 Back
BackHow Enzymes Work: Mechanisms, Structure, and Evolution
Study Guide - Smart Notes

Section 6.1: What Is an Enzyme?
Enzymes as Biological Catalysts
Enzymes are specialized biological catalysts that accelerate chemical reactions in living organisms. Unlike non-biological catalysts, enzymes exhibit remarkable specificity and efficiency under mild physiological conditions.
Definition: An enzyme is a protein (or, rarely, an RNA molecule) that catalyzes a specific biochemical reaction.
Specificity: Enzymes are highly specific for their substrates and products due to the unique arrangement of functional groups in their active sites.
Classification: Enzymes are classified into seven major groups based on the type of reaction they catalyze: oxidoreductases, transferases, hydrolases, lyases, isomerases, ligases, and translocases.
Regulation: Enzyme activity is tightly regulated to meet the needs of the organism and respond to environmental changes.
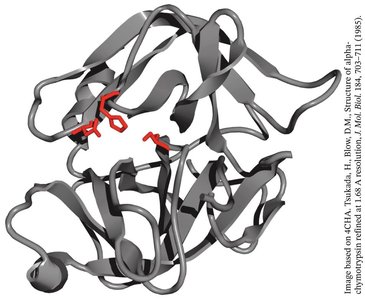
Enzyme Reaction Rates and Specificity
Enzymes dramatically increase reaction rates compared to uncatalyzed reactions, often by factors of 108 to 1012. Their specificity is illustrated by enzymes like chymotrypsin, which preferentially cleaves peptide bonds after phenolic amino acids.
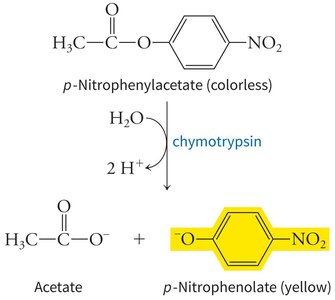
Example: Chymotrypsin catalyzes the hydrolysis of p-nitrophenylacetate to produce acetate and p-nitrophenolate (yellow), demonstrating substrate specificity and measurable product formation.
Enzyme Classification Table
Class of Enzyme | Type of Reaction Catalyzed |
|---|---|
Oxidoreductases | Oxidation–reduction reactions |
Transferases | Transfer of functional groups |
Hydrolases | Hydrolysis reactions |
Lyases | Group elimination to form double bonds |
Isomerases | Isomerization reactions |
Ligases | Bond formation coupled with ATP hydrolysis |
Translocases | Solute transport through membranes |
Section 6.2: Chemical Catalytic Mechanisms
Activation Energy and Reaction Rates
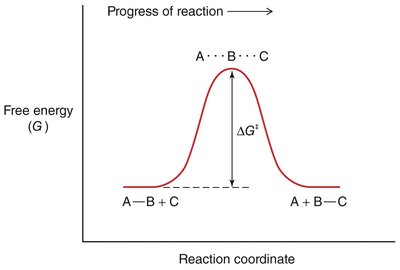
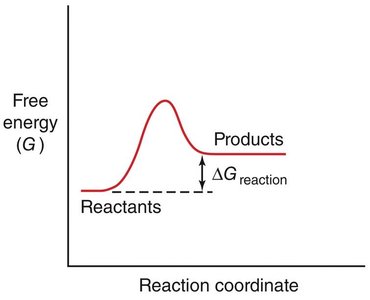
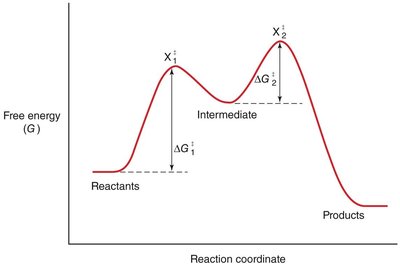
The rate of a chemical reaction depends on the activation energy (ΔG‡), which is the energy barrier that must be overcome for reactants to form products.
Transition State: The highest energy point along the reaction coordinate, where reactants are converted to products.
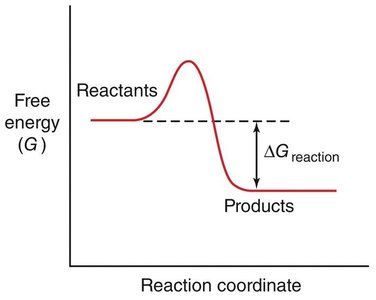
Spontaneity: The sign of ΔG determines whether a reaction is spontaneous (ΔG < 0) or non-spontaneous (ΔG > 0).
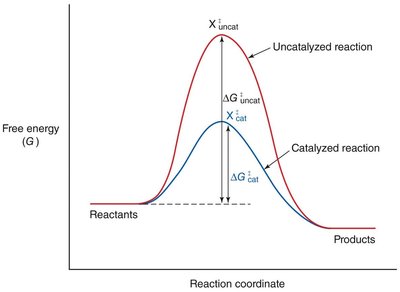
Enzyme Catalysis: Lowering Activation Energy
Enzymes accelerate reactions by lowering the activation energy, making it easier for reactants to reach the transition state.
Equation:
Effect: More reactant molecules achieve the transition state per unit time, increasing the reaction rate.
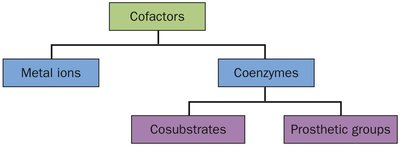
Cofactors in Enzyme Catalysis
Some enzymes require non-protein cofactors to assist in catalysis. Cofactors can be metal ions or organic molecules (coenzymes).
Metal ions: Participate in redox reactions or stabilize charged intermediates.
Coenzymes: Often derived from vitamins, may act as cosubstrates or prosthetic groups.
Types of Chemical Catalytic Mechanisms
Enzymes use three main mechanisms to catalyze reactions:
Acid-base catalysis: Proton transfer between enzyme and substrate.
Covalent catalysis: Formation of a transient covalent bond between enzyme and substrate.
Metal ion catalysis: Metal ions mediate redox reactions or stabilize charged groups.
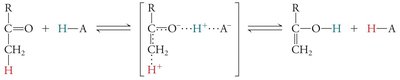
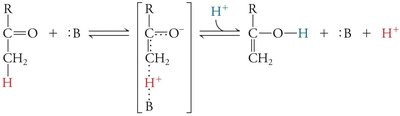
Covalent Catalysis and Reaction Intermediates
In covalent catalysis, a nucleophilic group in the enzyme forms a covalent intermediate with the substrate, resulting in a two-step reaction with two energy barriers.
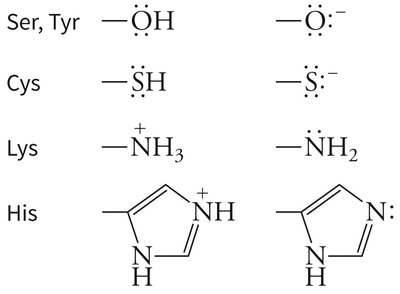
Example: Serine, cysteine, lysine, and histidine side chains can act as nucleophiles in covalent catalysis.
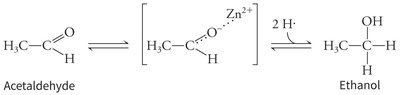
Metal Ion Catalysis Example
Example: Alcohol dehydrogenase uses a zinc ion to stabilize the transition state during the conversion of acetaldehyde to ethanol.
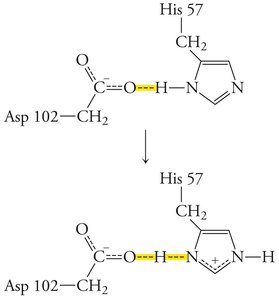
The Catalytic Triad of Chymotrypsin
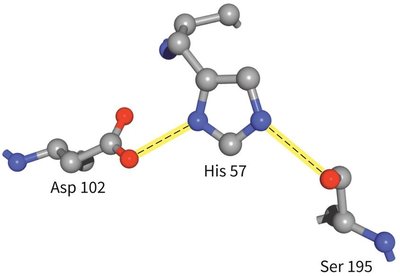
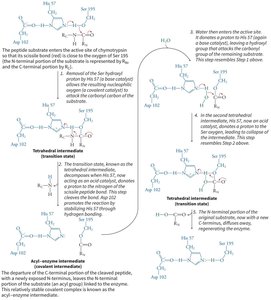
Chymotrypsin uses a hydrogen-bonded network of Asp 102, His 57, and Ser 195 to facilitate peptide bond hydrolysis.
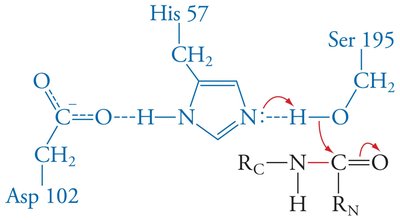
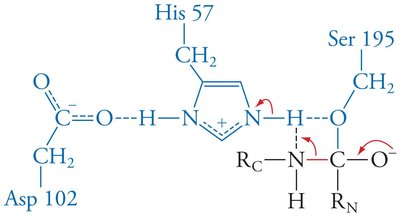
Mechanism: His 57 acts as a base, abstracting a proton from Ser 195, enabling nucleophilic attack on the substrate. Asp 102 stabilizes the positively charged His 57.
Section 6.3: Unique Properties of Enzyme Catalysts
Active Site Structure and Catalysis
The structure of the enzyme active site is crucial for catalysis, providing specificity and facilitating transition state stabilization.
Lock-and-key model: Proposes that the substrate fits precisely into the active site, but does not account for enzyme flexibility or product release.
Induced fit model: Suggests that enzymes undergo conformational changes upon substrate binding, optimizing interactions for catalysis.
Transition State Stabilization
Enzymes stabilize the transition state, often through features like the oxyanion hole in serine proteases, which forms hydrogen bonds with the substrate's anionic oxygen.
Low-barrier hydrogen bonds: Strong hydrogen bonds that further stabilize the transition state.
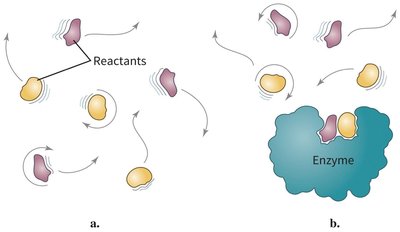
Proximity and Orientation Effects
Enzymes increase reaction rates by bringing reactants into close proximity and correct orientation, reducing the entropy barrier.
Example: Reactants in solution must collide with the correct orientation, but enzyme binding restricts their motion and aligns them for reaction.
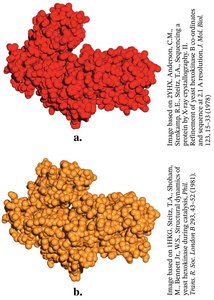
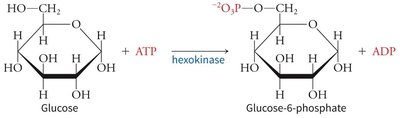
Induced Fit Model Example: Hexokinase
Hexokinase undergoes a conformational change upon glucose binding, enclosing the substrate and excluding water, which enhances catalysis.
Reaction:
Section 6.4: Chymotrypsin in Context
Enzyme Evolution: Divergent and Convergent
Enzymes can evolve similar mechanisms through divergent or convergent evolution.
Divergent evolution: Chymotrypsin, trypsin, and elastase share a common ancestor and retain similar structures and catalytic mechanisms.
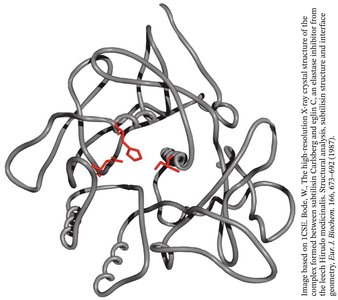
Convergent evolution: Subtilisin, a bacterial serine protease, evolved a similar catalytic triad and oxyanion hole independently of chymotrypsin.
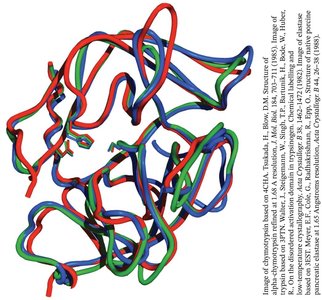
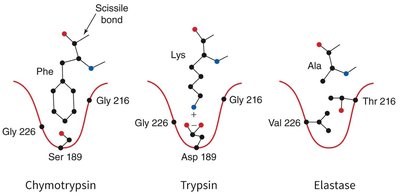
Specificity Pockets of Serine Proteases
The substrate specificity of serine proteases is determined by the chemical character of the specificity pocket at the active site.
Chymotrypsin: Prefers large hydrophobic side chains (e.g., Phe).
Trypsin: Prefers positively charged side chains (e.g., Lys, Arg).
Elastase: Prefers small side chains (e.g., Ala, Gly, Val).
Zymogen Activation and Protease Inhibition
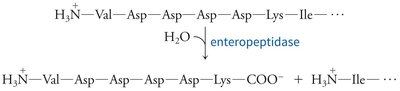
Proteases are often synthesized as inactive precursors (zymogens) and activated by proteolysis. Protease inhibitors regulate activity by blocking the active site.
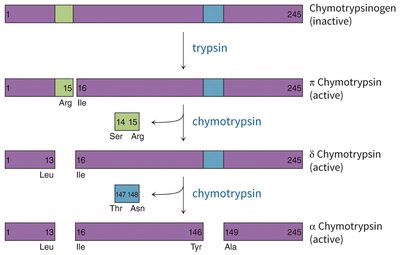
Zymogen activation: Chymotrypsinogen is activated by trypsin-mediated cleavage, followed by further processing to yield active chymotrypsin species.
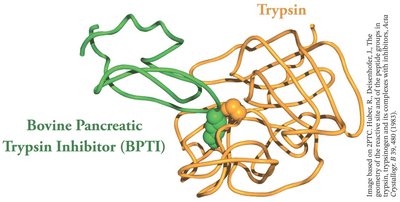
Protease inhibitors: Small proteins that bind to and block the active site of proteases, preventing unwanted proteolysis.
Optional: Section 6.5 Clinical Connection – Blood Coagulation
Blood Coagulation as a Protease Cascade
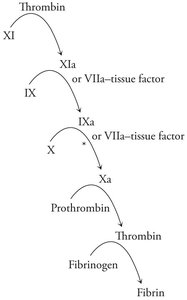
Blood coagulation is a cascade of proteolytic activations, resulting in the formation of a fibrin clot to stop bleeding.
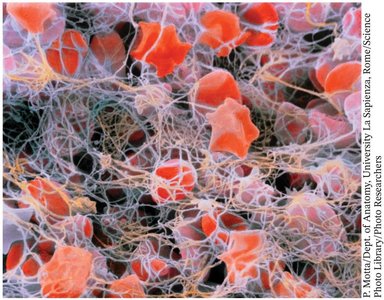
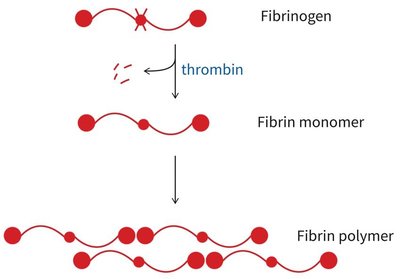
Fibrin: Polymerizes to form a mesh that traps blood cells at the site of injury.
Coagulation cascade: Sequential activation of proteases, starting with tissue factor exposure and culminating in thrombin-mediated conversion of fibrinogen to fibrin.
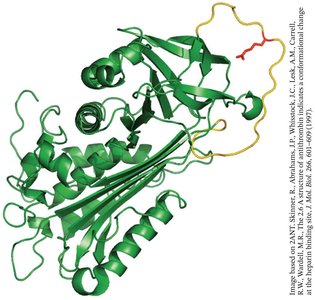
Antithrombin: Inhibits multiple coagulation proteases, preventing excessive clot formation.
 Additional info: Blood coagulation is not required for assessments but illustrates the physiological importance of protease regulation and cascade amplification in biochemistry.
Additional info: Blood coagulation is not required for assessments but illustrates the physiological importance of protease regulation and cascade amplification in biochemistry.
