 Back
BackGene Expression and Regulation: Mechanisms and Control in Eukaryotic Cells
Study Guide - Smart Notes

Gene Expression and Regulation in Eukaryotic Cells
Overview of the Genetic Code and Translation
Gene expression is the process by which information from a gene is used to synthesize functional gene products, primarily proteins. The translation of mRNA into protein involves the genetic code, tRNAs, and ribosomes.
Genetic Code: The genetic code is degenerate, meaning multiple codons can encode the same amino acid, but each codon specifies only one amino acid.
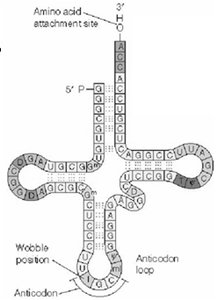
tRNA Structure and Function: Transfer RNAs (tRNAs) are adaptor molecules that match amino acids to codons in mRNA during translation. Each tRNA has an anticodon that pairs with a codon and an amino acid attachment site.
Wobble Hypothesis: The third position of the codon (wobble position) allows for non-standard base pairing, enabling one tRNA to recognize multiple codons.
Aminoacyl-tRNA: tRNAs are charged with their corresponding amino acids by aminoacyl-tRNA synthetases, forming aminoacyl-tRNAs (activated amino acids).
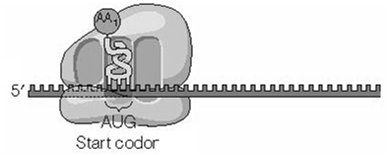
Translation Initiation: The ribosome assembles at the start codon (AUG) on the mRNA, and the first aminoacyl-tRNA binds to the P site.
mRNA Processing in Eukaryotes
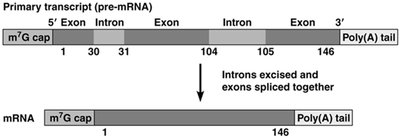
Eukaryotic mRNAs undergo extensive processing before translation, including capping, splicing, and polyadenylation.
Primary Transcript (pre-mRNA): Contains both exons (coding regions) and introns (non-coding regions).
RNA Splicing: Introns are excised, and exons are joined to form mature mRNA.
5' Cap and 3' Poly(A) Tail: The 5' end is capped with a methylguanosine, and the 3' end receives a poly(A) tail, both of which are important for mRNA stability and translation.
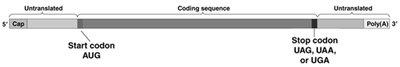
mRNA Structure: Mature mRNA contains untranslated regions (UTRs) at both ends, a coding sequence, a start codon (AUG), and a stop codon (UAG, UAA, or UGA).
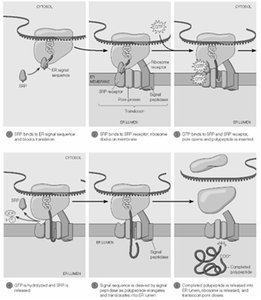
Cotranslational Import and Protein Targeting to the Endoplasmic Reticulum (ER)
Proteins destined for secretion or for the plasma membrane are synthesized with an N-terminal ER signal sequence that directs the ribosome to the ER membrane.
Signal Recognition Particle (SRP): Binds the signal sequence and pauses translation until the ribosome docks at the ER translocon.
Translocon: A protein-conducting channel in the ER membrane through which the nascent polypeptide enters the ER lumen.
GTP Hydrolysis: Provides energy for the transfer of the signal sequence and resumption of translation.
Signal Peptidase: Cleaves the signal sequence as the protein enters the ER.
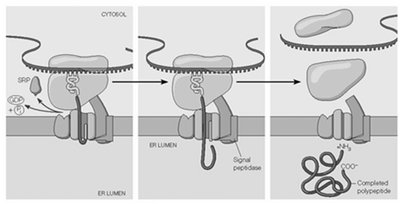
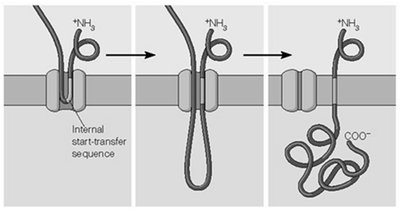
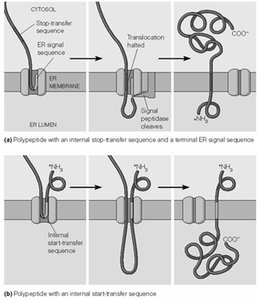
Transmembrane Protein Topology and ER Targeting
The arrangement of transmembrane domains in proteins is determined by the presence and position of start-transfer and stop-transfer sequences.
Single-Pass Proteins: Proteins with a signal peptide but no other transfer sequences are fully translocated into the ER lumen.
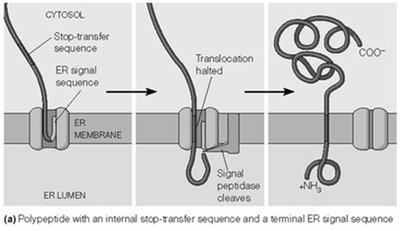
Stop-Transfer Sequence: Halts translocation, anchoring the protein in the membrane with a defined topology.
Internal Start-Transfer Sequence: Initiates translocation from within the polypeptide, resulting in different membrane orientations.
Regulation of Gene Expression
Gene expression is regulated at multiple levels, including chromatin structure, transcription, RNA processing, translation, and post-translational modifications.
Chromatin Structure: Chromatin can be condensed (heterochromatin, gene silencing) or decondensed (euchromatin, gene activation).
DNA Methylation: Addition of methyl groups to cytosine residues (especially in CpG islands) generally leads to gene silencing.
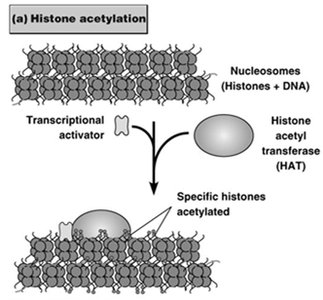
Histone Modifications: Acetylation of histones by HATs promotes gene activation; deacetylation by HDACs leads to repression. Methylation can have activating or repressing effects depending on the residue modified.
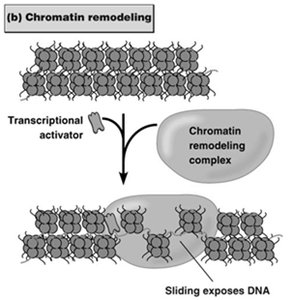
Chromatin Remodeling: ATP-dependent complexes reposition nucleosomes to expose or hide DNA regulatory elements.
Transcriptional Regulation
Transcription is the primary point of gene regulation, involving interactions between DNA elements and protein factors.
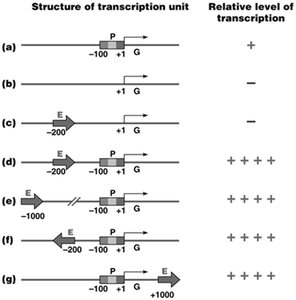
Cis-acting Elements: DNA sequences such as promoters, enhancers, silencers, and control elements.
Trans-acting Factors: Proteins including general transcription factors, activators, and repressors.
Enhancers: Can function at variable distances and orientations to increase transcription levels.
Combinatorial Control: Multiple transcription factors and elements interact to fine-tune gene expression in a tissue-specific manner.

Post-Transcriptional Regulation
Gene expression can also be regulated after transcription, including mRNA splicing, stability, and translation efficiency.
Alternative Splicing: Allows a single gene to produce multiple protein isoforms.
mRNA Stability: The half-life of mRNA affects how much protein is produced.
Translational Control: Regulatory proteins and small RNAs (miRNA, siRNA) can inhibit translation or promote mRNA degradation.

RNA Interference and Small RNAs
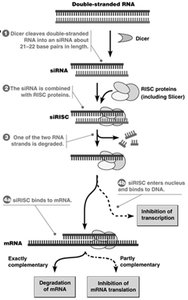
Small RNAs such as siRNAs and miRNAs regulate gene expression by targeting mRNAs for degradation or translational repression.
siRNA Pathway: Dicer processes double-stranded RNA into siRNAs, which are loaded into RISC to target complementary mRNAs for degradation.
miRNA Pathway: miRNAs are processed from hairpin precursors and guide RISC to partially complementary mRNAs, leading to translational inhibition or degradation.

Summary Table: Levels of Gene Expression Regulation
Level | Mechanisms |
|---|---|
Genome | Gene amplification, DNA rearrangement, chromatin decondensation, DNA methylation |
Transcription | Transcription factors, enhancers, silencers, promoter usage |
RNA Processing | Splicing, capping, polyadenylation, RNA editing |
Translation | mRNA stability, translation efficiency, miRNA/siRNA regulation |
Post-translation | Protein folding, modification, degradation (ubiquitin-proteasome system) |
Summary Diagram: Flow of Genetic Information and Regulation
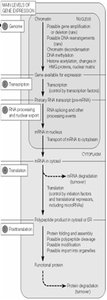
Additional info: This guide covers key mechanisms of gene expression and regulation, including the genetic code, translation, mRNA processing, protein targeting, chromatin structure, transcriptional and post-transcriptional regulation, and the role of small RNAs. Understanding these processes is fundamental for cell biology and molecular genetics.
