 Back
BackComprehensive Study Notes on Enzymes
Study Guide - Smart Notes

Introduction to Enzymes
Definition and Importance
Enzymes are biological catalysts that accelerate chemical reactions in living organisms without being consumed in the process. Most enzymes are globular proteins, though some, such as ribozymes, are RNA molecules with catalytic activity. Enzymes are essential for metabolic processes, including respiration, photosynthesis, digestion, and biosynthesis of macromolecules.
Classification of Enzymes
Enzymes are classified based on the type of reaction they catalyze. The table below summarizes the main classes:
Class of Enzyme | Type of Reaction Catalysed | Examples |
|---|---|---|
Oxidoreductase | Transfers electrons, oxygen, or hydrogen atoms (oxidation-reduction reactions) | Dehydrogenases, Oxidases |
Transferase | Transfers functional groups between molecules | Kinases, Phosphorylases |
Hydrolase | Hydrolysis reactions | Sucrase, Lipases, Proteases |
Lyase | Removes groups without hydrolysis | Decarboxylases |
Isomerase | Rearranges groups within a molecule | Isomerases |
Ligase | Forms bonds using ATP energy | Synthetases, Ligases |
Note: Enzyme names typically end with '-ase'.
Characteristics of Enzymes
General Properties
Specificity: Enzymes are highly specific to their substrates due to the unique structure of their active sites.
Efficiency: Enzymes have high turnover rates, meaning a small amount can catalyze the conversion of large quantities of substrate.
Chemical Stability: Enzymes remain chemically unchanged after the reaction and can be reused.
Sensitivity: Enzyme activity is affected by substrate concentration, enzyme concentration, temperature, and pH.
Cofactors: Some enzymes require non-protein components (cofactors) for activity. These may be inorganic ions, coenzymes, or prosthetic groups.
Regulation: Enzyme activity is tightly regulated by activators and inhibitors.
Equilibrium: Enzymes accelerate the attainment of equilibrium but do not alter the equilibrium position.
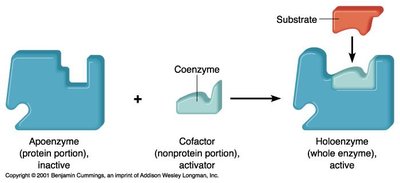
Cofactors: Apoenzyme and Holoenzyme
Apoenzyme: The protein portion of an enzyme, inactive without its cofactor.
Holoenzyme: The active enzyme with its cofactor bound.
Types of Cofactors:
Inorganic ions: e.g., Zn2+ for carbonic anhydrase.
Coenzymes: Organic molecules loosely bound, e.g., NAD+.
Prosthetic groups: Organic molecules tightly bound, e.g., haem in catalases.
Mode of Action of Enzymes
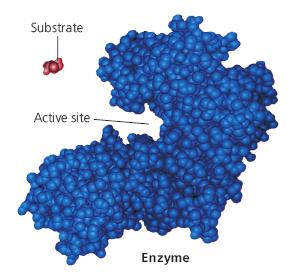
Active Site Structure
The active site is a specific region of the enzyme where substrate binding and catalysis occur. It is formed by a unique arrangement of amino acids, creating a three-dimensional pocket complementary to the substrate.
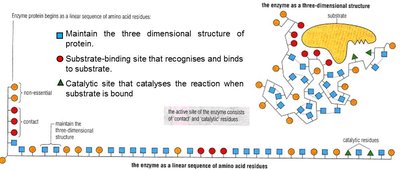
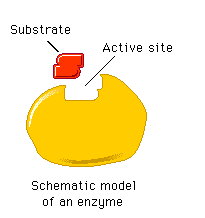
Enzyme Specificity
Determined by the fit between the enzyme's active site and its substrate (shape, size, charge, orientation).
Specificity arises from the enzyme's three-dimensional conformation, which is dictated by its amino acid sequence.
Enzyme-Substrate Complex
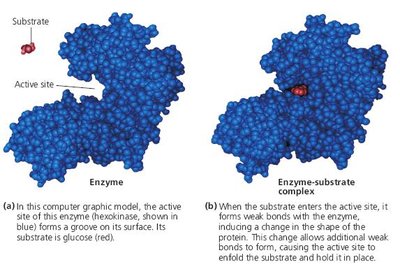
When a substrate binds to the enzyme's active site, an enzyme-substrate (E-S) complex forms. The substrate is converted to product, which then leaves the active site, allowing the enzyme to catalyze further reactions.
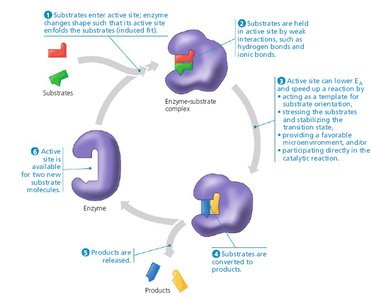
Lock-and-Key and Induced Fit Hypotheses
Lock-and-Key Hypothesis: The active site is perfectly complementary to the substrate, allowing precise binding.
Induced Fit Hypothesis: The active site is flexible and molds itself around the substrate upon binding, enhancing specificity and catalysis.
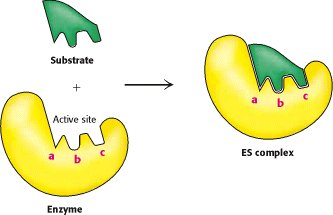
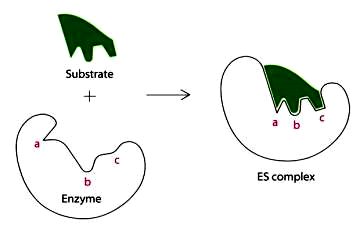
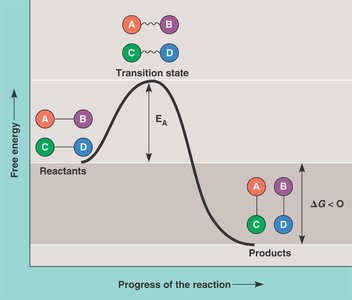
Activation Energy
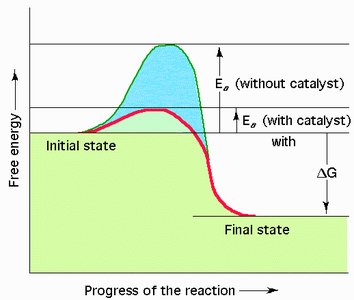
Activation energy (EA) is the minimum energy required for reactants to reach the transition state and undergo a chemical reaction. Enzymes lower the activation energy, allowing reactions to proceed faster at physiological temperatures.
Enzyme Kinetics
Measurement of Enzyme Kinetics
Reaction rates can be measured by the rate of product formation or substrate depletion.
Initial rates are determined from the linear portion of product or substrate vs. time graphs.
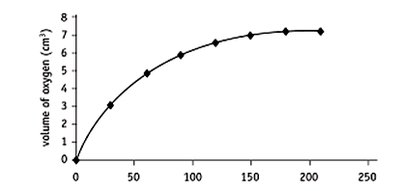
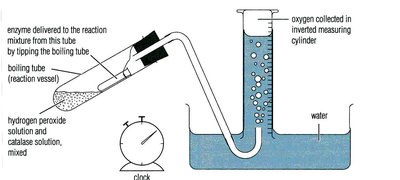
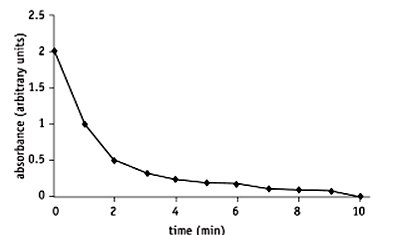
Measuring Substrate Disappearance
Example: Amylase catalyzing starch hydrolysis, monitored by iodine color change and absorbance.
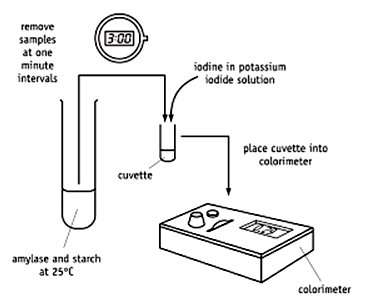
Michaelis Constant (Km)
Km is the substrate concentration at which the reaction rate is half of Vmax.
Low Km indicates high enzyme-substrate affinity; high Km indicates low affinity.
$K_m = [S] \text{ when } v = \frac{1}{2} V_{max}$
Factors Affecting Enzyme Activity
Substrate Concentration
At low substrate concentrations, the reaction rate increases proportionally with substrate concentration.
At high substrate concentrations, the enzyme becomes saturated, and the rate plateaus at Vmax.
Enzyme Concentration
At low enzyme concentrations, the reaction rate increases proportionally with enzyme concentration.
At high enzyme concentrations, the rate plateaus if substrate becomes limiting.
Temperature
Increasing temperature increases reaction rate up to an optimum, beyond which enzymes denature and activity drops sharply.
Q10 coefficient: Rate doubles for every 10°C rise within the optimal range.
$Q_{10} = \frac{\text{rate at } (x+10)^{\circ}C}{\text{rate at } x^{\circ}C}$
pH
Each enzyme has an optimum pH; deviations can reduce activity or denature the enzyme.
pH changes affect ionic and hydrogen bonds, altering the active site conformation.
Enzyme Inhibition
Reversible Inhibition
Inhibitors bind non-covalently and can dissociate, restoring enzyme activity.
Two main types: competitive and non-competitive inhibition.
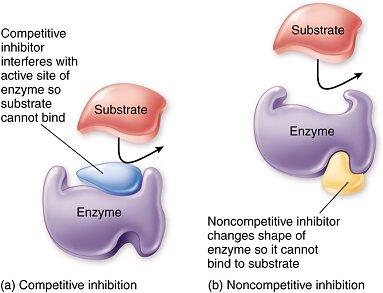
Competitive Inhibition
Inhibitor resembles substrate and binds to the active site, blocking substrate access.
Increases Km but does not affect Vmax.
Example: Malonate inhibits succinate dehydrogenase.
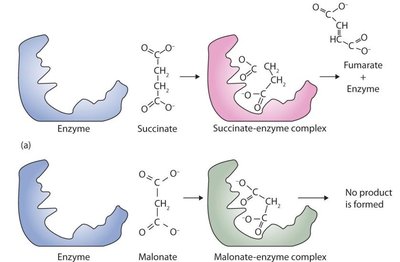
Non-Competitive Inhibition
Inhibitor binds to a site other than the active site, altering enzyme conformation.
Reduces Vmax but does not change Km.
Example: Cyanide inhibits cytochrome oxidase.
Irreversible Inhibition
Inhibitor binds covalently, permanently inactivating the enzyme.
Example: Heavy metals disrupt disulfide bonds in enzymes.
Allosteric Regulation of Enzymes
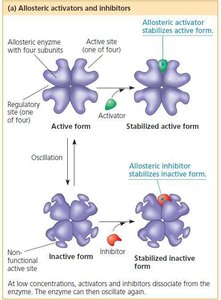
Allosteric Activation and Inhibition
Allosteric enzymes have multiple subunits and regulatory sites.
Binding of activators or inhibitors at allosteric sites stabilizes the active or inactive form of the enzyme, respectively.
Allows fine-tuned regulation of metabolic pathways.
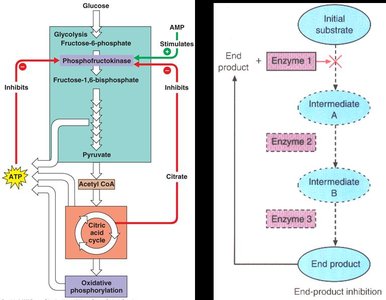
End-Product (Feedback) Inhibition
The final product of a metabolic pathway inhibits an earlier enzyme, preventing overproduction.
Example: ATP inhibits phosphofructokinase in glycolysis.
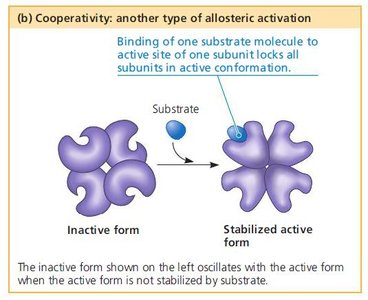
Cooperativity
Binding of a substrate to one active site increases the affinity of other active sites for the substrate.
Common in multi-subunit enzymes, enhancing sensitivity to substrate concentration.
Additional info: These notes provide a comprehensive overview of enzyme structure, function, kinetics, regulation, and inhibition, suitable for college-level cell biology students preparing for exams.
