 Back
BackGene Expression I: Transcription – Mechanisms and Regulation
Study Guide - Smart Notes

Gene Expression: I. Transcription
Introduction to Gene Expression
Gene expression is the process by which the information encoded in DNA is used to direct the synthesis of RNA and proteins. Transcription is the first step, where DNA serves as a template for RNA synthesis. This process is fundamental to all living cells and is tightly regulated to ensure proper cellular function.
Transcription: Synthesis of RNA from a DNA template.
Translation: Synthesis of proteins using the information in RNA.
Central Dogma: The directional flow of genetic information: DNA → RNA → Protein.
Exceptions: Reverse transcription (RNA → DNA) in retroviruses and RNA replication in some viruses.
18.1 The Directional Flow of Genetic Information
The Central Dogma of Molecular Biology
The central dogma describes the flow of genetic information from DNA to RNA to protein. While this is the general rule, exceptions exist, such as reverse transcription in retroviruses and RNA-dependent RNA synthesis in some viruses. However, information does not flow from protein back to nucleic acids.
mRNA (messenger RNA): Carries genetic information from DNA to ribosomes for protein synthesis.
rRNA (ribosomal RNA): Structural and catalytic component of ribosomes.
tRNA (transfer RNA): Brings amino acids to the ribosome during translation.
Non-coding RNAs: RNAs that are not translated into proteins (e.g., tRNA, rRNA).
Transcription and Translation in Prokaryotes vs. Eukaryotes
In prokaryotes, transcription and translation are coupled due to the absence of a nuclear envelope. In eukaryotes, these processes are separated by the nuclear membrane, allowing for additional regulation and RNA processing.
Prokaryotes: Transcription and translation occur simultaneously in the cytoplasm.
Eukaryotes: Transcription occurs in the nucleus; translation occurs in the cytoplasm.
18.2 Mechanisms of Transcription
Stages of Transcription
Transcription can be divided into four main stages: binding, initiation, elongation, and termination. Each stage involves specific molecular interactions and regulatory mechanisms.
Binding: RNA polymerase binds to the promoter region of DNA, causing local unwinding.
Initiation: RNA synthesis begins using one DNA strand as a template.
Elongation: RNA polymerase moves along the DNA, synthesizing RNA in the 5' to 3' direction.
Termination: RNA synthesis ends when the polymerase encounters a termination signal.
Promoters and RNA Polymerase in Prokaryotes
Promoters are specific DNA sequences that define where transcription starts. In bacteria, the promoter contains conserved sequences at -10 (Pribnow box/TATA box) and -35 positions. RNA polymerase, with the help of the sigma (σ) factor, recognizes these sequences and initiates transcription.
Core enzyme: Catalyzes RNA synthesis but lacks specificity for promoters.
Holoenzyme: Includes the sigma factor, which directs the polymerase to specific promoters.
Elongation and Proofreading
During elongation, RNA polymerase unwinds the DNA ahead and rewinds it behind. Proofreading mechanisms exist but are less stringent than in DNA replication, as errors in RNA are less detrimental.
Proofreading: RNA polymerase can backtrack and remove incorrect nucleotides (RNA backtracking).
Termination Mechanisms
Termination of transcription in bacteria can occur via two mechanisms:
Rho-independent: Formation of a GC-rich hairpin loop followed by a series of U's causes dissociation.
Rho-dependent: Rho protein binds to the RNA and unwinds it from the DNA template.
Transcription in Eukaryotes
Complexity of Eukaryotic Transcription
Eukaryotic transcription is more complex due to the presence of three RNA polymerases (I, II, III), diverse promoter structures, and extensive RNA processing. Transcription factors and chromatin structure play crucial roles in regulation.
RNA Polymerase I: Synthesizes most rRNA.
RNA Polymerase II: Synthesizes mRNA and some snRNA, miRNA.
RNA Polymerase III: Synthesizes tRNA, 5S rRNA, and other small RNAs.
Promoters and Control Elements
Eukaryotic promoters are more varied and can include core promoters (e.g., TATA box, Inr, BRE, DPE) and upstream control elements (e.g., CAAT box, GC box, enhancers). Enhancers can be located far from the gene and interact with promoters via DNA looping, facilitated by activator proteins.
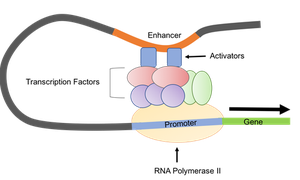
General Transcription Factors (GTFs): Required for RNA polymerase binding and initiation.
TFIID: Recognizes the TATA box via the TATA-binding protein (TBP) subunit.
TFIIH: Unwinds DNA and phosphorylates the C-terminal domain (CTD) of RNA polymerase II.
Chromatin Structure and Transcription Regulation
DNA is packaged with histone proteins, forming chromatin. Modifications such as acetylation (by HATs) and deacetylation (by HDACs) of histones regulate the accessibility of DNA to the transcription machinery. Acetylation loosens chromatin structure, promoting transcription, while deacetylation tightens it, repressing transcription.
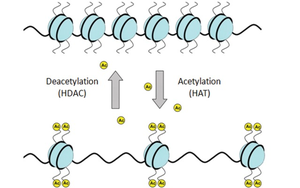
HATs (Histone Acetyltransferases): Add acetyl groups, activating transcription.
HDACs (Histone Deacetylases): Remove acetyl groups, repressing transcription.
18.3 RNA Processing and Turnover
RNA Processing in Eukaryotes
Primary transcripts (pre-mRNA) undergo several modifications before becoming mature mRNA capable of being translated. These modifications are essential for mRNA stability, export, and translation.
5' Capping: Addition of a methylated guanosine cap to the 5' end, protecting mRNA and aiding in ribosome binding.
3' Polyadenylation: Addition of a poly(A) tail to the 3' end, enhancing stability and export.
Splicing: Removal of non-coding introns and joining of exons by the spliceosome.
5' Cap and Poly(A) Tail Functions
The 5' cap and poly(A) tail protect mRNA from degradation and are involved in translation initiation. Poly(A) binding proteins (PABPs) interact with the tail and translation initiation factors, forming a closed-loop structure that enhances translation efficiency.
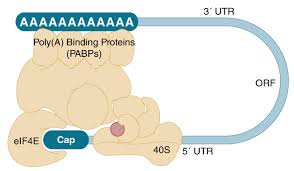
Cap: Recognized by translation initiation factors (eIF4E).
Poly(A) tail: Binds PABPs, stabilizing mRNA and facilitating translation.
Splicing and Alternative Splicing
Introns are removed from pre-mRNA by spliceosomes, which are complexes of snRNPs and proteins. Alternative splicing allows a single gene to produce multiple protein isoforms, increasing proteomic diversity.
Splice sites: Typically defined by GU at the 5' end and AG at the 3' end of introns.
Branch point: An A residue near the 3' end of the intron is essential for lariat formation during splicing.
Alternative splicing: Regulated by splicing enhancers/silencers and associated proteins.
mRNA Export and Quality Control
Only fully processed, mature mRNAs are exported from the nucleus via the nuclear pore complex. Quality control mechanisms ensure that only properly spliced and modified mRNAs reach the cytoplasm.
Nuclear Export Factor 1 (NXF1): Interacts with the cap-binding complex and exon junction complex to mediate export.
mRNA Turnover and Stability
mRNA molecules have variable half-lives, influencing gene expression levels. In eukaryotes, mRNA stability is regulated by the length of the poly(A) tail and the presence of the 5' cap. Degradation can occur from either end by RNases.
Prokaryotic mRNA: Short half-life (minutes).
Eukaryotic mRNA: Longer half-life (hours to days).
Epigenetic Regulation and mRNA Imprinting
Epigenetic mechanisms, such as histone modifications and mRNA imprinting, regulate gene expression without altering the DNA sequence. mRNA imprinting involves marking mRNAs to influence their stability and translation, potentially affecting development and disease.
Epigenetic regulation: Includes DNA methylation, histone modification, and mRNA imprinting.
mRNA imprinting: Co-transcriptional modification of mRNA, influencing gene expression in response to environmental factors.
Summary Table: Key Differences in Transcription and mRNA Processing
Feature | Prokaryotes | Eukaryotes |
|---|---|---|
RNA Polymerases | One type | Three types (I, II, III) |
Promoter Structure | Simple, conserved (-10, -35) | Complex, varied (TATA, Inr, etc.) |
Transcription Factors | Sigma factor | Multiple GTFs, regulatory proteins |
RNA Processing | Minimal | 5' cap, poly(A) tail, splicing |
mRNA Stability | Short half-life | Longer half-life |
Key Terms and Concepts
Transcription unit: DNA sequence transcribed into a single RNA molecule.
Promoter: DNA sequence where RNA polymerase binds to initiate transcription.
Enhancer: Regulatory DNA sequence that increases transcription efficiency.
Spliceosome: Complex responsible for removing introns from pre-mRNA.
Alternative splicing: Process by which different combinations of exons are joined to produce multiple mRNA variants from a single gene.
Epigenetics: Heritable changes in gene expression not involving changes to the DNA sequence.
Additional info: This summary integrates foundational concepts and mechanisms of transcription and RNA processing, with emphasis on regulatory complexity in eukaryotes and the importance of post-transcriptional modifications for gene expression control.
