 Back
Back21: Cell Signaling Pathways and the Cytoskeleton in Eukaryotic Cells
Study Guide - Smart Notes

Cell Signaling Pathways
Overview of Intracellular Signaling Pathways
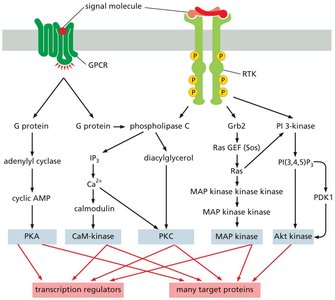
Cell signaling is essential for cells to perceive and respond to their microenvironment. Major classes of cell-surface receptors, including G protein-coupled receptors (GPCRs) and receptor tyrosine kinases (RTKs), initiate complex intracellular signaling cascades that regulate gene expression and cellular behavior.
GPCRs activate heterotrimeric G proteins, which in turn regulate enzymes such as adenylyl cyclase and phospholipase C, leading to the production of second messengers (e.g., cAMP, IP3, Ca2+).
RTKs activate signaling proteins through phosphorylation, including the MAP kinase and PI3-kinase pathways.
Both pathways converge on kinases that regulate transcription factors and other target proteins, ultimately altering gene expression and cell function.
Cytokine Receptors and the JAK-STAT Pathway
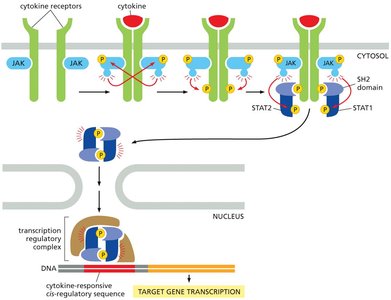
Cytokine receptors are critical for immune signaling and gene expression regulation. Unlike RTKs, cytokine receptors lack intrinsic enzymatic activity and rely on associated Janus kinases (JAKs) to transmit signals.
Cytokines are small proteins (e.g., interleukins, interferons, TNFs) that bind to cytokine receptors, triggering JAK activation.
Activated JAKs phosphorylate STAT proteins (Signal Transducers and Activators of Transcription), which dimerize and translocate to the nucleus to regulate gene transcription.
Signal termination is mediated by protein phosphatases.
Receptor Serine/Threonine Kinases and TGF-β Signaling
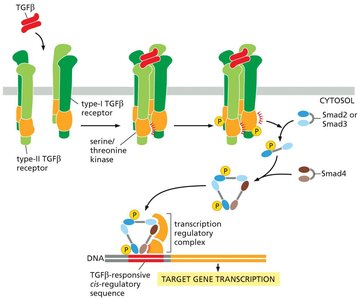
Receptor serine/threonine kinases, such as those in the TGF-β superfamily, provide a direct route for signal transduction to the nucleus. These receptors are activated by ligands like TGF-β, leading to phosphorylation of SMAD proteins.
TGF-β receptors phosphorylate receptor-regulated SMADs (R-SMADs), which then form complexes with co-SMADs and translocate to the nucleus.
These complexes regulate the transcription of TGF-β-responsive genes, influencing cell growth, differentiation, and apoptosis.
Evolution of Cell Communication in Plants and Animals
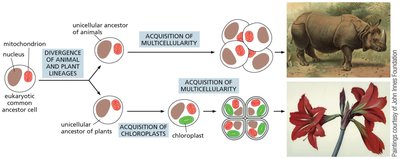
Multicellularity and cell communication evolved independently in plants and animals, diverging from a common unicellular eukaryotic ancestor over a billion years ago. Plant and animal cells utilize distinct sets of receptors and signaling mechanisms.
Plants predominantly use serine/threonine kinases (e.g., LRR receptor kinases) and have few GPCRs.
Plant cells do not use RTKs, steroid-hormone-type nuclear receptors, or cAMP signaling as animals do.
Plant Hormone Signaling Pathways
Ethylene Signaling
Ethylene is a gaseous plant hormone that regulates growth and stress responses. The ethylene signaling pathway involves receptors homologous to bacterial histidine kinases.
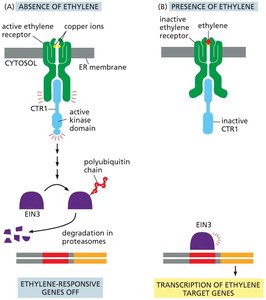
In the absence of ethylene, the receptor activates CTR1 kinase, leading to degradation of the transcription factor EIN3 and repression of ethylene-responsive genes.
When ethylene is present, the receptor is inactivated, CTR1 is turned off, EIN3 accumulates, and ethylene target genes are transcribed.
Auxin Signaling
Auxin is a key plant hormone that regulates cell elongation and gene expression. The auxin receptor is located in the nucleus and controls the degradation of transcriptional repressors.
In the absence of auxin, Aux/IAA proteins repress auxin response factors (ARFs), keeping target genes off.
Auxin binding leads to ubiquitination and degradation of Aux/IAA proteins, allowing ARFs to activate gene transcription.
Phytochrome Signaling
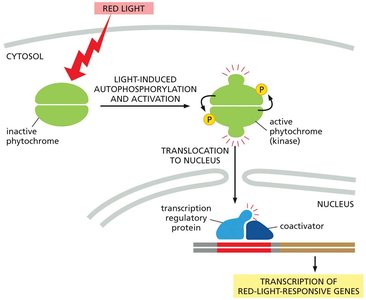
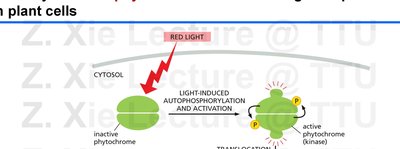
Phytochromes are red/far-red light photoreceptors that regulate plant development. They are dimeric, cytoplasmic serine/threonine kinases that translocate to the nucleus upon activation by red light.
Light-induced autophosphorylation activates phytochromes, which then move to the nucleus and regulate transcription of light-responsive genes.
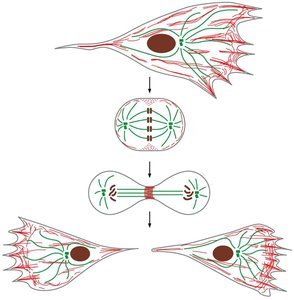
The Cytoskeleton
Introduction to the Cytoskeleton
The cytoskeleton is a dynamic network of protein filaments that provides structural support, enables cell motility, and facilitates intracellular transport and cell division. It is composed of three main types of protein filaments: microtubules, actin filaments, and intermediate filaments.
Supports cell shape and structure
Enables cell movement and division
Facilitates transport of organelles and vesicles
Highly dynamic and reorganized during cellular processes
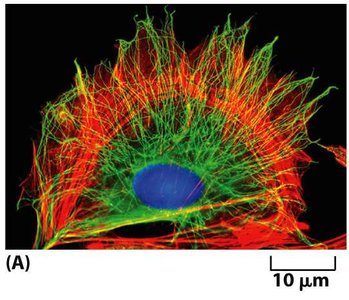
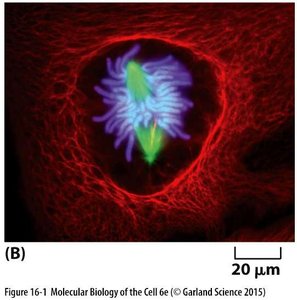
Organization of the Cytoskeleton in Epithelial Cells
Epithelial cells display a polarized organization of cytoskeletal filaments, which are essential for maintaining tissue structure and function.
Intermediate filaments provide mechanical strength.
Microtubules are attached to organizing centers (MTOCs).
Actin filaments are concentrated in the cell cortex and support cell shape and motility.
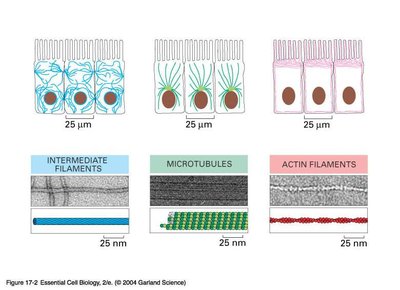
Types of Cytoskeletal Filaments
The three main types of cytoskeletal filaments differ in structure and function:
Filament Type | Structure | Main Function | Location |
|---|---|---|---|
Intermediate Filaments | Rope-like | Mechanical strength | Throughout cytoplasm |
Microtubules | Long, hollow cylinders | Organelle transport, cell division | Radiate from MTOCs |
Actin Filaments | Helical polymers | Cell shape, motility, cytokinesis | Cell cortex |
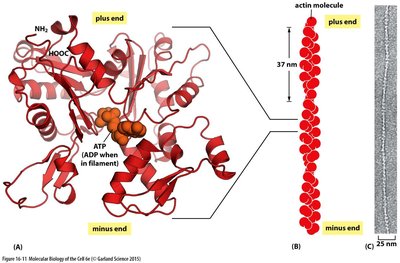
Actin Filaments: Structure and Function
Actin filaments (microfilaments) are found in all eukaryotic cells and are crucial for muscle contraction, cell movement, and cytokinesis. They serve as tracks for motor proteins and are involved in the formation of cellular structures such as the contractile ring during cell division.
Actin monomers (G-actin) polymerize to form filamentous actin (F-actin).
Filaments have structural polarity with distinct plus and minus ends.
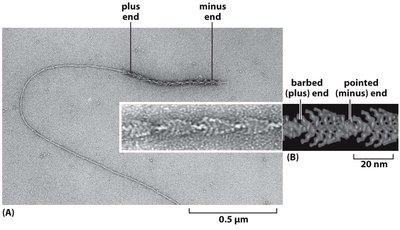

Actin Polymerization and Dynamics
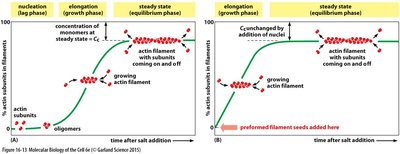
Actin polymerization is a dynamic process involving nucleation, elongation, and steady-state phases. ATP hydrolysis drives treadmilling, where subunits add at the plus end and dissociate from the minus end.
Nucleation is the rate-limiting step, requiring the formation of a stable actin nucleus.
Elongation proceeds rapidly once nucleation occurs.
Steady-state is reached when the rate of addition equals the rate of loss.
Actin Nucleation Factors
Actin nucleation is facilitated by proteins such as the Arp2/3 complex and formins. The Arp2/3 complex initiates the formation of branched actin networks, essential for cell motility and shape changes.
Arp2/3 complex binds to the side of existing filaments, creating a treelike web of actin.
Formins promote the formation of unbranched actin filaments.
*Additional info: The dynamic assembly and disassembly of actin filaments are regulated by a variety of accessory proteins, which control filament length, branching, and interactions with other cellular structures.*
