 Back
BackChromatin Organization and Function: Structure, Packaging, and Regulation of the Genome
Study Guide - Smart Notes

Chromatin Organization and the Genome
Physical Scale and Organization of the Genome
The eukaryotic genome is composed of long DNA polymers organized into chromosomes, which are further packaged within the nucleus. This organization is essential for the regulation, replication, and segregation of genetic material, especially in multicellular organisms where a single genome encodes the blueprints for diverse cell types.
Genome: The complete set of genetic material in an organism.
Chromosome: A single, long DNA molecule with associated proteins, carrying part of the genetic information.
Gene: A segment of DNA that encodes a functional product, typically a protein or RNA.
Chromosome number and size are characteristic of each species (e.g., humans have 46 chromosomes, budding yeast have 16, fission yeast have 3).
Genome size varies widely among species, but does not always correlate with gene number.
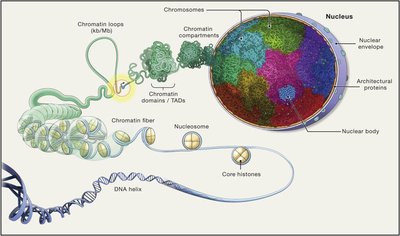
Genome Size and Non-Coding DNA
Genome size (C-value) and the proportion of non-coding DNA vary greatly among organisms. In complex eukaryotes, much of the genome does not encode proteins or functional RNAs, but may play roles in regulation and chromatin structure.
Human haploid genome: ~3.1 × 109 base pairs.
Non-coding DNA increases with organismal complexity.
Gene density is much higher in simple eukaryotes (e.g., S. cerevisiae: 483 genes per million base pairs) than in humans (12–15 genes per million base pairs).
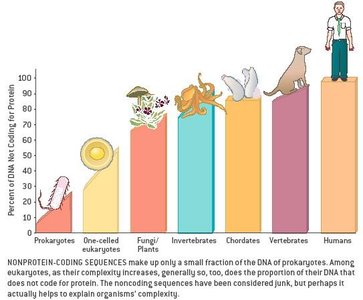
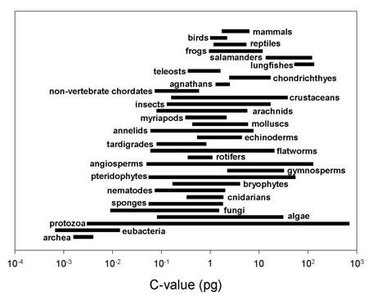
Gene Density and Genome Organization
Gene density varies not only between species but also within the genome of a single organism. Regions of high and low gene density reflect functional compartmentalization and regulatory complexity.
Gene-rich regions are interspersed with large stretches of non-coding DNA.
Comparative genomics reveals differences in gene organization among bacteria, yeast, flies, and humans.
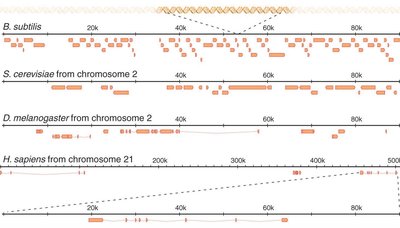
Chromatin Structure and Packaging
Levels of Chromatin Organization
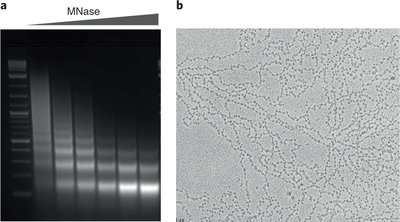
To fit the long DNA molecules into the small nuclear volume, eukaryotic cells employ hierarchical packaging strategies. Chromatin is the complex of DNA and proteins (mainly histones) that forms chromosomes.
Nucleosome: The basic unit of chromatin, consisting of ~146 bp of DNA wrapped around a histone octamer.
Chromatin fiber: Nucleosomes are further folded into higher-order structures, such as the 10 nm "beads-on-a-string" fiber and the 30 nm fiber.
Histone H1 promotes higher-order packaging beyond the nucleosome.

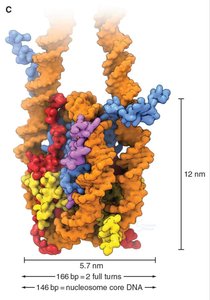
Nucleosome Structure and Histone Composition
The nucleosome core particle is composed of an octamer of histone proteins: two each of H2A, H2B, H3, and H4. DNA wraps around this core, and histone tails extend outward, subject to post-translational modifications that regulate chromatin function.
Nucleosomal wrapping shortens DNA length by about seven-fold.
Histone modifications (e.g., acetylation, methylation) influence gene expression and chromatin accessibility.
Higher-Order Chromatin Organization
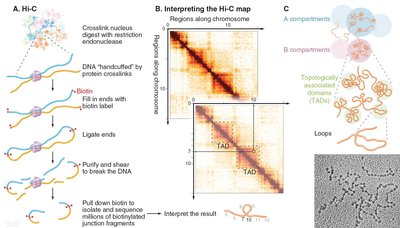
Chromatin Loops, TADs, and Compartments
Beyond the nucleosome, chromatin is organized into loops, topologically associating domains (TADs), and compartments. These structures facilitate gene regulation and genome stability.
Chromatin loops: Bring distant genomic regions into proximity, often mediated by protein complexes such as cohesin.
TADs: Domains within which chromatin interactions are frequent; boundaries are often defined by architectural proteins (e.g., CTCF, cohesin).
Compartments: Larger-scale segregation of active (A) and inactive (B) chromatin.
Role of SMC Complexes (Cohesin and Condensin)
Structural Maintenance of Chromosomes (SMC) complexes, such as cohesin and condensin, are ATPases that mediate the formation and maintenance of chromatin loops and higher-order structures. These complexes are essential for chromosome segregation and genome organization.
Cohesin extrudes DNA loops and helps establish TAD boundaries.
Condensin is critical for mitotic chromosome condensation and stability.

Centromeres and Chromosome Segregation
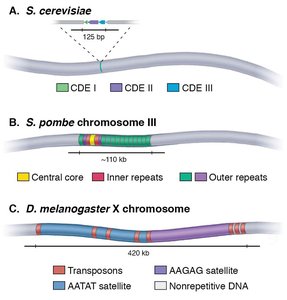
Centromere Structure and Function
The centromere is a specialized chromosomal region that forms the kinetochore, the attachment site for spindle microtubules during cell division. Centromeres are typically composed of repetitive DNA sequences and specialized nucleosomes containing CENP-A, a histone H3 variant.
Centromere DNA sequence and size vary among species (e.g., humans: alpha-satellite repeats; budding yeast: 125 bp point centromeres).
CENP-A nucleosomes are recognized by kinetochore proteins, ensuring proper chromosome segregation.
Clinical Relevance: Autoimmunity and Centromere Proteins
Autoimmune diseases such as scleroderma can involve the production of antibodies against centromeric proteins. Detection of anti-nuclear antibodies (ANA) is a diagnostic marker for such disorders.
Patients with scleroderma may have antibodies targeting CENP proteins.
ANA tests are commonly used in clinical diagnostics for autoimmune diseases.
Summary Table: Hierarchy of Chromatin Organization
Level | Structure | Function |
|---|---|---|
DNA Helix | Double-stranded DNA | Genetic information storage |
Nucleosome | DNA wrapped around histone octamer | Basic unit of chromatin, compaction |
Chromatin Fiber | 10 nm/30 nm fiber | Further compaction, regulation |
Loops | Chromatin loops (kb–Mb) | Gene regulation, enhancer-promoter contacts |
TADs | Topologically associating domains | Domain-level regulation, insulation |
Compartments | A/B compartments | Active/inactive chromatin segregation |
Chromosome | Entire DNA molecule with proteins | Genome transmission, segregation |
Additional info: Chromatin organization is dynamic and responsive to cellular signals, allowing for regulated access to genetic information during processes such as transcription, replication, and repair.
