 Back
BackGene Expression: Mechanisms and Regulation in Eukaryotes and Prokaryotes
Study Guide - Smart Notes

Gene Expression
Overview of Gene Expression
Gene expression is the process by which information encoded in a gene is used to direct the synthesis of a functional gene product, typically a protein or functional RNA. This process involves transcription (copying DNA into RNA) and, in eukaryotes, extensive RNA processing before translation.
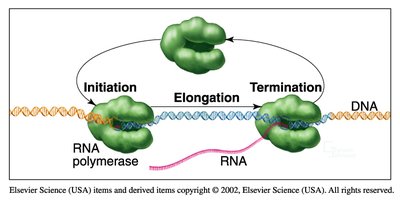
Transcription is catalyzed by RNA polymerases, which synthesize RNA from a DNA template.
Gene expression is tightly regulated at multiple steps, especially at the initiation of transcription.
I. Transcription Units
Definition and Organization
A gene is a segment of DNA that encodes a functional RNA or protein, including adjacent regulatory sequences. The transcription unit encompasses both regulatory and coding sequences.
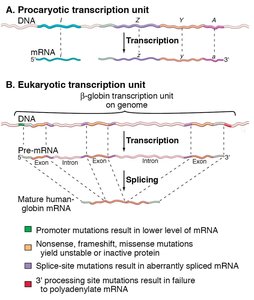
In prokaryotes, multiple genes may be transcribed together as an operon, producing a single mRNA that can encode several proteins.
In eukaryotes, each gene typically has its own transcription unit, and the initial RNA transcript (pre-mRNA) undergoes processing (e.g., splicing) to become mature mRNA.
II. Classes of RNA
Types and Functions
Cells produce several classes of RNA, each with distinct roles:
Ribosomal RNA (rRNA): Structural and catalytic components of ribosomes; most abundant RNA type.
Transfer RNA (tRNA): Adaptor molecules in translation, bringing amino acids to the ribosome.
Messenger RNA (mRNA): Encodes protein sequences; processed from pre-mRNA in eukaryotes.
Small nuclear RNAs (snRNAs): Involved in splicing and RNA processing.
Noncoding RNAs (ncRNAs): Regulatory RNAs, including microRNAs (miRNAs), which modulate gene expression.
III. RNA Polymerases
Structure and Specificity
Eukaryotic cells have three main RNA polymerases, each responsible for transcribing different classes of genes:
RNA polymerase I: Synthesizes most rRNAs.
RNA polymerase II: Synthesizes mRNAs and some snRNAs.
RNA polymerase III: Synthesizes tRNAs, 5S rRNA, and other small RNAs.
All RNA polymerases are multi-subunit enzymes related to DNA polymerases but do not require a primer to initiate synthesis. 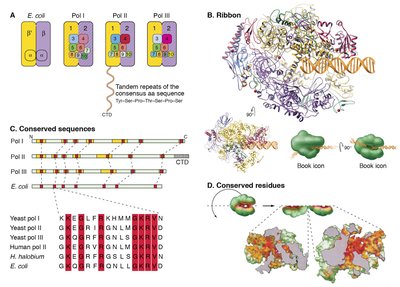
IV. DNA Elements That Control Transcription Initiation
Regulatory DNA Sequences
Transcription initiation is the primary regulatory step in gene expression. Regulation is achieved through:
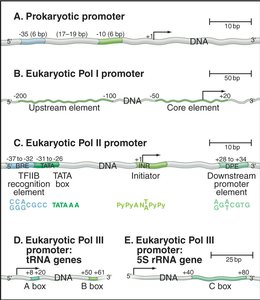
Promoters: DNA sequences immediately upstream of the transcription start site that recruit RNA polymerase and general transcription factors (GTFs). Many contain a TATA box recognized by the TATA-binding protein (TBP).
Promoter proximal elements: Short DNA sequences near the promoter that bind specific transcription factors to enhance initiation.
Enhancers: Distant regulatory elements that can increase transcription from a promoter, regardless of orientation or distance.
V. Initiation of Transcription
Assembly of the Preinitiation Complex
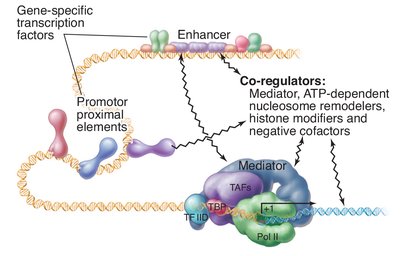
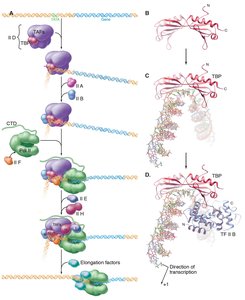
Initiation involves the assembly of a multiprotein complex at the promoter:
TBP binds the TATA box, bending the DNA.
General transcription factors and mediator complex assemble, recruiting RNA polymerase II.
Helicases use ATP to unwind DNA, allowing RNA polymerase to access the template strand and begin RNA synthesis.
VI. Transcription Elongation, Pausing, and Termination
Mechanisms and Regulation
Elongation: RNA polymerase synthesizes RNA at 50–100 nucleotides per second, using ATP, GTP, UTP, and CTP as substrates. The C-terminal domain (CTD) of Pol II is phosphorylated to allow promoter escape and processive elongation.
Pausing and Editing: Polymerase may pause or backtrack for error correction.
Termination: Specific DNA sequences or protein factors trigger dissociation of the polymerase and release of the RNA transcript. Mechanisms differ among Pol I, II, and III.
VII. Families of Transcription Factors
DNA-Binding Domains and Specificity
Transcription factors are grouped into families based on their DNA-binding domains:
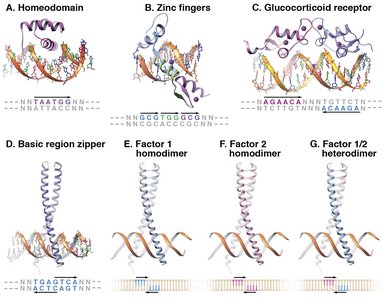
Helix-Turn-Helix (Homeodomain): Recognizes specific DNA sequences via an α-helix in the major groove.
Zinc Finger: Multiple small domains stabilized by zinc ions, each recognizing a short DNA sequence.
Leucine Zipper: Coiled-coil domains that dimerize and bind DNA as homo- or heterodimers.
Steroid Receptors: Homodimers that bind palindromic DNA sequences in response to hormone binding.
VIII. Experimental Identification of DNA Regulatory Elements and Transcription Factors
Key Techniques
Promoter Bashing: Systematic deletion of regulatory DNA to identify sequences required for gene expression.
Chromatin Immunoprecipitation (ChIP): Crosslinking proteins to DNA, immunoprecipitating with specific antibodies, and sequencing the associated DNA to map protein-DNA interactions genome-wide.
IX. Gene-Specific Regulation of Transcription
Combinatorial Control and Expression Profiling
Each gene is regulated by a unique combination of transcription factors binding to its promoter, proximal elements, and enhancers.
Gene expression can be measured using RNA sequencing (RNAseq), which quantifies mRNA levels genome-wide.
Transcriptional repression can occur via competition for DNA binding or chromatin modification.
X. Influence of Chromatin Structure on Transcription
Histone Modifications and Nucleosome Remodeling
Chromatin structure determines the accessibility of DNA to transcription factors:
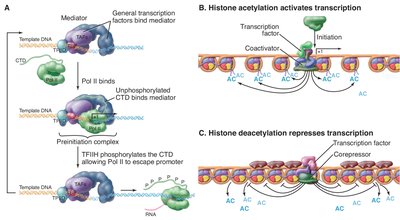
Histone acetylation by acetyltransferases opens chromatin, facilitating transcription.
Histone deacetylation by deacetylases compacts chromatin, repressing transcription.
Nucleosome remodeling complexes use ATP to reposition nucleosomes, aiding transcription factor access and elongation.
XI. Regulation of Gene Expression by Signaling Pathways
Signal-Dependent Transcriptional Control
Gene expression is modulated by intracellular and extracellular signals:
Steroid hormones activate transcription factors by releasing them from cytoplasmic sequestration.
Growth factors and other signals activate protein kinases, which phosphorylate transcription factors, altering their activity and nuclear localization.
Gene regulatory circuits enable complex, cell-type-specific expression patterns, often studied using single-cell RNA sequencing.
XII. Genome Editing Tools
Targeted Modification of DNA
Gene editing technologies introduce site-specific double-strand breaks (DSBs) in DNA, which are repaired by the cell:
Zinc finger nucleases: Engineered proteins that recognize specific DNA sequences.
CRISPR/Cas9: Uses a guide RNA to direct the Cas9 nuclease to a specific DNA sequence for cleavage.
XIII. Epigenetic Modifications and Nucleosome Position
Epigenetic Regulation of Transcription
Histone acetylation and methylation are coordinated with RNA polymerase II recruitment and chromatin compaction.
Chromatin remodeling complexes are recruited by epigenetic marks, influencing gene accessibility and expression.
Key Terms and Concepts
Chromatin Immunoprecipitation (ChIP)
Coactivator/Corepressor
C-terminal domain (CTD)
Enhancer
General Transcription Factors (GTFs)
Initiation, Elongation, Termination
Promoter, Promoter Proximal Elements
RNA Polymerase I, II, III
TATA Box, TBP
Transcription Factor
Noncoding RNA (ncRNA)
Sample Table: Comparison of RNA Polymerases
Polymerase | Main Products | Promoter Elements |
|---|---|---|
RNA Pol I | rRNA (except 5S) | Upstream element, core element |
RNA Pol II | mRNA, some snRNA | TATA box, initiator, downstream elements |
RNA Pol III | tRNA, 5S rRNA, 7S RNA | A box, B box, C box |
Key Equations
RNA synthesis reaction:
Phosphorylation of Pol II CTD:
