 Back
BackDNA Replication and the Eukaryotic Cell Cycle: Mechanisms and Regulation
Study Guide - Smart Notes

DNA Sequencing and Bioinformatics
Pyrosequencing
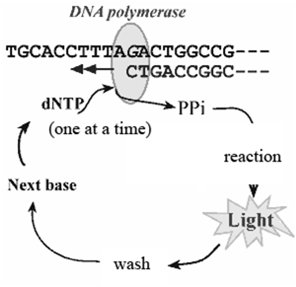
Pyrosequencing is a DNA sequencing technique that detects the incorporation of nucleotides during DNA synthesis by measuring the release of pyrophosphate, which is converted into a detectable light signal. This method allows for the parallel sequencing of millions of DNA fragments.
Process: DNA polymerase adds nucleotides one at a time to a growing DNA strand. Each incorporation releases pyrophosphate (PPi), which, through a series of enzymatic reactions involving luciferase, produces a flash of light.
Detection: The light intensity corresponds to the number of nucleotides incorporated, allowing the sequence to be determined.
Applications: Used for rapid sequencing of short DNA regions, mutation detection, and microbial identification.
DNA Databases and BLAST
Bioinformatics tools such as BLAST (Basic Local Alignment Search Tool) are essential for analyzing DNA and protein sequences. BLAST compares a query sequence to sequences in a database to identify homologous regions.
Nucleotide BLAST (blastn): Compares nucleotide sequences.
Protein BLAST: Compares amino acid sequences.
Applications: Gene identification, evolutionary studies, and functional annotation.
The Eukaryotic Cell Cycle
Overview of Cell Cycle Phases
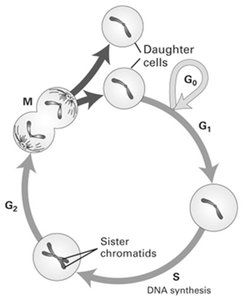
The eukaryotic cell cycle is a series of ordered events that lead to cell division and the production of two daughter cells. It ensures the accurate duplication and segregation of genetic material.
M phase (Mitosis): The cell physically divides to produce two daughter cells.
Interphase: Consists of G1, S, and G2 phases.
G1 phase: Cell growth and preparation for DNA replication.
S phase: DNA synthesis; the amount of DNA doubles.
G2 phase: Further growth and preparation for mitosis.
G0 phase: A quiescent state where cells exit the cycle, either temporarily (e.g., stem cells) or permanently (e.g., neurons).
Timing of Cell Cycle Phases
Doubling time: The time required for a cell to divide (18–24 hours for typical mammalian cells).
G1 phase: Highly variable, typically 8–10 hours.
S phase: 6–8 hours.
G2 phase: 4–6 hours.
M phase: 30–45 minutes.
G0 Phase and Cell Differentiation
Cells in the G0 phase have exited the cell cycle. Some are terminally differentiated (e.g., neurons), while others (e.g., stem cells) can re-enter the cycle in response to signals such as growth factors.
Permanent G0: Cells that no longer divide unless transformed (e.g., cancerous transformation).
Temporary G0: Cells that can re-enter the cycle for tissue repair or replenishment.
G1 Phase: Cell Growth
During G1, the cell increases in size and prepares for DNA replication. The length of G1 can vary greatly depending on cell type and conditions.
Fast-dividing cells: May have a very short or absent G1 phase (e.g., embryonic cells).
Preparation for S phase: Synthesis of proteins and organelles required for DNA replication.
S Phase and G2 Phase
In S phase, DNA replication occurs, doubling the genetic material. The G2 phase is a second growth phase where the cell prepares for mitosis, ensuring DNA replication is complete and repairing any DNA damage.
DNA content: Doubles from 2N to 4N after S phase.
G2 phase: Checkpoints ensure readiness for mitosis.
Mechanisms of DNA Replication
Semiconservative Replication
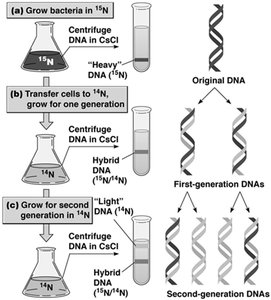
DNA replication is semiconservative, meaning each daughter DNA molecule consists of one parental and one newly synthesized strand. This was demonstrated by the Meselson-Stahl experiment using isotopic labeling and density gradient centrifugation.
Semiconservative model: Each new DNA double helix contains one old and one new strand.
Meselson-Stahl experiment: Used 15N and 14N isotopes to distinguish parental and daughter DNA strands.
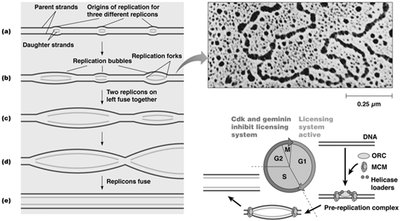
Multiple Replicons in Eukaryotic Genomes
Eukaryotic chromosomes contain multiple origins of replication, forming replicons. Replication bubbles form at these origins, allowing rapid and efficient duplication of large genomes.
Replicons: Units of DNA in which replication is initiated.
Replication bubbles: Regions where DNA is locally unwound for replication.
Temporal regulation: Early and late replicating domains are spatially separated in the nucleus.
Directionality of DNA Synthesis
DNA polymerase synthesizes new DNA in the 5' to 3' direction by adding nucleotides to the free 3' hydroxyl group. This directionality is due to the enzyme's mechanism and the chemical properties of nucleotides.
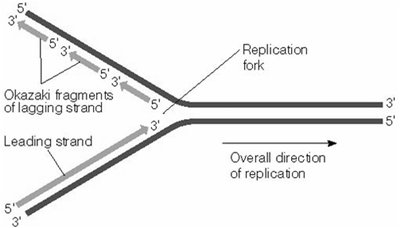
Leading strand: Synthesized continuously in the direction of the replication fork.
Lagging strand: Synthesized discontinuously as Okazaki fragments, later joined by DNA ligase.
Proteins Involved in DNA Replication
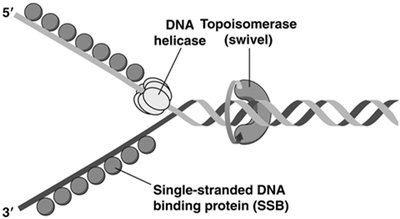
Several proteins coordinate the unwinding and synthesis of DNA at the replication fork:
DNA helicase: Unwinds the double helix.
Topoisomerase: Relieves supercoiling tension ahead of the fork.
Single-stranded DNA binding proteins (SSB): Stabilize unwound DNA.
Primase: Synthesizes short RNA primers to initiate DNA synthesis.
DNA polymerase: Extends the DNA strand from the primer.
DNA ligase: Joins Okazaki fragments on the lagging strand.
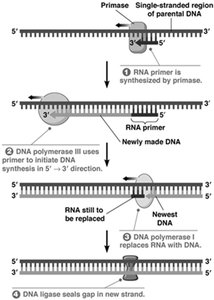
RNA Primers and Proofreading
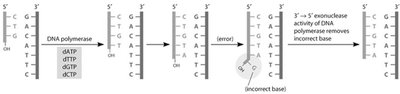
DNA polymerase requires a primer with a free 3' OH group to initiate synthesis. Primase synthesizes short RNA primers, which are later removed and replaced with DNA. DNA polymerase also has proofreading activity to correct errors.
Primer removal: 5' to 3' exonuclease activity removes RNA primers.
Proofreading: 3' to 5' exonuclease activity corrects misincorporated nucleotides.
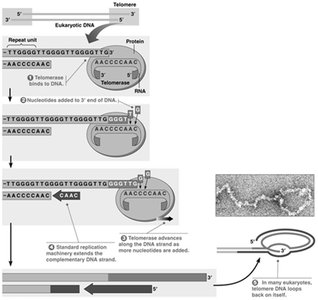
Telomeres and Telomerase
Linear chromosomes face the "end-replication problem," where the lagging strand cannot be fully replicated at the 3' end. Telomerase extends telomeres, repetitive DNA sequences at chromosome ends, to prevent loss of genetic information.
Telomerase: A ribonucleoprotein enzyme that adds telomeric repeats to the 3' end of chromosomes.
Importance: Maintains chromosome integrity and prevents cellular aging.
DNA Topology: Supercoiling and Topoisomerases
Supercoiling of DNA
DNA can exist in relaxed or supercoiled forms. Supercoiling compacts DNA and is regulated by topoisomerases.
Positive supercoiling: Overwinding of the DNA helix.
Negative supercoiling: Underwinding of the DNA helix.
Topoisomerases: Enzymes that introduce or remove supercoils by creating transient breaks in the DNA backbone.
Type I topoisomerases: Make single-strand breaks.
Type II topoisomerases: Make double-strand breaks (e.g., DNA gyrase in bacteria).
Additional info: Topoisomerases are essential for DNA replication, transcription, and chromosome segregation, as they prevent excessive supercoiling and tangling of DNA.
