 Back
BackEnzymes: The Catalysts of Life – Comprehensive Study Notes
Study Guide - Smart Notes

Enzymes: The Catalysts of Life
Introduction to Enzymes
Enzymes are biological catalysts that dramatically increase the rate of chemical reactions in cells, making the difference between reactions that can occur and those that will occur under physiological conditions. They are essential for life, as they enable reactions to proceed rapidly and efficiently at the temperatures and conditions found within living organisms.
Activation Energy and the Metastable State
Many reactions in cells are thermodynamically feasible but do not occur at appreciable rates due to high activation energy (EA). For example, the hydrolysis of ATP is energetically favorable ( kcal/mol), but ATP remains stable in water for days without catalysis. The activation energy barrier must be overcome for the reaction to proceed.
Activation Energy (EA): The minimum energy required to initiate a chemical reaction.
Metastable State: A state where reactants are stable and do not react spontaneously.
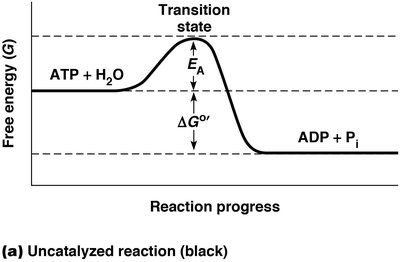
Effect of Temperature on Reaction Rate
Increasing temperature raises the kinetic energy of molecules, increasing the proportion of molecules with enough energy to overcome EA. However, cells are isothermal (constant temperature), so this method is not practical for biological systems.
Isothermal: Maintaining a constant temperature (homeostasis).
Thermal Activation: More molecules reach EA at higher temperatures.
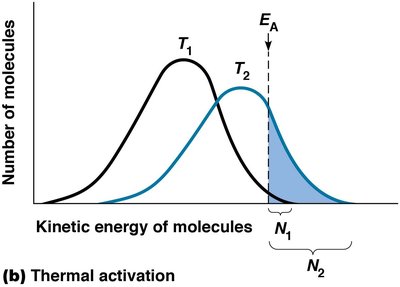
Lowering Activation Energy: Catalysts
Catalysts, including enzymes, lower the activation energy by providing a surface for reactants to bind and interact, facilitating the reaction. Catalysts are not consumed or altered in the reaction.
Properties of Catalysts:
Increase reaction rates by lowering EA.
Form transient, reversible complexes with substrates.
Change the rate at which equilibrium is achieved, not the equilibrium position.
Enzymes as Biological Catalysts
Most enzymes are proteins, but some RNA molecules (ribozymes) also have catalytic activity. Enzymes are highly specific for their substrates due to the unique structure of their active sites.
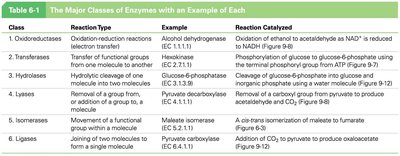
Major Classes of Enzymes
Enzymes are classified into six major classes based on the type of reaction they catalyze:
Class | Reaction Type | Example | Reaction Catalyzed |
|---|---|---|---|
Oxidoreductases | Oxidation-reduction reactions | Alcohol dehydrogenase | Oxidation of ethanol to acetaldehyde |
Transferases | Transfer of functional groups | Hexokinase | Phosphorylation of glucose |
Hydrolases | Hydrolytic cleavage | Glucose-6-phosphatase | Cleavage of glucose-6-phosphate |
Lyases | Removal/addition of groups | Pyruvate decarboxylase | Removal of carboxyl group from pyruvate |
Isomerases | Isomerization | Malate isomerase | Cis-trans isomerization of malate |
Ligases | Joining molecules | Pyruvate carboxylase | Addition of CO2 to pyruvate |

The Active Site of Enzymes
The active site is a specific region formed by the three-dimensional folding of the enzyme, where substrates bind and catalysis occurs. The active site is usually a groove or pocket with high affinity for the substrate.
Enzyme Specificity: The shape and chemistry of the active site confer high substrate specificity.
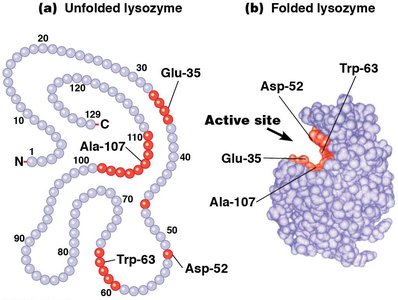
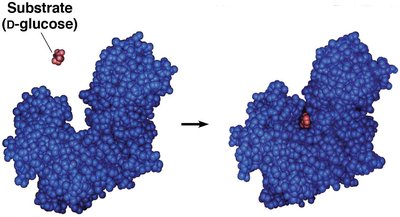
Enzyme-Substrate Interaction and Induced-Fit Model
Enzymes bind substrates through noncovalent interactions, positioning them optimally for catalysis. The induced-fit model describes how the enzyme changes shape upon substrate binding, bringing necessary amino acid side chains into the active site.
Cofactors and Prosthetic Groups
Some enzymes require nonprotein cofactors for catalytic activity. These include metal ions and small organic molecules called coenzymes, often derived from vitamins. The presence of cofactors explains the need for trace vitamins and minerals.
Enzyme Inhibition
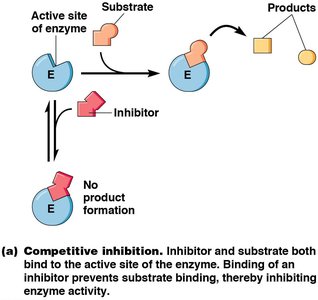
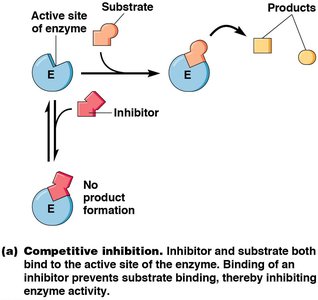
Enzyme inhibitors can be irreversible (covalently binding and permanently inactivating the enzyme) or reversible (noncovalently binding and able to dissociate). Reversible inhibitors are classified as competitive or noncompetitive:
Competitive Inhibitors: Bind to the active site, preventing substrate binding. Increase Km, no effect on Vmax.
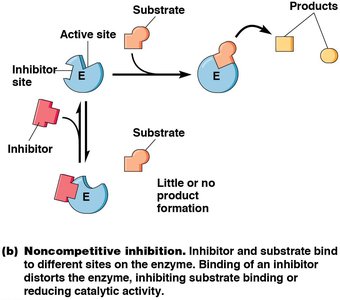
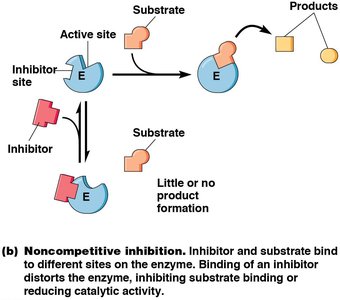
Noncompetitive Inhibitors: Bind to a different site, altering enzyme activity. Decrease Vmax, no effect on Km.
Enzyme Regulation
Enzyme activity is regulated to meet cellular needs. Regulation can occur at the substrate level or through allosteric mechanisms:
Substrate-Level Regulation: Increased substrate increases reaction rate; increased product decreases rate.
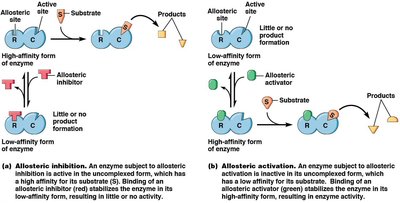
Allosteric Regulation: Enzymes have two conformations, one with high substrate affinity and one with low. Allosteric effectors (activators or inhibitors) bind to regulatory sites, altering enzyme activity.
Feedback Inhibition
Feedback inhibition is a regulatory mechanism where the end product of a pathway inhibits an earlier step, preventing overproduction and maintaining homeostasis.
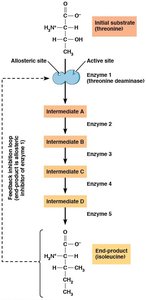
Covalent Modification
Enzyme activity can also be regulated by covalent modification, such as phosphorylation, methylation, or acetylation. These modifications alter enzyme activity and are reversible.
Enzyme Kinetics
Enzyme kinetics describes the quantitative aspects of catalysis and substrate conversion. Reaction rates depend on substrate, product, and inhibitor concentrations. The Michaelis–Menten equation models the relationship between reaction velocity and substrate concentration:
Km (Michaelis constant): Substrate concentration at half-maximal velocity; indicates enzyme-substrate affinity.
Vmax: Maximum reaction velocity when the enzyme is saturated with substrate.
kcat: Turnover number; rate at which substrate is converted to product by a single enzyme at Vmax.
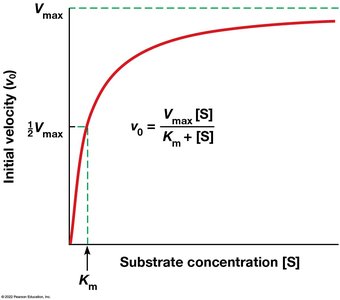
Michaelis–Menten Equation:
Examples and Applications
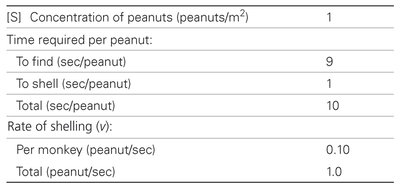
Enzyme kinetics can be illustrated using analogies, such as monkeys shelling peanuts, to demonstrate the effects of substrate concentration and enzyme efficiency.
[S] (peanuts/m2) | Time to find (sec/peanut) | Time to shell (sec/peanut) | Total (sec/peanut) | Rate per monkey (peanut/sec) | Total (peanut/sec) |
|---|---|---|---|---|---|
1 | 9 | 1 | 10 | 0.10 | 1.0 |
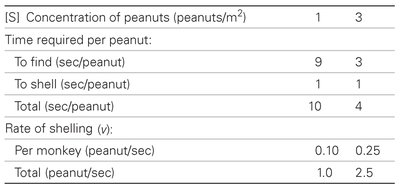
[S] (peanuts/m2) | Time to find (sec/peanut) | Time to shell (sec/peanut) | Total (sec/peanut) | Rate per monkey (peanut/sec) | Total (peanut/sec) |
|---|---|---|---|---|---|
1 | 9 | 1 | 10 | 0.10 | 1.0 |
3 | 3 | 1 | 4 | 0.25 | 2.5 |
Enzyme Inhibition Kinetics
Competitive Inhibition: Increases Km, no effect on Vmax.
Noncompetitive Inhibition: Decreases Vmax, no effect on Km.
Summary Table: Enzyme Kinetic Parameters
Enzyme Name | Substrate | Km (M) | kcat (sec-1) |
|---|---|---|---|
Acetylcholinesterase | Acetylcholine | 9 × 10-5 | 1.4 × 104 |
Carbonic anhydrase | CO2 | 1 × 10-2 | 1 × 106 |
Fumarase | Fumarate | 5 × 10-6 | 8 × 102 |
Triose phosphate isomerase | Glyceraldehyde-3-phosphate | 5 × 10-4 | 4.3 × 103 |
β-lactamase | Benzylpenicillin | 2 × 10-5 | 2 × 103 |
Conclusion
Enzymes are fundamental to cellular life, enabling rapid, specific, and regulated biochemical reactions. Understanding their structure, function, regulation, and kinetics is essential for cell biology and biochemistry.
