 Back
BackProtein Function and Evolution: Immunoglobulins, Oxygen Transport, and Allosteric Regulation
Study Guide - Smart Notes

Protein Function and Evolution
Transcription and Translation: From Gene to Protein
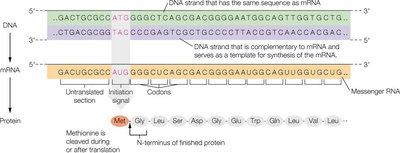
Proteins are synthesized through the processes of transcription and translation, where genetic information encoded in DNA is transcribed into messenger RNA (mRNA) and then translated into a polypeptide chain. This process is fundamental to the expression of genetic information and the production of functional proteins.
Transcription: DNA is used as a template to synthesize mRNA, which carries the genetic code from the nucleus to the ribosome.
Translation: The mRNA sequence is read in codons (triplets of nucleotides), each specifying an amino acid, to assemble the polypeptide chain.
Initiation Signal: Translation begins at a specific start codon (AUG), which codes for methionine.
Post-Translational Processing: After translation, the polypeptide may undergo folding, formation of disulfide bonds, and enzymatic modifications to become a functional protein.
Additional info: The genetic code is universal and conserved across species, allowing for the transfer of genetic information between organisms.
Post-Translational Processing of Proteins
After synthesis, many proteins undergo post-translational modifications that are essential for their final structure and function. These modifications include folding, formation of disulfide bonds, proteolytic cleavage, and chemical modifications of amino acid residues.
Folding: Newly synthesized polypeptides fold into their native three-dimensional structures, often assisted by chaperone proteins.
Disulfide Bonds: Covalent bonds between cysteine residues stabilize protein structure, especially in secreted proteins like insulin.
Proteolytic Cleavage: Specific enzymes cleave peptide bonds to activate or mature proteins (e.g., conversion of proinsulin to insulin).
Enzymatic Modifications: Addition of functional groups (phosphorylation, glycosylation) can regulate protein activity, localization, or stability.
Additional info: Post-translational modifications are critical for protein diversity and regulation in cellular processes.
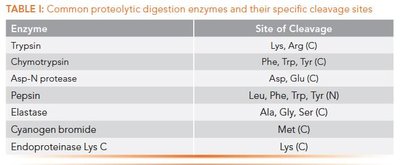
Common Proteolytic Digestion Enzymes
Enzyme | Site of Cleavage |
|---|---|
Trypsin | Lys, Arg (C) |
Chymotrypsin | Phe, Trp, Tyr (C) |
Asp-N protease | Asp, Glu (N) |
Pepsin | Leu, Phe, Trp, Tyr (N) |
Elastase | Ala, Gly, Ser (C) |
Cyanogen bromide | Met (C) |
Endoproteinase Lys C | Lys (C) |
Insulin Processing: A Model for Protein Maturation
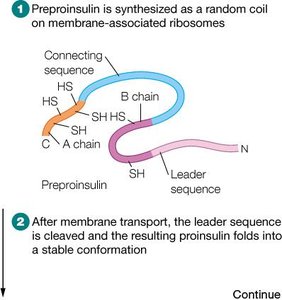
Insulin is synthesized as a single polypeptide precursor (preproinsulin) that undergoes sequential processing to yield the active hormone. This process illustrates the importance of post-translational modifications in protein maturation.
Preproinsulin: Contains a signal peptide that directs the nascent chain to the endoplasmic reticulum.
Proinsulin: After removal of the signal peptide, the chain folds and forms disulfide bonds.
Mature Insulin: Proteolytic cleavage removes the connecting peptide (C-peptide), resulting in two chains (A and B) linked by disulfide bonds.
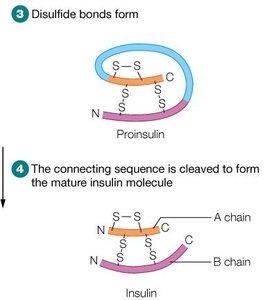
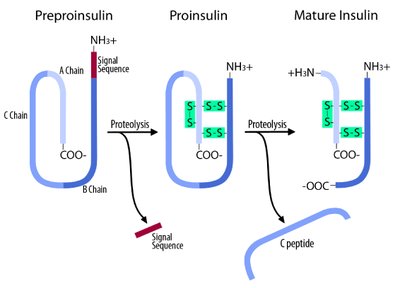
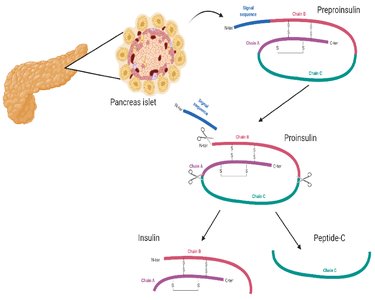
Example: Proinsulin is stored in the pancreas and rapidly converted to insulin when needed, preventing toxicity from excess active hormone.
Protein Sequence Alignment and Evolution
Sequence Identity vs. Sequence Similarity
Comparing protein sequences helps reveal evolutionary relationships and functional similarities. Sequence identity refers to exact matches of amino acids, while sequence similarity considers conservative substitutions with similar chemical properties.
Sequence Alignment: Tools like BLAST compare protein sequences to identify regions of similarity and infer evolutionary relationships.
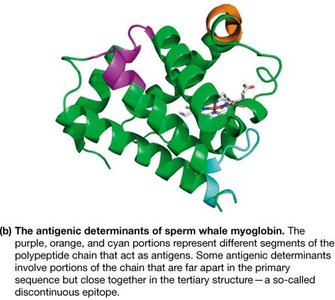
Globins: Myoglobin and hemoglobin share sequence and structural similarities, reflecting their common evolutionary origin.
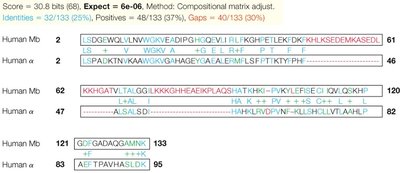

Additional info: Sequence similarity can indicate conserved functional domains even when overall identity is low.
Antibody Structure and Function
Immunoglobulins: Structure and Classes
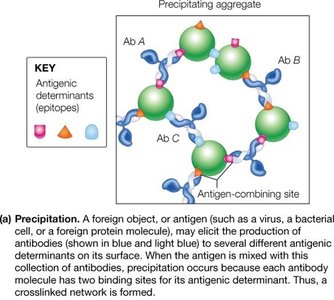
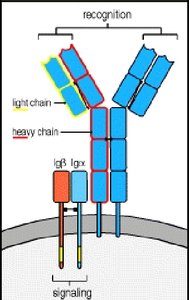
Antibodies, or immunoglobulins, are proteins produced by B lymphocytes that specifically recognize and bind antigens. They play a central role in the adaptive immune response.
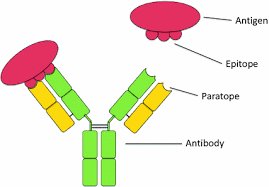
Structure: Each antibody consists of two identical heavy chains and two identical light chains, forming a Y-shaped molecule stabilized by disulfide bonds.
Variable Domains: The antigen-binding sites are located at the tips of the Y, formed by variable regions of both heavy and light chains.
Constant Domains: The stem of the Y (Fc region) mediates effector functions and determines the antibody class.
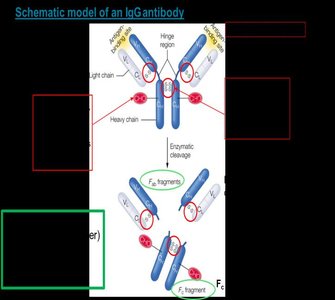
Classes of Immunoglobulins:
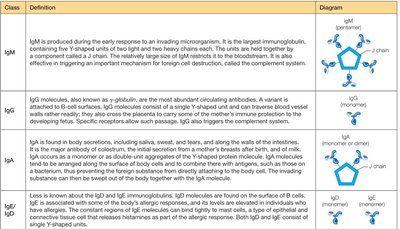
Class | Definition | Diagram |
|---|---|---|
IgM | Produced early in response; pentameric; triggers complement system | Pentamer |
IgG | Most abundant; monomeric; crosses placenta; triggers complement | Monomer |
IgA | Found in secretions; monomer/dimer; protects mucosal surfaces | Monomer/Dimer |
IgD | Surface of B cells; monomer/dimer | Monomer/Dimer |
IgE | Allergic responses; binds mast cells; monomer | Monomer |
Antigen Recognition and Immune Response
Antibodies recognize specific regions (epitopes) on antigens. The adaptive immune response is divided into humoral (antibody-mediated) and cellular (T-cell-mediated) branches.
Epitope: The specific part of an antigen recognized by an antibody.
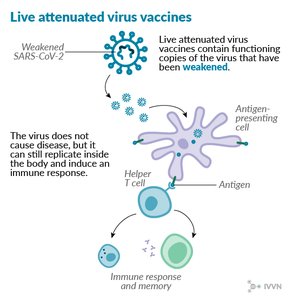
Humoral Response: B cells produce antibodies that bind and neutralize pathogens.
Cellular Response: T cells recognize and destroy infected or abnormal cells.
Memory: After initial exposure, memory cells enable a rapid and robust response upon re-exposure to the same antigen.
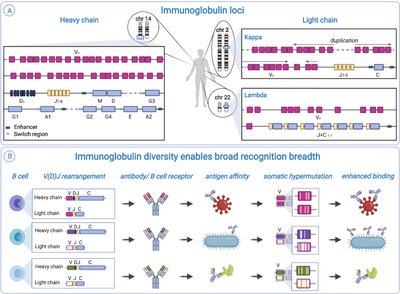
Development and Diversity of Antibodies
The immune system generates a vast diversity of antibodies through gene rearrangement and somatic hypermutation, allowing recognition of a wide array of antigens.
VDJ Recombination: Variable (V), Diversity (D), and Joining (J) gene segments recombine to create unique antibody genes in B cells.
Somatic Hypermutation: Further diversification occurs in activated B cells, enhancing antigen binding.
Clonal Selection: B cells with high-affinity receptors proliferate and differentiate into plasma cells (antibody-secreting) and memory cells.
Oxygen Transport Proteins: Myoglobin and Hemoglobin
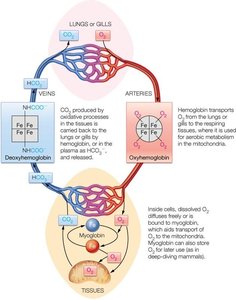
Role of Globins in Oxygen Transport and Storage
Myoglobin and hemoglobin are globular proteins essential for oxygen transport and storage in animals. Myoglobin is a monomer found in muscle, while hemoglobin is a tetramer found in red blood cells.
Myoglobin: Binds oxygen with high affinity; serves as an oxygen reservoir in muscle tissue.
Hemoglobin: Transports oxygen from lungs to tissues; exhibits cooperative binding, allowing efficient oxygen delivery and release.
Heme Group: Both proteins contain a heme prosthetic group with an Fe(II) ion that binds oxygen reversibly.


Oxygen Binding and Allosteric Regulation
Hemoglobin displays allosteric regulation, where binding of oxygen to one subunit increases the affinity of the remaining subunits (cooperative binding). This is reflected in the sigmoidal oxygen-binding curve of hemoglobin, compared to the hyperbolic curve of myoglobin.
Cooperative Binding: Facilitates efficient oxygen loading in the lungs and unloading in tissues.
T-State (Tense): Low-affinity, deoxygenated conformation.
R-State (Relaxed): High-affinity, oxygenated conformation.
Allosteric Effectors: Molecules such as H+, CO2, Cl-, and 2,3-BPG modulate hemoglobin's oxygen affinity, adapting oxygen delivery to physiological needs.
The Bohr Effect and Allosteric Modulation
The Bohr effect describes how decreases in pH (increased H+ concentration) and increases in CO2 concentration reduce hemoglobin's affinity for oxygen, promoting oxygen release in metabolically active tissues.
Bohr Effect Equation:
2,3-Bisphosphoglycerate (2,3-BPG): Binds to deoxyhemoglobin, stabilizing the T-state and facilitating oxygen release, especially under conditions of low oxygen availability (e.g., high altitude).
Fetal Hemoglobin (HbF): Has higher oxygen affinity due to reduced binding of 2,3-BPG, allowing efficient oxygen transfer from mother to fetus.
Additional info: Allosteric regulation of hemoglobin is a classic example of protein structure-function relationship and physiological adaptation.
Summary Table: Factors Affecting Hemoglobin-Oxygen Dissociation Curve
Factor | Effect on Curve | Physiological Significance |
|---|---|---|
pH (Bohr Effect) | Right shift (lower affinity) | Promotes O2 release in tissues |
CO2 | Right shift (lower affinity) | Promotes O2 release in tissues |
2,3-BPG | Right shift (lower affinity) | Facilitates O2 unloading |
Temperature | Right shift (lower affinity) | Increased O2 delivery during fever/exercise |
Fetal Hemoglobin | Left shift (higher affinity) | Enhances O2 uptake from maternal blood |
