 Back
BackGene Regulation in Prokaryotes and Eukaryotes: The Lac Operon and Epigenetic Mechanisms
Study Guide - Smart Notes

Gene Regulation in Bacteria: The Lac Operon
Introduction to Operons
Operons are a hallmark of prokaryotic gene regulation, allowing coordinated expression of genes with related functions. The lac operon is a classic example, providing insight into inducible gene regulation in Escherichia coli.
Operon: A cluster of genes under the control of a single promoter and operator, transcribed as a single polycistronic mRNA.
Advantage: Enables simultaneous regulation of multiple functionally related genes, optimizing resource use.
Polycistronic mRNA: A single mRNA molecule that encodes multiple proteins, each with its own start and stop codon.
Note: Operons are rare in eukaryotes.
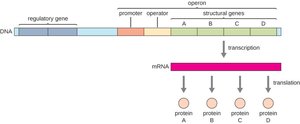
Types of Operon Regulation
Operons can be regulated by two main mechanisms: repressible and inducible systems.
Repressible operons: Typically control anabolic pathways (e.g., trp operon); default state is "on" but can be turned "off" by a repressor when the end product is abundant.
Inducible operons: Typically control catabolic pathways (e.g., lac operon); default state is "off" but can be turned "on" in the presence of a substrate.
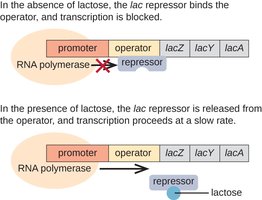
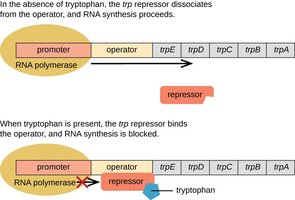
The Lac Operon: Structure and Function
The lac operon consists of three structural genes (lacZ, lacY, lacA) involved in lactose metabolism, regulated by a single promoter and operator.
lacZ: Encodes β-galactosidase, which cleaves lactose into glucose and galactose.
lacY: Encodes lactose permease, which transports lactose into the cell.
lacA: Encodes thiogalactoside transacetylase (function less central to lactose metabolism).
lacI: Encodes the lac repressor protein, which regulates the operon but is not part of it.
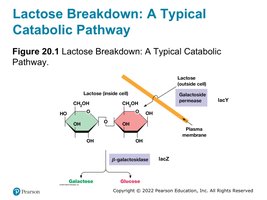
Negative Regulation by the Lac Repressor
The lac operon is negatively regulated by the lac repressor, a protein encoded by the lacI gene. In the absence of lactose, the repressor binds the operator, blocking RNA polymerase and preventing transcription. When lactose (specifically allolactose) is present, it binds the repressor, causing a conformational change that releases the repressor from the operator, allowing transcription.
Allosteric regulation: The repressor protein changes shape upon binding the inducer (allolactose).
Polycistronic mRNA: One transcript encodes all three proteins.

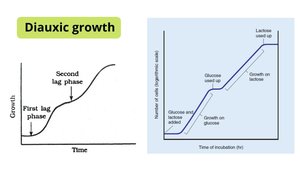
Diauxic Growth and Catabolite Repression
When both glucose and lactose are present, E. coli preferentially uses glucose. Only after glucose is depleted does lactose utilization begin, resulting in a biphasic (diauxic) growth curve. This is due to catabolite repression, where glucose inhibits the lac operon.
Mutations Affecting the Lac Operon
lacZ- or lacY-: Defective enzymes, unable to metabolize lactose.
Operator-constitutive (Oc): Mutation in operator prevents repressor binding, causing constitutive expression.
Promoter mutations (P-): Decrease RNA polymerase binding, reducing transcription.
lacI-: Nonfunctional repressor, operon always "on" (constitutive expression).
lacIs (superrepressor): Repressor cannot bind inducer, operon always "off" (gain-of-function mutation).
Gene Regulation in Eukaryotes: Epigenetic Mechanisms
Introduction to Epigenetics
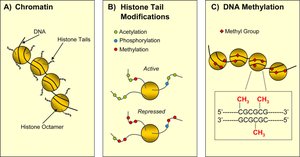
Epigenetic regulation refers to heritable changes in gene expression that do not involve changes to the DNA sequence. In eukaryotes, chromatin structure and chemical modifications to DNA and histones play central roles in regulating gene accessibility and transcription.
Chromatin: DNA packaged with histone proteins into nucleosomes.
Epigenetic modifications: Include DNA methylation and histone modifications (acetylation, methylation, phosphorylation, ubiquitination).
Histone Modifications and Chromatin Remodeling
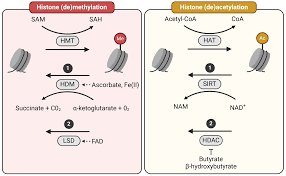
Histone modifications alter chromatin structure, influencing gene expression. Acetylation of histones generally promotes transcription by loosening chromatin, while methylation can either activate or repress genes depending on the context.
HAT (Histone Acetyltransferase): Adds acetyl groups, promoting open chromatin and active transcription.
HDAC (Histone Deacetylase): Removes acetyl groups, leading to condensed chromatin and gene repression.
HMT (Histone Methyltransferase): Adds methyl groups, often leading to gene silencing.
HDM (Histone Demethylase): Removes methyl groups.
Chromatin remodeling complexes (e.g., SWI/SNF): Use ATP to reposition nucleosomes, regulating DNA accessibility.

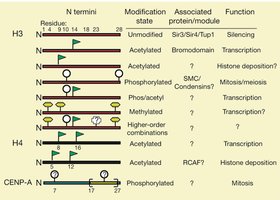
The Histone Code Hypothesis
The "histone code" hypothesis proposes that specific combinations of histone modifications constitute a code that is read by other proteins to regulate chromatin structure and gene expression.
Different modifications (acetylation, methylation, phosphorylation, ubiquitination) on histone tails can recruit specific proteins that activate or repress transcription.
DNA Methylation
DNA methylation involves the addition of methyl groups to cytosine bases, typically at CpG dinucleotides. This modification is associated with transcriptional repression and is a key mechanism of long-term gene silencing in eukaryotes.
CpG islands: Regions rich in CpG sites, often found near gene promoters; methylation here strongly represses transcription.
MeCP2: Protein that binds methylated DNA and recruits histone-modifying enzymes, further repressing gene expression.
Epigenetic Inheritance and Imprinting
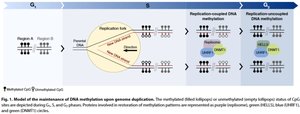
Epigenetic marks such as DNA methylation can be inherited through cell divisions. Genomic imprinting is a phenomenon where certain genes are expressed in a parent-of-origin-specific manner, often regulated by DNA methylation.
Maintenance methylases: Enzymes that copy methylation patterns to daughter strands during DNA replication.
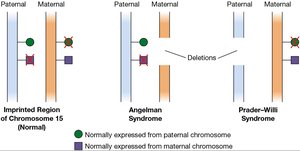
Imprinting syndromes: Disorders such as Prader-Willi and Angelman syndromes result from abnormal imprinting and gene expression on chromosome 15.
X Chromosome Inactivation
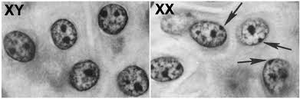
In female mammals, one X chromosome is randomly inactivated in each cell during early development, ensuring dosage compensation between males (XY) and females (XX). The inactive X becomes highly methylated and condensed into a Barr body.
Xist RNA: A long noncoding RNA that coats the inactive X chromosome, recruiting chromatin-modifying proteins to silence it.
Dosage compensation: Ensures equal expression of X-linked genes in males and females.
Mosaicism: Females are mosaics for X-linked gene expression due to random X inactivation.

Transcriptional Control in Eukaryotes
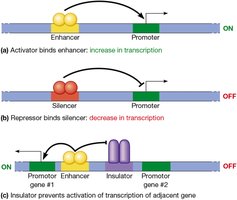
Transcriptional control is a major level of gene regulation in eukaryotes, involving core promoters, proximal control regions, and distant regulatory elements such as enhancers and silencers.
Transcription factors: Proteins that bind specific DNA sequences to regulate transcription.
Enhancers: Distant DNA elements that increase transcription by interacting with activator proteins.
Silencers: DNA elements that decrease transcription by recruiting repressors.
Insulators: DNA sequences that block the influence of enhancers on adjacent genes.
Summary Table: Key Mechanisms of Gene Regulation
Mechanism | Prokaryotes | Eukaryotes |
|---|---|---|
Operons | Common (e.g., lac, trp) | Rare |
Polycistronic mRNA | Yes | No (mostly monocistronic) |
Epigenetic regulation | No | Yes (DNA methylation, histone modification) |
Transcription factors | Simple | Complex, many types |
Enhancers/Silencers | No | Yes |
Practice Questions and Answers
Protein phosphorylation, folding, and proteolytic cleavage are examples of: Post-translational control of gene expression and activity.
XIST in mammals: Produces a noncoding RNA that coats and silences one X chromosome.
Dosage compensation in mammals: Ensures gene expression from the X chromosome is equal between males and females.
Epigenetic modifications associated with X inactivation: Histone methylation and DNA methylation.
X chromosome inactivation in placental mammals: Is random, with either maternal or paternal X silenced.
Histone acetylation and methylation: Both are common in vivo modifications.
Negative control: An active regulatory protein turns off gene expression.
Post-transcriptional control in eukaryotes: Includes alternative mRNA splicing, control of nuclear export, and differential mRNA degradation.
Alternative splicing: Allows a single gene to produce multiple protein products.
Enhancers: Increase transcription by interacting with specific transcription factors.
Chromatin remodeling: Modifies nucleosome positions to allow or restrict DNA access.
Histone acetylation: Enhances gene expression by loosening chromatin structure.
Chromatin remodeling: Is not a mechanism of post-transcriptional regulation.
