 Back
BackControl of Gene Expression: Mechanisms and Regulation
Study Guide - Smart Notes

Gene Expression and Cell Differentiation
Overview of Gene Expression
Gene expression is the process by which cells selectively direct the synthesis of proteins and RNAs encoded in their genome. This selective expression is fundamental for cell differentiation, allowing cells to develop specialized functions.
Cell differentiation is achieved by changes in gene expression.
Gene expression involves multiple steps, each regulated to ensure proper cellular function.
Regulation can occur at transcription, RNA processing, transport, translation, and post-translational levels.
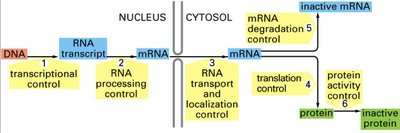
Transcriptional Control
Transcriptional Switches and Promoters
Transcriptional control is a primary mechanism for regulating gene expression, typically occurring at the initiation step.
Promoters are DNA sequences where RNA synthesis begins, including a transcription initiation site and upstream regions required for RNA polymerase recognition.
Upstream regions contain recognition sites for proteins that associate with the active polymerase, such as sigma factors in bacteria and general transcription factors in eukaryotes.
DNA regulatory sequences act as switches to turn genes on or off, ranging from short (10 nt pairs in bacteria) to long (10,000 nt pairs in eukaryotes).
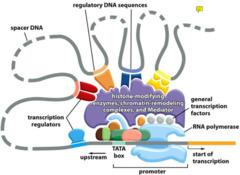
Transcription regulators bind to regulatory DNA sequences, controlling transcription initiation.
Gene Regulation in Bacteria: The Lac Operon
Operon Structure and Function
Bacteria regulate gene expression in response to environmental conditions, such as available food sources.
An operon is a cluster of genes under the control of a single promoter, common in bacteria but rare in eukaryotes.
The lac operon is a classic example, controlling genes involved in lactose metabolism.
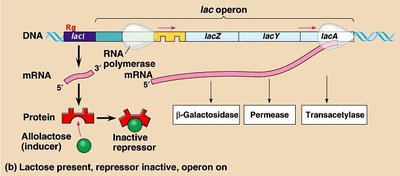
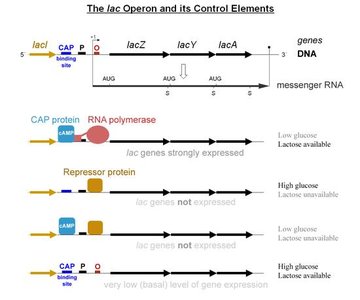
Lac Operon Control Elements
The lac operon is regulated by the presence of lactose and glucose, involving CAP protein, repressor protein, and RNA polymerase.
When lactose is present, the repressor is inactivated, allowing transcription of lac genes.
CAP protein enhances expression when glucose is low.
Repressor protein blocks expression when lactose is absent.
Eukaryotic Transcriptional Control
Regulators, Activators, and Repressors
Eukaryotic cells use transcriptional regulators, activators, and repressors to modulate gene expression.
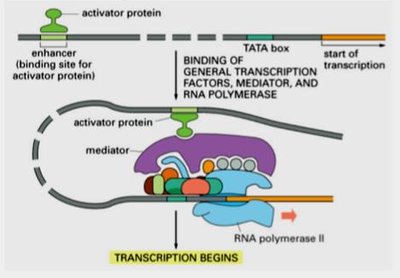
Transcriptional activators bind to enhancers, increasing gene expression.
Repressors inhibit transcription by binding to specific DNA sequences.
Complexes of general transcription factors, mediator proteins, and RNA polymerase II assemble at the promoter to initiate transcription.
Epigenetic Regulation
DNA Methylation
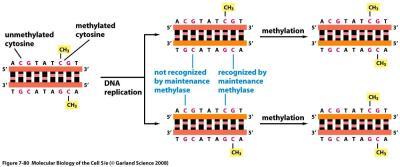
DNA methylation is a heritable modification that can silence gene expression.
Methylation of cytosine residues in DNA can prevent transcription factor binding.
Maintenance methylases ensure methylation patterns are preserved during DNA replication.
Histone Modification
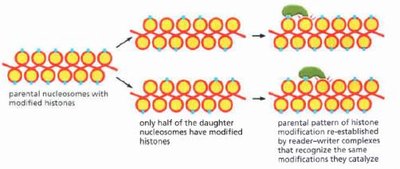
Histone modifications affect chromatin structure and gene accessibility.
Parental nucleosomes with modified histones can transmit modification patterns to daughter cells.
Reader-writer complexes recognize and re-establish histone modifications after DNA replication.
Post-Transcriptional Controls
Mechanisms of Post-Transcriptional Regulation
Post-transcriptional controls operate after transcription has begun and are crucial for regulating gene expression.
Alternative splicing allows a single gene to produce multiple protein variants.
Post-translational modifications alter protein function after synthesis.
mRNA degradation and translation control determine the amount of protein produced.
MicroRNAs (miRNAs) and RNA interference (RNAi) are important mechanisms for silencing gene expression.
MicroRNAs (miRNAs)
miRNAs are small RNA molecules that base-pair with specific mRNAs, reducing their stability and translation.
In humans, miRNAs regulate at least 1/3 of all protein-coding genes.
RNA Interference (RNAi)
RNAi is a defense mechanism against foreign RNA molecules.
Double-stranded RNAs are cut into short fragments (~22 nt) by Dicer.
Small interfering RNAs (siRNAs) are incorporated into RISC complexes, which use single-stranded RNA to target and destroy complementary foreign RNA.
Summary Table: Levels of Gene Expression Control
Level | Mechanism | Example |
|---|---|---|
Transcriptional | Regulation of RNA synthesis initiation | Promoter, enhancers, repressors |
RNA Processing | Modification of RNA transcript | Splicing, capping, polyadenylation |
RNA Transport | Localization and export of mRNA | mRNA export from nucleus |
Translation | Control of protein synthesis | miRNA, ribosome binding |
mRNA Degradation | Stability and breakdown of mRNA | Deadenylation, miRNA targeting |
Protein Activity | Post-translational modification | Phosphorylation, ubiquitination |
Key Equations and Concepts
Central Dogma of Molecular Biology:
Gene Regulation Equation:
Additional info:
Epigenetic mechanisms such as DNA methylation and histone modification are crucial for long-term regulation and inheritance of gene expression patterns.
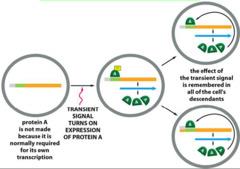
Transcriptional memory can ensure that transient signals affecting gene expression are remembered in cell lineages.
