 Back
BackGene Expression II: The Genetic Code and Protein Synthesis (Translation)
Study Guide - Smart Notes

Gene Expression II: The Genetic Code and Protein Synthesis (Translation)
Introduction to Translation
Translation is the process by which the genetic information encoded in messenger RNA (mRNA) is used to assemble a specific sequence of amino acids, forming a polypeptide chain. This process is central to gene expression and is carried out by a coordinated set of molecular components.
mRNA: Contains codons that specify the amino acid sequence of the protein.
tRNA: Adaptor molecules that match amino acids to codons in mRNA via their anticodon loop.
Aminoacyl tRNA Synthetases (AASs): Enzymes that attach the correct amino acid to its corresponding tRNA.
Ribosome: The molecular machine that catalyzes peptide bond formation and ensures the correct reading of the mRNA.
The Genetic Code
Structure and Properties of the Genetic Code
The genetic code is a set of rules that defines how the sequence of nucleotides in mRNA is translated into the sequence of amino acids in a protein. It is nearly universal among organisms.
Triplet Code: Each amino acid is specified by a sequence of three nucleotides (codon).
Degeneracy: Most amino acids are encoded by more than one codon.
Nonoverlapping: Codons are read one after another, without overlap.
Unambiguous: Each codon specifies only one amino acid.
Start and Stop Codons: AUG is the start codon; UAA, UAG, and UGA are stop codons.
Example: The codon AUG codes for methionine and also serves as the start signal for translation.
Reading Frames
The reading frame determines how the nucleotide sequence is divided into codons. Shifting the reading frame changes the resulting amino acid sequence.

Codon Usage Bias
Codon usage bias refers to the preference for certain codons over others that encode the same amino acid. This bias varies among species and can affect gene expression efficiency.
Codon | Human | Drosophila | E. coli |
|---|---|---|---|
AGA | 22% | 10% | 1% |
AGG | 23% | 8% | 1% |
CGA | 10% | 8% | 4% |
CGC | 22% | 49% | 43% |
CGG | 14% | 8% | 2% |
CGU | 9% | 18% | 49% |
Total number of arginine codons | 2403 | 506 | 149 |
Total number of genes | 195 | 46 | 149 |
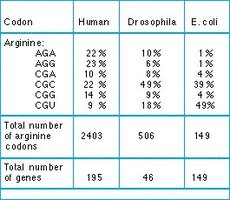
Additional info: Codon usage bias can influence the efficiency of translation and is important in biotechnology applications, such as optimizing gene expression in heterologous systems.
Translation: The Cast of Characters
tRNA Structure and Function
Transfer RNAs (tRNAs) are adaptor molecules that bring amino acids to the ribosome and match them to the codons in mRNA via their anticodon loop. The third position of the codon (wobble position) allows for some flexibility in base pairing, enabling one tRNA to recognize multiple codons.
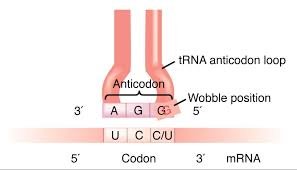
Messenger RNA (mRNA) Structure
mRNA molecules contain untranslated regions (UTRs) at both the 5' and 3' ends, a coding region, and modifications such as a 5' cap and a 3' poly(A) tail in eukaryotes. These features are essential for mRNA stability, translation initiation, and regulation.
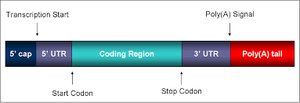
Monocistronic vs. Polycistronic mRNA
Most eukaryotic mRNAs are monocistronic, encoding a single polypeptide. In contrast, many prokaryotic mRNAs are polycistronic, encoding multiple polypeptides from a single transcript (operons).
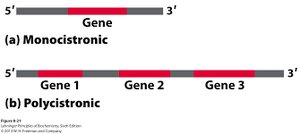
The Mechanism of Translation
Stages of Translation
Translation occurs in three main stages:
Initiation: Assembly of the translation machinery at the start codon.
Elongation: Sequential addition of amino acids to the growing polypeptide chain.
Termination: Release of the completed polypeptide upon encountering a stop codon.
Initiation Complex Formation
During initiation, initiation factors, the small ribosomal subunit, initiator tRNA, and mRNA assemble to form the initiation complex. In eukaryotes, the 5' cap and poly(A) tail of mRNA are important for efficient initiation.
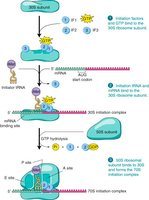
Elongation and Peptide Bond Formation
Elongation involves the binding of aminoacyl-tRNA to the A site, peptide bond formation catalyzed by rRNA (a ribozyme), and translocation of the ribosome along the mRNA. The process is highly accurate and energy-dependent.
Termination
When a stop codon enters the A site, release factors bind and trigger the release of the polypeptide and disassembly of the translation complex.
Mutations and Translation
Types of Mutations
Missense Mutation: A codon change results in a different amino acid.
Nonsense Mutation: A codon change creates a premature stop codon.
Silent Mutation: A codon change does not alter the amino acid (often due to degeneracy).
Nonstop Mutation: A stop codon is mutated to an amino acid codon, resulting in a longer protein.
Posttranslational Processing
Types of Posttranslational Modifications
After translation, polypeptides often undergo modifications that are essential for their function. These include cleavage of signal sequences, formation of disulfide bonds, and addition of chemical groups such as phosphate, methyl, acetyl, or ubiquitin.
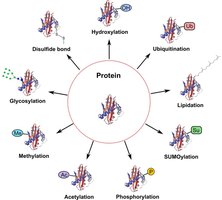
Proteolytic Cleavage: Removal of specific segments to activate the protein (e.g., insulin maturation).
Chemical Modifications: Phosphorylation, methylation, acetylation, glycosylation, ubiquitination, etc.
Protein Splicing: Removal of inteins and joining of exteins.
Sample Questions
Which of the following is the correct order of the main steps in translation? Answer: Initiation → Elongation → Termination
The genetic code is described as "degenerate." What does this mean? Answer: Multiple codons can code for the same amino acid.
What is the role of tRNA in translation? Answer: To carry amino acids to the ribosome and match them to the mRNA codon.
A point mutation changes a codon from UUU (phenylalanine) to UGA (a stop codon). What type of mutation is this? Answer: Nonsense mutation.
What happens during the termination step of translation? Answer: A stop codon is recognized by release factors, leading to disassembly of the ribosome.
