 Back
BackStructure and Properties of DNA: Biochemistry Study Notes
Study Guide - Smart Notes

Structure and Properties of DNA
Central Dogma of Molecular Biology
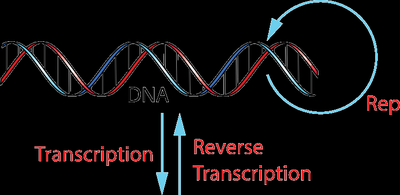
The central dogma describes the flow of genetic information within a biological system. It outlines the processes by which the information in DNA is transcribed into RNA and then translated into protein, with some exceptions such as reverse transcription in retroviruses.
Replication: DNA makes a copy of itself during cell division.
Transcription: DNA is used as a template to synthesize messenger RNA (mRNA).
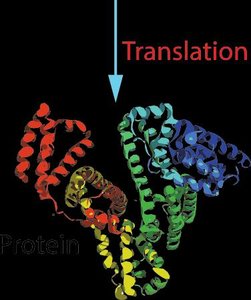
Translation: mRNA is decoded by ribosomes to produce proteins.
Reverse Transcription: In some viruses, RNA can be reverse-transcribed to DNA.
Primary Structure of DNA
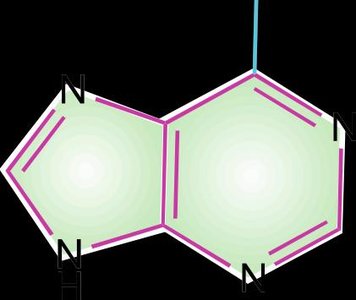
The primary structure of DNA refers to the linear sequence of nucleotides, each consisting of a nitrogenous base, a deoxyribose sugar, and a phosphate group. The sequence encodes genetic information.
Nucleoside: A molecule consisting of a nitrogenous base and a sugar.
Nucleotide: A nucleoside with one or more phosphate groups attached.
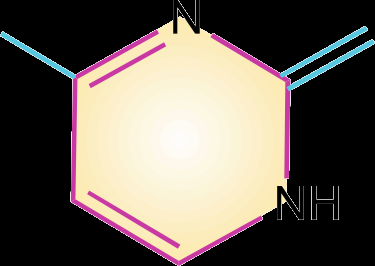
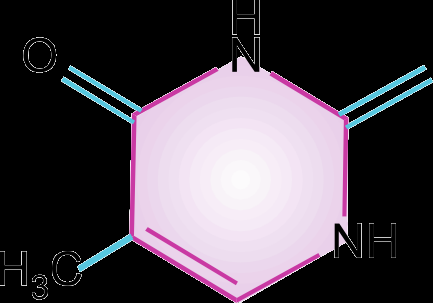
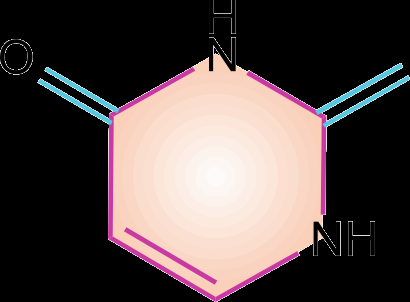
Pyrimidines: Cytosine (C), Thymine (T, in DNA), and Uracil (U, in RNA).
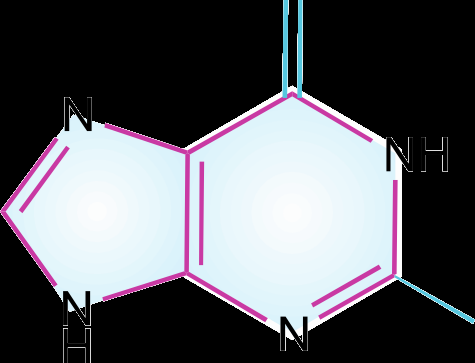
Purines: Adenine (A) and Guanine (G).
Phosphodiester bond: Links the 3'-carbon of one sugar to the 5'-carbon of the next.
Secondary Structure of DNA
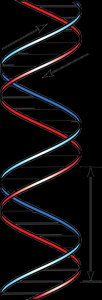
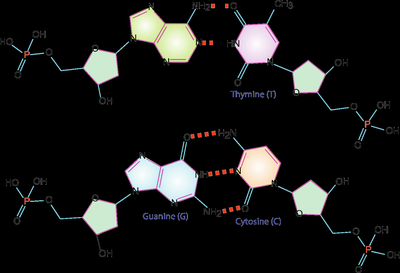
The secondary structure of DNA is the double helix, as described by Watson and Crick. Two antiparallel strands are held together by hydrogen bonds between complementary bases (A-T and G-C).
Watson-Crick base pairs: Adenine pairs with Thymine (A-T), Guanine pairs with Cytosine (G-C).
Major and minor grooves: Allow proteins to interact with specific base sequences.
Helical parameters: 10 base pairs per turn, 2.0 nm diameter, 0.34 nm between bases.
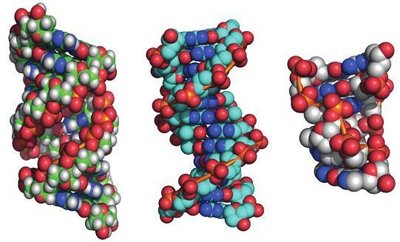
Forms of DNA Helices
DNA can adopt several conformations, with the B-form being the most common under physiological conditions. The A-form and Z-form are alternative structures that occur under specific conditions.
B-form: Right-handed, 10.5 bases per turn, predominant in cells.
A-form: Right-handed, more compact, 11 bases per turn, forms in dehydrated conditions.
Z-form: Left-handed, zigzag backbone, 12 bases per turn, forms in high salt or with certain sequences.
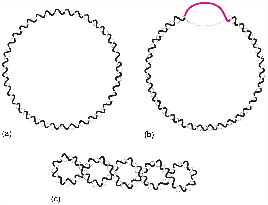
Tertiary Structure of DNA: Supercoiling
Supercoiling refers to the overwinding or underwinding of DNA, which is important for DNA packaging and regulation. Circular DNA molecules, such as plasmids, often exhibit supercoiling.
Negative supercoiling: Underwound DNA, facilitates strand separation.
Positive supercoiling: Overwound DNA, occurs ahead of replication forks.
Topoisomerases: Enzymes that regulate supercoiling by cutting and rejoining DNA strands.
Denaturation and Renaturation of DNA
Denaturation is the process by which double-stranded DNA unwinds and separates into single strands due to the disruption of hydrogen bonds, typically by heat, pH, or chemicals. Renaturation is the reformation of the double helix from single strands.
Melting temperature (Tm): The temperature at which half of the DNA is denatured; increases with GC content.
Hyperchromic shift: Increase in absorbance at 260 nm upon denaturation.
Renaturation: Depends on sequence complexity and concentration.
DNA Methylation and Genomic Imprinting
Methylation of DNA bases, especially cytosine and guanine, plays a role in gene regulation, protection from restriction enzymes, and genomic imprinting. Abnormal methylation patterns can lead to diseases.
Dam methylase: Methylates adenine in GATC sequences.
Genomic imprinting: Parent-specific methylation patterns; errors can cause syndromes such as Prader-Willi and Angelman.
Base Analogues and Antiviral Drugs
Base analogues are structurally similar to normal DNA bases and can be incorporated into DNA, often disrupting replication or function. Some are used as antiviral or anticancer drugs.
Acyclovir: Used for herpes virus infections.
AZT (3'-deoxy-3'-azidothymidine): Used in HIV treatment.
5-Fluorouracil: Thymine analogue, inhibits thymidylate synthetase.
Chromosomes, Chromatin, and Nucleosomes
In eukaryotes, DNA is packaged with histone proteins to form chromatin. The basic unit of chromatin is the nucleosome, which consists of DNA wrapped around histone proteins.
Histones: Basic proteins (H1, H2A, H2B, H3, H4) that package and order DNA.
Nucleosome: Bead-like structure, 10 nm in diameter.
30 nm fiber: Higher-order chromatin structure, stabilized by H1 histone.
Repetitive DNA Sequences
Repetitive DNA sequences are classified based on their reassociation kinetics and copy number in the genome.
Highly repetitive: Tandem repeats, 10-15% of mammalian DNA.
Moderately repetitive: Interspersed repeats, 25-40% of DNA.
Single copy: Unique sequences, 50-60% of DNA.
Plasmids and Conjugation
Plasmids are small, circular DNA molecules found in bacteria. They can be transferred between cells via conjugation, a process important for genetic exchange and antibiotic resistance.
Conjugation: Transfer of plasmid DNA from donor to recipient bacterium through a conjugation tube.
Replication: Plasmid DNA replicates independently of chromosomal DNA.

Enzymatic Manipulation of DNA
Various enzymes are used to manipulate DNA in the laboratory and in cells.
DNases/RNases: Degrade DNA/RNA by breaking phosphodiester bonds.
Restriction enzymes: Cut DNA at specific sequences, producing sticky or blunt ends.
DNA ligases: Join DNA fragments covalently.
DNA polymerases: Synthesize DNA from a template.
Reverse transcriptase: Synthesizes DNA from an RNA template.
DNA Cloning and Polymerase Chain Reaction (PCR)
DNA cloning allows for the amplification and manipulation of specific DNA sequences. PCR is a technique used to amplify DNA exponentially using cycles of denaturation, annealing, and extension.
Cloning vectors: Plasmids, bacteriophages, artificial chromosomes.
PCR steps: Denaturation (95°C), annealing (55°C), extension (72°C), repeated for 30-40 cycles.
Applications: Gene mapping, sequencing, diagnostics.
Detection of Genetic Mutations: RFLP and Southern Blot
Restriction Fragment Length Polymorphism (RFLP) analysis and Southern blotting are used to detect specific mutations, such as the point mutation responsible for sickle cell anemia.
RFLP: Detects differences in DNA fragment lengths after restriction enzyme digestion.
Southern blot: Transfers DNA fragments to a membrane for hybridization with labeled probes.
Sickle cell anemia: Caused by a single nucleotide mutation in the β-globin gene, leading to abnormal hemoglobin and sickled red blood cells.
Additional info: These notes cover the structure, properties, and manipulation of DNA, which are foundational topics in biochemistry and molecular biology. The included images directly support the understanding of DNA structure, base pairing, and the central dogma.
