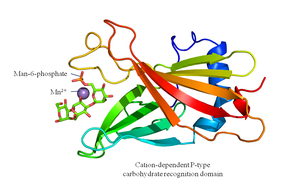
Blood Group Determination
Blood group antigens are determined by specific glycosylation patterns on red blood cell surfaces. 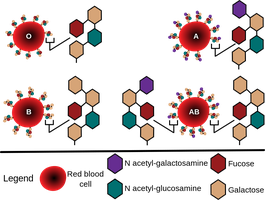
Phosphorylation
Overview of Phosphorylation
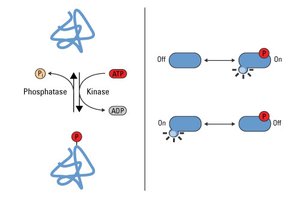
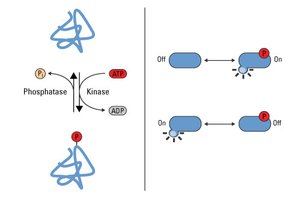
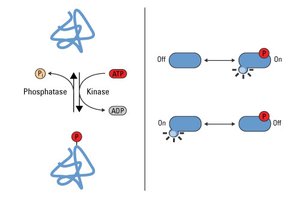
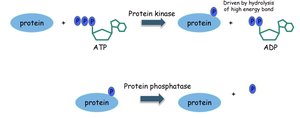
Phosphorylation is the addition of a phosphate group (PO43-) to an amino acid residue, typically serine, threonine, or tyrosine, catalyzed by kinases. It is a reversible modification, with phosphatases removing phosphate groups.
Key Enzymes: Kinases (add phosphate), phosphatases (remove phosphate).
Biological Roles: Regulation of enzyme activity, protein-protein interactions, subcellular localization, degradation, and signal transduction.
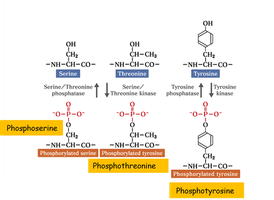
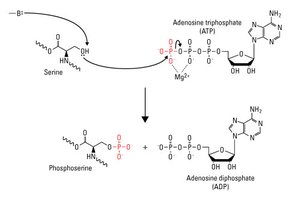
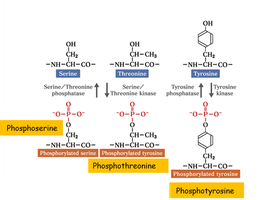
Phosphorylation – Amino Acids
Phosphorylation alters the properties of serine, threonine, and tyrosine side chains by adding a strong negative charge and bulk, which can affect protein structure and function.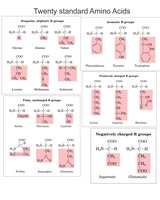 
Functional Consequences of Phosphorylation
Activation/Inhibition: Alters enzyme activity (e.g., glycogen phosphorylase activation).
Protein-Protein Interactions: Creates docking sites for signaling molecules (e.g., SH2 domains).
Subcellular Localization: Directs proteins to specific compartments.
Degradation: Can mark proteins for degradation via the ubiquitin-proteasome pathway.
Phosphorylation Cascades and Kinome
The human kinome consists of hundreds of kinases, enabling highly specific and complex regulation of cellular processes through phosphorylation cascades. 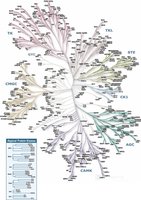
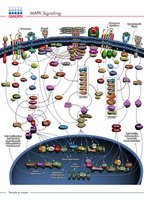
MAPK Signaling Pathway
Mitogen-activated protein kinase (MAPK) signaling is a key phosphorylation cascade involved in cell growth, differentiation, and stress responses. 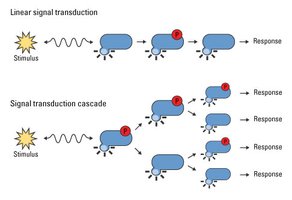
Ubiquitylation
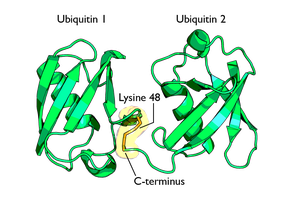
Overview of Ubiquitylation
Ubiquitylation is the covalent attachment of ubiquitin, a small regulatory protein, to lysine residues on target proteins. It is a reversible PTM, primarily involved in protein degradation via the ubiquitin-proteasome pathway.
Key Enzymes: E1 activating enzyme, E2 conjugating enzyme, E3 ligase, deubiquitinases.
Biological Roles: Protein degradation, stem cell regulation, transcription, DNA repair, replication, trafficking, virus budding, signal transduction.
Disease Relevance: Dysfunction leads to cancer, metabolic syndromes, inflammatory disorders, diabetes, neurodegenerative diseases.
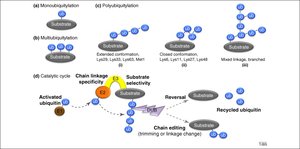
Types of Ubiquitylation
Monoubiquitylation: Single ubiquitin attached to a substrate.
Multiubiquitylation: Multiple ubiquitins attached at different sites.
Polyubiquitylation: Chains of ubiquitin attached to a single site, with various linkage types (extended, closed, forked).
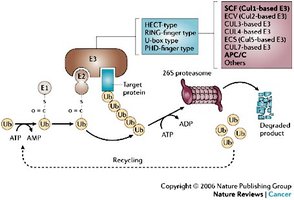
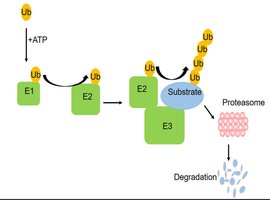
Mechanism of Ubiquitylation
Ubiquitin is activated by E1, transferred to E2, and then attached to the substrate by E3 ligase. E3 confers substrate specificity, while deubiquitinases can remove or edit ubiquitin chains. 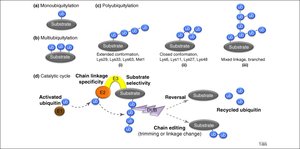
Protein Degradation via Ubiquitin-Proteasome Pathway
Ubiquitylated proteins are targeted to the proteasome for degradation, recycling ubiquitin and regulating protein levels.
Lipidation (GPI Anchor)
Overview of GPI Anchors
Glycosyl-phosphatidylinositol (GPI) anchors are glycolipids attached to the C-terminus of proteins, embedding them in the membrane. This modification affects protein localization and function.
Acetylation
Overview of Acetylation
Acetylation is the addition of an acetyl group (COCH3) to lysine side chains, catalyzed by lysine acetyltransferases (KAT) and histone acetyltransferases (HAT), using acetyl-CoA as a cofactor. Deacetylases (HDACs) remove acetyl groups.
Biological Roles: Chromatin stability, protein-protein interaction, cell cycle control, metabolism, nuclear transport, actin nucleation.
Methylation
Overview of Methylation
Methylation is the addition of a methyl group (-CH3) to amino acid residues, catalyzed by methyltransferases. It is a reversible PTM, often occurring on lysine and arginine residues, especially in histone proteins.
Biological Roles: Gene regulation, protein stability, signaling, protein-protein interactions.
Gene Expression: Histone methylation controls chromatin structure and transcription.
Protein Stability: Methylation can protect proteins from degradation.
Signal Transduction: Modulates signaling pathways.
PTMs Together: Coordinated Signaling
Integration of Multiple PTMs
Multiple PTMs can occur on a single protein, coordinating complex signaling pathways and cellular responses. For example, phosphorylation, ubiquitylation, and proteasomal activation are integrated in apoptosis and NF-κB signaling.
Summary
PTMs are essential for protein function, regulation, and cellular signaling.
Major types include glycosylation, phosphorylation, ubiquitylation, lipidation, acetylation, and methylation.
PTMs contribute to proteome complexity and are critical in health and disease.
