 Back
BackGene Regulation in Prokaryotes and Eukaryotes: Operons, Regulatory Mechanisms, and Molecular Biology Techniques
Study Guide - Smart Notes

Gene Regulation: Overview
Types of Gene Regulation
Gene regulation is essential for cellular function, allowing cells to respond to environmental changes and developmental cues. Regulation can be classified as negative or positive, depending on whether gene expression is turned off or on by regulatory proteins.
Negative Regulation: Genes are ON unless a regulatory protein binds to turn them OFF. Common in bacteria.
Positive Regulation: Genes are OFF until a regulatory protein turns them ON. Occurs in eukaryotes.

Prokaryotic Gene Regulation: The Operon Model
Operon Structure and Function
An operon is a cluster of genes under the control of a single promoter and regulatory elements, allowing coordinated expression. Operons are common in prokaryotes and some organelles (e.g., mitochondria).
Components: Promoter, operator (controlling site), regulatory gene (repressor), coding sequences, terminator.
Inducers: Molecules that initiate transcription by inactivating repressors.
Induction: Synthesis of gene products in response to an inducer.
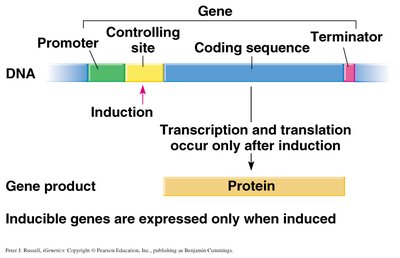
Lac Operon: Inducible System in E. coli
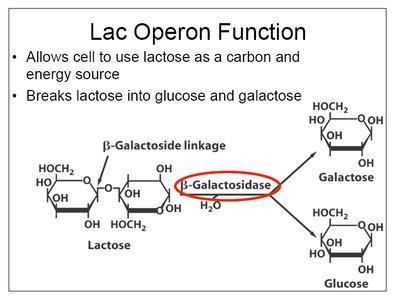
The lac operon enables E. coli to metabolize lactose when glucose is absent. It is a classic example of negative and positive regulation.
Structural genes: lacZ (β-galactosidase), lacY (permease), lacA (transacetylase).
Regulatory genes: lacI (repressor), operator, promoter.
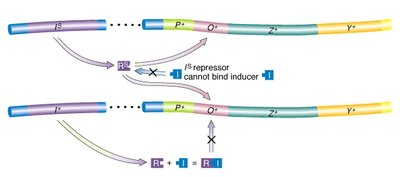
Inducer: Allolactose (a lactose isomer) binds the repressor, inactivating it and allowing transcription.
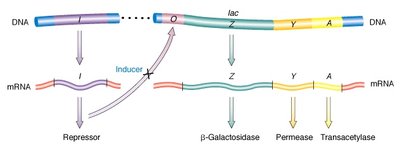
Mutational Analysis of the Lac Operon
Mutations in the lac operon can affect gene expression in different ways:
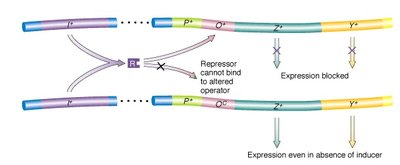
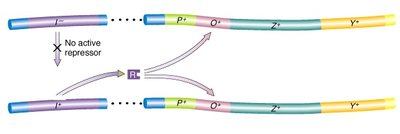
Operator mutations (lacO): Prevent repressor binding, causing constitutive expression of downstream genes.
Repressor mutations (lacI): Inactivate the repressor, also leading to constitutive expression.
Promoter mutations (Plac): Prevent RNA polymerase binding, blocking expression of all structural genes.
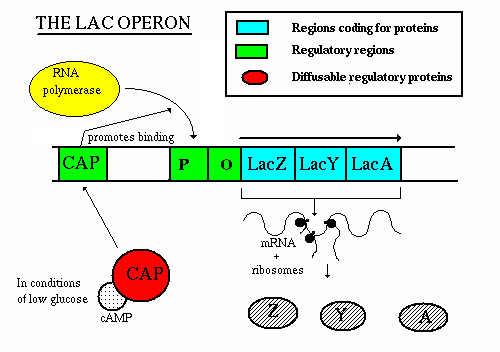
Positive Control: Catabolite Activation
When glucose is scarce, cAMP levels rise, and the cAMP-CAP complex binds upstream of the lac promoter, enhancing RNA polymerase binding and transcription. This is an example of positive regulation.
CAP (catabolite activator protein): Binds cAMP and DNA, promoting transcription in low glucose conditions.
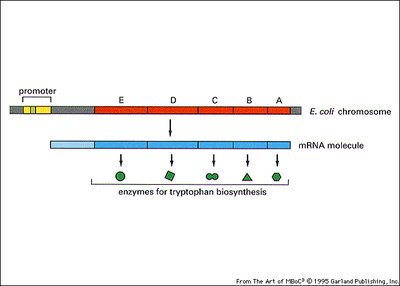
Trp Operon: Repressible System in E. coli
The trp operon encodes enzymes for tryptophan biosynthesis. It is regulated by a repressor that is activated by tryptophan (the co-repressor).
Negative regulation: In the presence of tryptophan, the repressor binds the operator, blocking transcription.
In the absence of tryptophan: The repressor is inactive, and transcription proceeds.
Eukaryotic Gene Regulation
Levels of Regulation
Eukaryotic gene regulation is more complex due to multicellularity and developmental requirements. Regulation occurs at multiple levels:
Pre-transcriptional: DNA methylation, histone modification, chromatin structure.
Transcriptional: Transcription factors, enhancers, silencers, response elements.
Translational: mRNA stability, alternative splicing, miRNA inhibition.
Post-translational: Protein modification, subunit assembly, degradation.
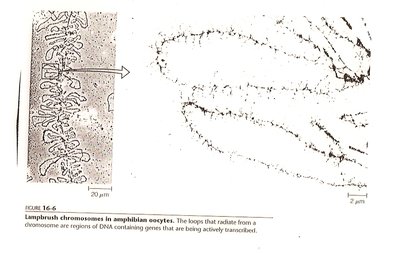
Pre-Transcriptional Regulation
DNA methylation: Addition of methyl groups to CpG islands silences genes.
Histone modification: Acetylation activates, deacetylation represses transcription.
Chromatin remodeling: Alters DNA accessibility for transcription machinery.
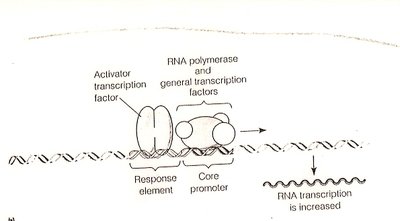
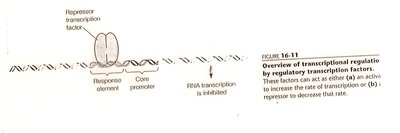
Transcriptional Regulation
Transcription factors: Proteins that bind DNA and regulate RNA polymerase activity.
Enhancers and silencers: DNA elements that increase or decrease transcription rates.
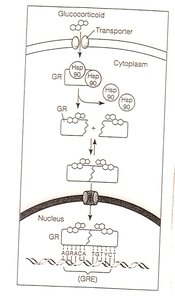
Response elements: Specific DNA sequences recognized by regulatory proteins.
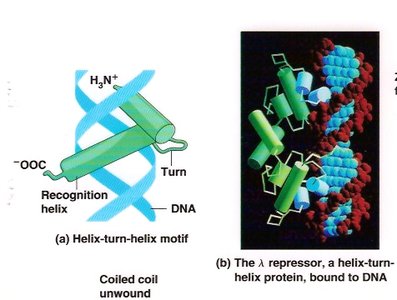
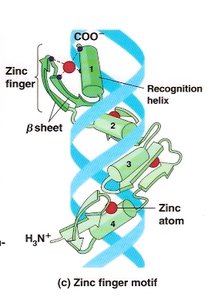
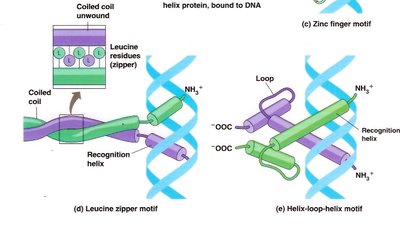
Transcription Factor Motifs
Helix-turn-helix: Common DNA-binding motif in prokaryotic repressors.
Zinc finger: DNA-binding motif stabilized by zinc ions.
Leucine zipper and helix-loop-helix: Motifs that mediate dimerization and DNA binding in eukaryotic transcription factors.
Translational and Post-Translational Regulation
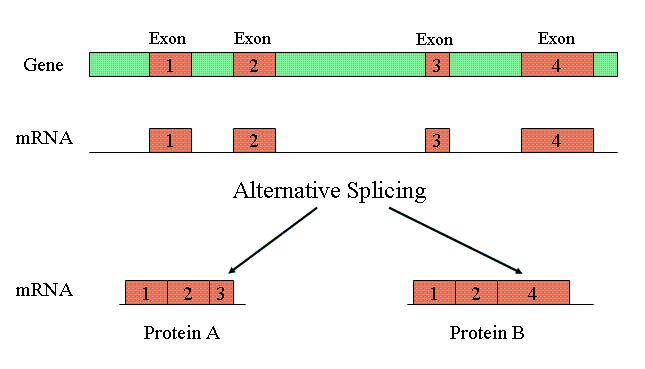
Alternative splicing: Generates multiple proteins from one gene.
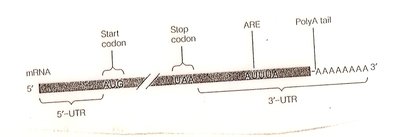
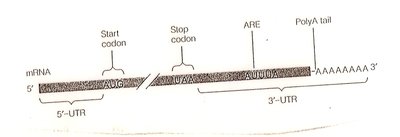
mRNA stability: AU-rich elements (AREs) and poly-A tail length affect mRNA degradation.
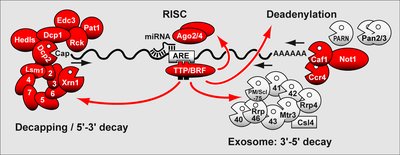
miRNA inhibition: MicroRNAs bind mRNA, blocking translation or promoting degradation.
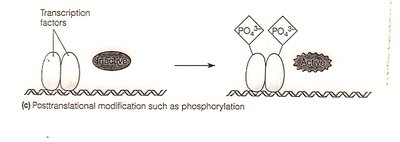
Protein modification: Phosphorylation, methylation, and other modifications alter protein function.
Molecular Biology Techniques for Studying Gene Expression
PCR and RT-PCR
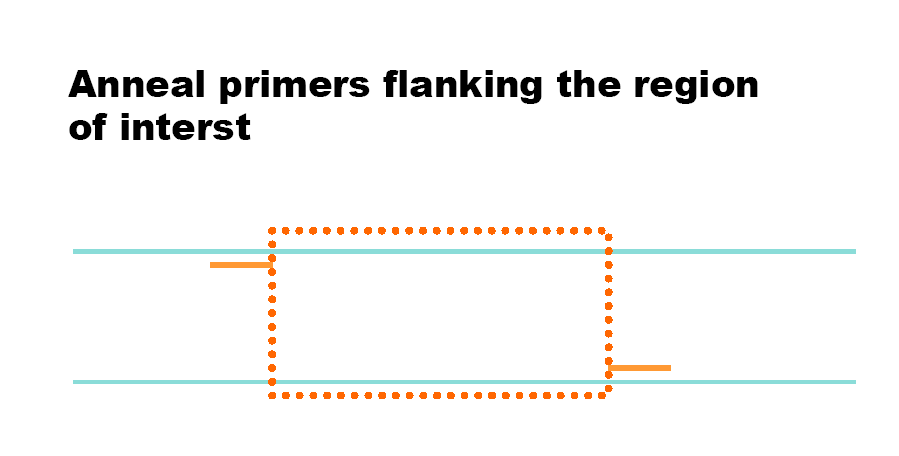
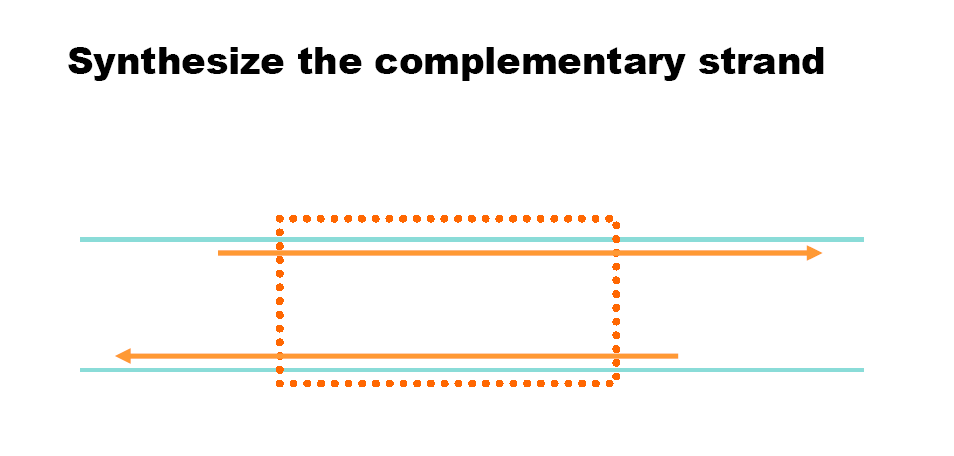
Polymerase Chain Reaction (PCR) is a technique to amplify DNA. Reverse Transcriptase PCR (RT-PCR) uses mRNA as a template to generate cDNA, allowing analysis of gene expression.
PCR steps: Denaturation (~94°C), annealing (~55°C), extension (~72°C).
RT-PCR: Converts mRNA to cDNA, then amplifies cDNA.
qRT-PCR: Quantifies mRNA levels in real time.
Other Techniques
Northern blot: Detects specific RNA molecules.
Western blot: Detects specific proteins.
DNA microarrays: Analyze expression of thousands of genes simultaneously.
Summary Table: Key Differences in Gene Regulation
Feature | Prokaryotes | Eukaryotes |
|---|---|---|
Organization | Operons (polycistronic) | Monocistronic, complex regulation |
Regulation Level | Mainly transcriptional | Multiple (epigenetic, transcriptional, post-transcriptional, translational, post-translational) |
Regulatory Proteins | Repressors, activators | Transcription factors, coactivators, repressors |
Examples | Lac operon, trp operon | Alternative splicing, miRNA regulation |
