 Back
Back17: Energy Conversion, Photosynthesis, and Protein Sorting in Eukaryotic Cells
Study Guide - Smart Notes

Energy Conversion in Photosynthesis
Photosynthetic Electron Transport Chain
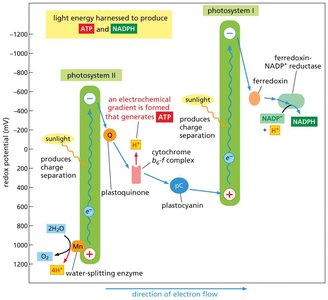
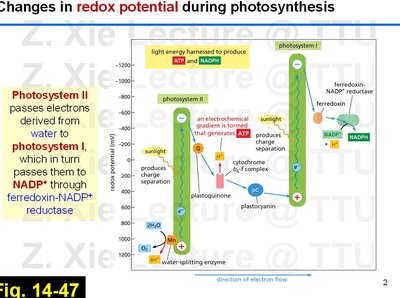
The photosynthetic electron transport chain is a series of protein complexes and mobile carriers embedded in the thylakoid membrane of chloroplasts. It is responsible for converting light energy into chemical energy in the form of ATP and NADPH.
Photosystem II (PSII): Absorbs light energy, which excites electrons derived from water, releasing O2 and protons (H+).
Electron Transport: Electrons are transferred from PSII to plastoquinone (Q), then through the cytochrome b6-f complex, plastocyanin (pC), and finally to Photosystem I (PSI).
Photosystem I (PSI): Absorbs additional light energy, further exciting electrons, which are then transferred to ferredoxin (Fd) and ultimately to NADP+ via ferredoxin-NADP+ reductase (FNR), forming NADPH.
ATP Synthesis: The movement of electrons is coupled to the translocation of protons across the thylakoid membrane, generating a proton gradient that drives ATP synthesis via ATP synthase.
Redox Potential: The flow of electrons is energetically downhill, from a low to a high redox potential, enabling the synthesis of ATP and NADPH.
Electron Flow and Proton Movement
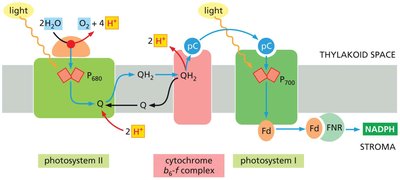
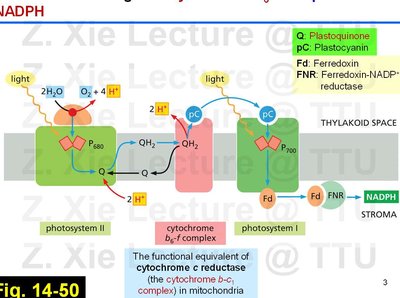
Electron flow through the cytochrome b6-f complex is analogous to the cytochrome b-c1 complex in mitochondria. This process is tightly coupled to proton translocation, which is essential for chemiosmotic ATP synthesis.
Key Carriers: Plastoquinone (Q), plastocyanin (pC), and ferredoxin (Fd) shuttle electrons between complexes.
Proton Gradient: Protons are pumped from the stroma into the thylakoid space, creating an electrochemical gradient.
ATP and NADPH Formation: The proton gradient powers ATP synthase, while electrons reduce NADP+ to NADPH.
Comparison of Mitochondria and Chloroplasts
Energy Conversion Mechanisms
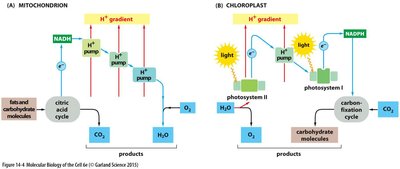
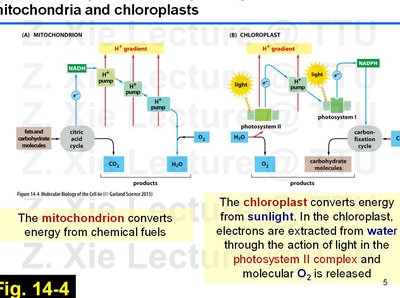
Both mitochondria and chloroplasts use electron transport chains and chemiosmotic gradients to synthesize ATP, but they differ in their energy sources and products.
Mitochondria: Convert energy from chemical fuels (e.g., glucose) via the citric acid cycle and oxidative phosphorylation, producing ATP, CO2, and H2O.
Chloroplasts: Convert energy from sunlight via the light reactions of photosynthesis, producing ATP, NADPH, and O2.
Proton Gradients: In mitochondria, protons are pumped into the intermembrane space; in chloroplasts, into the thylakoid space.
Chemiosmotic Theory and Experimental Evidence
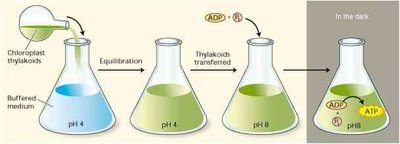
The chemiosmotic theory, proposed by Peter Mitchell, states that ATP synthesis is driven by a proton gradient across a membrane. André Jagendorf provided direct experimental evidence for this mechanism in chloroplasts.
Jagendorf Experiment: Isolated chloroplast thylakoids were equilibrated at pH 4, then transferred to pH 8 buffer containing ADP and Pi. The resulting proton gradient drove ATP synthesis in the dark, confirming the chemiosmotic mechanism.
Carbon Fixation and the Calvin Cycle
Role of Rubisco
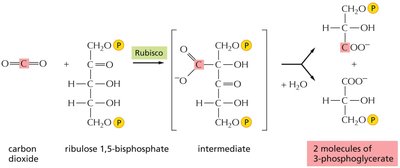
Ribulose bisphosphate carboxylase/oxygenase (Rubisco) catalyzes the first step of carbon fixation in the Calvin cycle, converting CO2 and ribulose 1,5-bisphosphate into two molecules of 3-phosphoglycerate.
Rubisco: The most abundant protein on Earth, but relatively slow in catalytic activity.
Location: Functions in the stroma of chloroplasts.
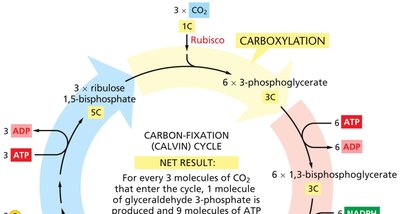
The Calvin Cycle
The Calvin cycle is the set of biochemical reactions that assimilate atmospheric CO2 into organic molecules using ATP and NADPH produced in the light reactions.
Carboxylation: CO2 is fixed by Rubisco to form 3-phosphoglycerate.
Reduction: 3-phosphoglycerate is converted to glyceraldehyde 3-phosphate using ATP and NADPH.
Regeneration: Ribulose 1,5-bisphosphate is regenerated to continue the cycle.
Products: For every 3 CO2 molecules fixed, 1 molecule of glyceraldehyde 3-phosphate is produced, which can be used to synthesize sugars, fats, and amino acids.
Genetic Systems of Mitochondria and Chloroplasts
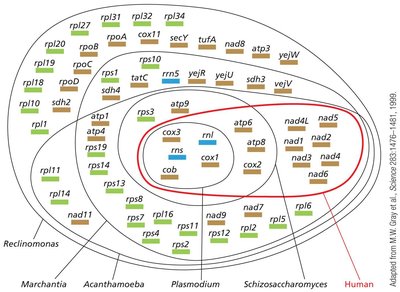
Organelle Genomes
Mitochondria and chloroplasts contain their own genomes, which encode a small subset of the proteins required for organelle function. Most organellar proteins are encoded by nuclear DNA and imported into the organelles.
Genome Structure: Both organelles have circular DNA molecules, with mitochondria typically having smaller genomes than chloroplasts.
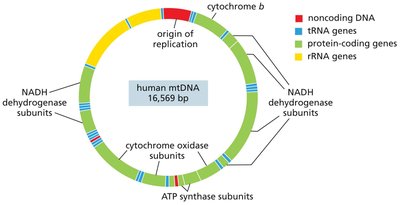
Gene Content: Human mitochondrial DNA (~16.6 kb) encodes 13 proteins, 22 tRNAs, and 2 rRNAs. Chloroplast genomes are larger and encode more genes.
Inheritance: Organelle genes are usually maternally inherited in animals and plants.
Protein Sorting and Intracellular Membrane Trafficking
Major Intracellular Compartments
Eukaryotic cells contain distinct membrane-enclosed organelles, each specialized for particular functions. These compartments are separated by selectively permeable membranes.
Nucleus: Contains the genetic material and is surrounded by a double membrane (nuclear envelope).
Endoplasmic Reticulum (ER): Site of protein and lipid synthesis; rough ER is studded with ribosomes, while smooth ER is involved in lipid metabolism.
Golgi Apparatus: Modifies, sorts, and packages proteins and lipids for delivery to various destinations.
Lysosomes: Contain hydrolytic enzymes for degradation of macromolecules.
Protein Sorting Mechanisms
Proteins are directed to their correct cellular locations by sorting signals within their amino acid sequences. Three main mechanisms are used for protein import into organelles:
Gated Transport: Proteins move between the cytosol and nucleus through nuclear pore complexes.
Protein Translocation: Proteins are translocated across membranes into the ER, mitochondria, or chloroplasts, often in an unfolded state.
Vesicular Transport: Proteins are transported between organelles via membrane-bound vesicles.
Nuclear Import and Export
Nuclear pore complexes (NPCs) regulate the movement of molecules between the nucleus and cytosol. Nuclear localization signals (NLS) direct proteins to the nucleus, where they are recognized by nuclear import receptors (importins).
Nuclear Localization Signal (NLS): A specific amino acid sequence that targets proteins to the nucleus.
Importins: Receptors that bind NLS-containing proteins and mediate their transport through the NPC.
Ran GTPase Cycle: Provides directionality for nuclear import and export by regulating the binding and release of cargo proteins.
Summary Table: Comparison of Mitochondria and Chloroplasts
Feature | Mitochondrion | Chloroplast |
|---|---|---|
Energy Source | Chemical fuels (e.g., glucose) | Sunlight |
Main Products | ATP, CO2, H2O | ATP, NADPH, O2 |
Electron Donor | NADH, FADH2 | H2O |
Location of Proton Gradient | Across inner mitochondrial membrane | Across thylakoid membrane |
ATP Synthase Location | Inner mitochondrial membrane | Thylakoid membrane |
