 Back
BackCell Motility: Microtubule- and Actin-Based Movement in Eukaryotic Cells
Study Guide - Smart Notes

Cell Motility: Overview
Cell motility encompasses the diverse mechanisms by which cells and their internal components move. This includes movement at the subcellular, cellular, and tissue levels, driven primarily by cytoskeletal elements and associated motor proteins.
Subcellular motility: Separation of chromosomes during cell division, intracellular transport of vesicles and organelles.
Cellular motility: Cilia/flagella-based movement, amoeboid movement, chemotaxis, and tumor metastasis.
Tissue-level motility: Muscle contraction, resulting from the coordinated action of many muscle cells.
Microtubule-Based Motility
Motor Proteins and Microtubules
Microtubule-based motility is powered by motor proteins that convert chemical energy from ATP hydrolysis into mechanical work, enabling the transport of cellular cargo along microtubule tracks.
Kinesins: Move toward the plus end of microtubules, typically transporting cargo away from the cell center.
Dyneins: Move toward the minus end of microtubules, often transporting cargo toward the cell center.
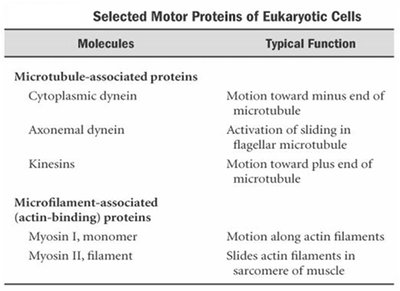
Kinesin Structure and Function
Kinesin-1 is a well-studied motor protein with two globular heads (motor domains), a coiled-coil stalk, and a light-chain tail that binds cargo. The heads contain microtubule-binding and ATPase sites, generating force through ATP hydrolysis.
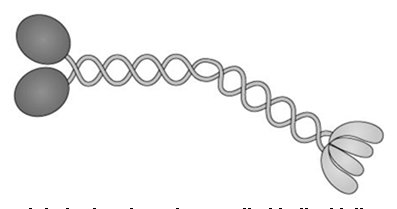
Kinesin Movement Mechanism
Kinesin moves along microtubules in a processive, stepwise manner, with each step corresponding to the hydrolysis of one ATP molecule and a displacement of approximately 8 nm. The movement is often described as a "walking" motion, where one head detaches and moves forward to bind a new tubulin subunit.
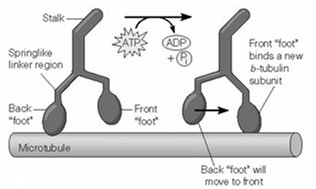
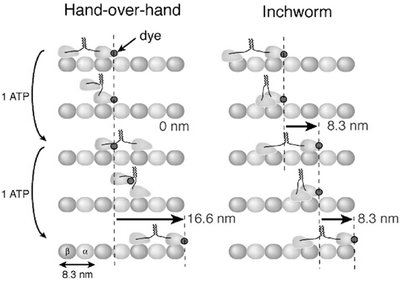
Hand-Over-Hand vs. Inchworm Models
Two models have been proposed for kinesin movement: the hand-over-hand and inchworm mechanisms. Experimental evidence supports the hand-over-hand model, where the two heads alternate in leading, resulting in 8-nm steps per ATP hydrolyzed.
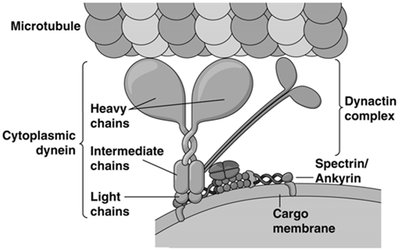
Dynein Structure and Function
Cytoplasmic dynein is a large, multi-subunit complex with heavy, intermediate, and light chains. The heavy chains bind to microtubules and generate movement toward the minus end, while the light chains interact with the dynactin complex, which links dynein to cargo membranes.
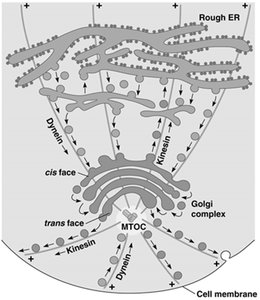
Microtubule-Based Vesicle Transport
Microtubules serve as tracks for the transport of vesicles and organelles within cells. Kinesin and dynein mediate anterograde and retrograde transport, respectively, ensuring the proper distribution of cellular components.
Cilia and Flagella: Structure and Function
General Features
Cilia and flagella are motile appendages of eukaryotic cells, composed of a core axoneme connected to a basal body. Cilia are typically short and numerous, while flagella are longer and fewer in number.
Axoneme Structure
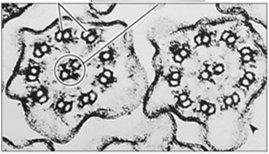
The axoneme is the core structure of cilia and flagella, characterized by a "9+2" arrangement of microtubules: nine outer doublets surrounding a central pair. Dynein arms attached to the A tubule of each doublet interact with the B tubule of the adjacent doublet, generating sliding forces that are converted into bending motions by nexin links and other cross-linking proteins.
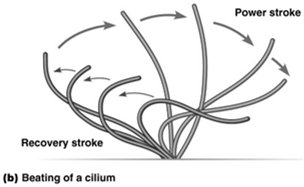
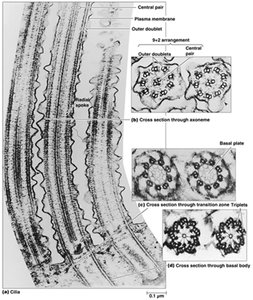
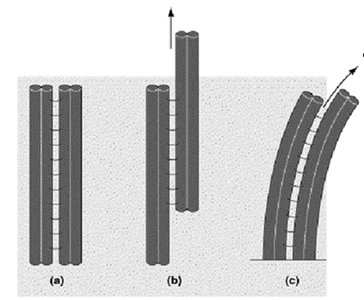
Mechanism of Ciliary and Flagellar Movement
Dynein arms "walk" along adjacent microtubule doublets, causing them to slide against each other. This sliding is constrained by cross-linking proteins, resulting in the bending of the entire axoneme and the characteristic beating motion of cilia and flagella.
Actin-Based Motility and Muscle Contraction
Myosin Structure and Function
Myosins are actin-based motor proteins with a head domain (containing actin-binding and ATPase sites), a neck region (binding light chains), and a variable tail domain. Myosin II forms bipolar filaments in muscle cells, enabling contraction.
Organization of Skeletal Muscle Tissue
Skeletal muscle is composed of bundles of myofibers (multinucleated muscle cells), which contain myofibrils made up of repeating sarcomeres—the fundamental contractile units. Sarcomeres are defined by Z lines and contain thick (myosin) and thin (actin) filaments.
Structure of the Sarcomere
Thick filaments: Composed of myosin II, bundled together by myomesin.
Thin filaments: Composed of actin, attached to Z-disks, and decorated by tropomyosin and troponin. Capping proteins (Cap Z and tropomodulin) regulate filament length.
Major Protein Components of Vertebrate Skeletal Muscle
The following table summarizes the major protein components and their functions in vertebrate skeletal muscle:
Protein | Function |
|---|---|
G-actin | Major component of thin filaments |
Myosin | Major component of thick filaments |
Tropomyosin | Binds along thin filaments |
Troponin | Calcium regulation of contraction |
Titin | Links thick filaments to Z line |
Nebulin | Stabilizes thin filaments |
Myomesin | Myosin-binding protein at M line |
α-actinin | Bundles actin filaments at Z line |
Ca2+ ATPase | Transports Ca2+ into SR for relaxation |
CapZ | Caps actin at Z line |
Tropomodulin | Maintains thin filament length |
Sliding Filament Model of Muscle Contraction
Muscle contraction occurs as thick and thin filaments slide past each other, increasing their overlap. The I band shortens, Z lines are pulled together, and the sarcomere contracts. The process is driven by the cyclic interaction of myosin heads with actin filaments, powered by ATP hydrolysis.
Each myosin head binds to actin, undergoes a power stroke, and releases upon ATP binding.
ATP hydrolysis "cocks" the myosin head for the next cycle.
Regulation of Muscle Contraction
In striated muscle, contraction is regulated by the troponin-tropomyosin complex. In the absence of Ca2+, tropomyosin blocks myosin binding sites on actin. When Ca2+ binds to troponin C, tropomyosin shifts, exposing binding sites and allowing contraction.
Ca2+ increase: Binds troponin C, enables contraction.
Ca2+ decrease: Troponin-tropomyosin blocks myosin, muscle relaxes.
Smooth Muscle and Non-Muscle Cell Motility
Smooth muscle contraction is regulated differently, involving calmodulin and myosin light chain kinase (MLCK), rather than troponin. Actin-based motility in non-muscle cells includes cell crawling via lamellipodia and filopodia, driven by actin polymerization and myosin activity.
Chemotaxis
Chemotaxis is the directed movement of cells in response to chemical gradients, involving cytoskeletal rearrangements and actin-based protrusions.
Summary Table: Motor Proteins and Their Functions
Molecules | Typical Function |
|---|---|
Cytoplasmic dynein | Motion toward minus end of microtubule |
Axonemal dynein | Activation of sliding in flagellar microtubule |
Kinesins | Motion toward plus end of microtubule |
Myosin I, monomer | Motion along actin filaments |
Myosin II, filament | Slides actin filaments in sarcomere of muscle |

Key Concepts and Review Questions
All motor proteins convert chemical energy (ATP) into motion.
All myosins are motor proteins that bind to and exert force on microfilaments (actin).
Calcium regulation of muscle contraction occurs via conformational changes in tropomyosin, mediated by troponin binding Ca2+.
