 Back
BackIntracellular Compartments and Protein Sorting: The Endomembrane System and ER Protein Trafficking
Study Guide - Smart Notes

Intracellular Compartments and Protein Sorting
The Endomembrane System
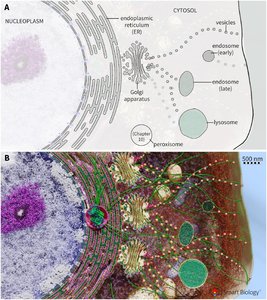
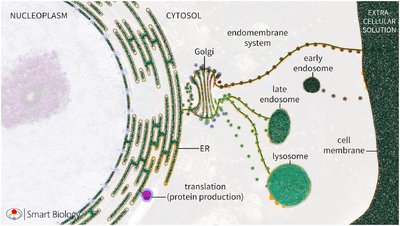
The endomembrane system is a network of membrane-bound organelles that coordinate the synthesis, modification, and transport of proteins and lipids within eukaryotic cells. This system is essential for maintaining cellular organization and function.
Key Organelles:
Endoplasmic Reticulum (ER): Site of protein and lipid synthesis.
Outer Nuclear Membrane: Continuous with the ER.
Golgi Apparatus: Central hub for protein sorting and modification.
Endosomes: Involved in sorting and recycling of internalized material.
Lysosomes: Responsible for degradation of macromolecules.
Peroxisomes: Involved in lipid metabolism and detoxification.
Cell Membrane: Boundary of the cell, involved in transport and signaling.
Environmental Similarities: Organelle lumens share chemical similarities with the extracellular matrix, creating a "non-cytosolic environment" distinct from the cytosol.
Structural and Functional Characteristics of Key Organelles
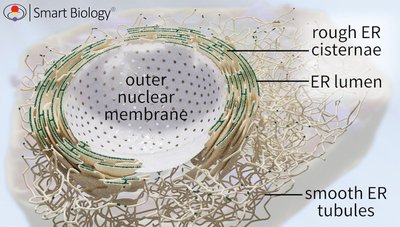
The ER is structurally diverse, consisting of rough and smooth domains, each with specialized functions.
Rough Endoplasmic Reticulum (RER):
Continuous membrane network connected to the nuclear envelope.
"Rough" appearance due to ribosomes bound to its cytosolic surface.
Two interconnected domains:
Cisternae: Flattened, stacked sacs near the nucleus, providing large luminal spaces for protein synthesis and folding.
Tubules: Narrow, branching membranes linking the RER to the Smooth ER (SER) and transitional ER regions.
Smooth Endoplasmic Reticulum (SER):
Lacks ribosomes; consists of interconnected tubules.
Functions in lipid synthesis, detoxification, calcium storage, and carbohydrate metabolism.
Produces membrane lipids essential for vesicle formation and endomembrane maintenance.
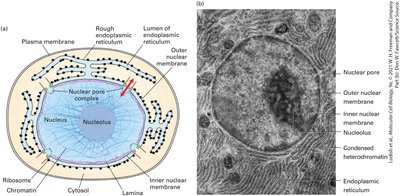
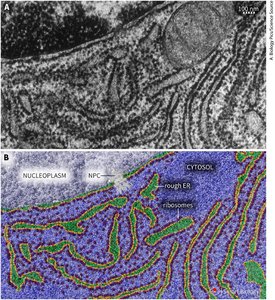
RER Functions and Protein Trafficking
The RER is the entry point for proteins into the endomembrane system, responsible for initial folding, quality control, and early post-translational modifications such as N-linked glycosylation. Properly folded proteins are packaged into vesicles for transport to the Golgi apparatus.
Protein Folding and Quality Control: Chaperones and enzymes ensure proper folding and modification.
Post-Translational Modifications: Includes N-linked glycosylation, disulfide bond formation, and proteolytic cleavage.
Vesicle Formation: Proteins are sorted and packaged for transport to the Golgi.
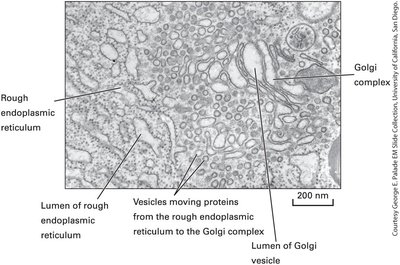
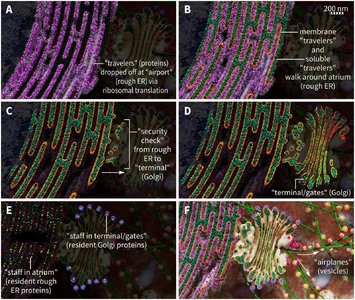
Transitional ER and ER Exit Sites (ERES)
Transitional ER regions are smooth, ribosome-free areas at the junctions between rough and smooth ER, often near the cis face of the Golgi apparatus. These specialized sites are where proteins and lipids exit the ER.
ER Exit Sites: Locations where vesicles bud off to transport proteins and lipids to the Golgi.
Golgi Apparatus: Series of flattened, stacked membrane-bound sacs (cisternae) that serve as the central hub for protein sorting and modification.
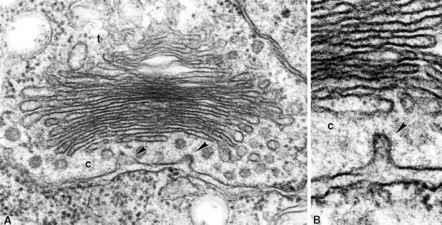
Protein Trafficking into the Endoplasmic Reticulum
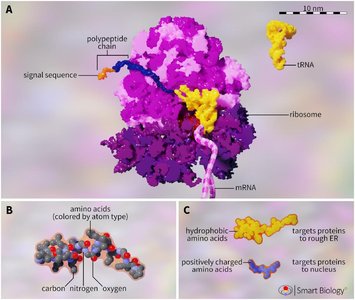
Signal Sequences and Targeting
Proteins destined for the endomembrane system contain specific signal sequences within their amino acid chains, usually at the N-terminus. These sequences direct the protein to its final organelle destination.
Hydrophobic Signal Sequences: Target proteins to the ER.
Positively Charged Signal Sequences: Target proteins to the nucleus.
Translation Initiation: All proteins begin translation on cytosolic free ribosomes; the N-terminus determines the pathway.
Free Ribosomes vs Attached Ribosomes
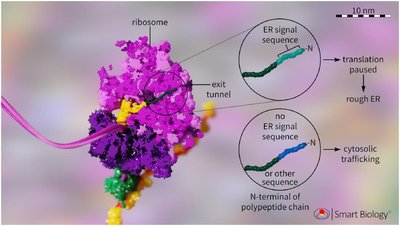
Ribosomes can be free in the cytosol or attached to the ER. The presence of an ER signal sequence causes ribosomes to attach to the ER membrane, initiating the endomembrane pathway.
Free Ribosomes: Synthesize proteins destined for the cytosol or organelles such as mitochondria and peroxisomes.
Attached Ribosomes: Synthesize proteins destined for secretion, membrane insertion, or organelle lumens.
Signal Recognition Particle (SRP) and Cotranslational Translocation
The Signal Recognition Particle (SRP) is a ribonucleoprotein that recognizes the ER signal sequence and pauses translation. The SRP-ribosome complex is then targeted to the ER membrane, where translation resumes and the polypeptide is translocated into the ER lumen.
SRP Functions:
Binds to ER signal sequence.
Pauses translation.
Targets ribosome to SRP receptor on ER membrane.
Cotranslational Translocation: Polypeptide is synthesized and translocated into the ER lumen simultaneously.
Signal Peptidase: Cleaves off ER signal sequence once translocation is complete.
Synthesis of Soluble and Integral Membrane Proteins
Proteins synthesized in the ER can be soluble (released into the lumen) or integral membrane proteins (embedded in the membrane). Internal hydrophobic signal-anchor sequences act as both ER targeting signals and the first transmembrane domain, and are not cleaved.
Soluble Proteins: Translocated into the ER lumen; signal sequence is cleaved.
Integral Membrane Proteins: Inserted into the ER membrane; internal signal-anchor sequences establish membrane topology.
Processing of Newly Synthesized Proteins in the Endoplasmic Reticulum
Proteolytic Cleavage, Disulfide Bond Formation, and Glycosylation
Newly synthesized proteins undergo several modifications in the ER to ensure proper folding, stability, and function.
Proteolytic Cleavage: Signal peptidase removes signal sequences.
Disulfide Bond Formation: Protein disulfide isomerase (PDI) forms and rearranges disulfide bonds, stabilizing protein structure and protecting against oxidative damage.
N-linked Glycosylation: Oligosaccharyltransferase adds sugar chains to asparagine residues (Asn-X-Ser/Thr, X ≠ Pro).
Molecular Chaperones: Heat shock proteins (e.g., Hsp70/BiP), calnexin, and calreticulin assist in protein folding and prevent aggregation.
Glycosylation and Its Functions
Glycosylation is the covalent attachment of oligosaccharide chains to proteins, producing glycoproteins. This modification protects proteins, forms the glycocalyx, and enables cell recognition.
N-linked Glycosylation: Most common; occurs in the ER.
O-linked Glycosylation: Occurs in the Golgi; sugars added to serine or threonine residues.
Functions:
Protection from harsh conditions and enzymatic degradation.
Formation of glycocalyx for cell surface protection.
Cell recognition via specific sugar patterns.
Quality Control and Checkpoints in the Endomembrane System
Protein Quality Control: The Five Checkpoints
Proteins must pass several quality control checkpoints to ensure only properly folded and modified proteins proceed through the endomembrane system.
Checkpoint | Purpose | Fate of Misfolded Proteins |
|---|---|---|
1. Rough ER retention (BiP & calnexin) | Identify and retain misfolded or improperly modified proteins | Remain in ER for folding or repair |
2. Retrograde retrieval from Golgi | Retrieve misfolded proteins or ER-resident proteins | Returned to ER for correction or degradation |
3. Unfolded Protein Response (UPR) | Senses buildup of misfolded proteins, increases folding capacity | Boosts chaperone production, expands ER |
4. Dislocation & degradation | Removes irreparably misfolded proteins from ER | Exported to cytosol, ubiquitinated, degraded by proteasomes |
5. Apoptosis | Protects organism if damage is unresolvable | Triggers programmed cell death |
Chaperone Proteins and Folding
Chaperone proteins such as BiP, GRP94, calnexin, calreticulin, and PDI assist in protein folding and prevent aggregation. Lectin chaperones recognize glycosylated proteins, ensuring proper folding and quality control.
General Chaperones: BiP, GRP94, GRP170
Lectin Chaperones: Calnexin, calreticulin
Enzymatic Chaperones: Protein disulfide isomerase (PDI), peptidyl–prolyl cis–trans isomerase
Unfolded Protein Response (UPR) and Degradation
If misfolded proteins accumulate, the Unfolded Protein Response (UPR) is triggered, increasing chaperone production and expanding the ER. Irreparably misfolded proteins are removed from the ER, ubiquitinated, and degraded by proteasomes. If damage is unresolvable, apoptosis is triggered to protect the organism.
UPR Mechanism: Bip dissociates from Ire1, triggering a signaling cascade that increases folding capacity.
Dislocation: Misfolded proteins are exported to the cytosol for degradation.
Apoptosis: Programmed cell death if protein damage cannot be resolved.
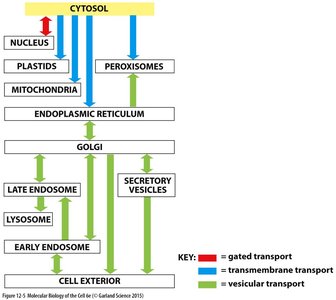
Summary Table: Trafficking Pathways in the Endomembrane System
Organelle | Transport Type | Direction |
|---|---|---|
Nucleus | Gated transport | Cytosol → Nucleus |
ER | Transmembrane transport | Cytosol → ER |
Golgi | Vesicular transport | ER → Golgi |
Endosomes/Lysosomes | Vesicular transport | Golgi → Endosomes/Lysosomes |
Cell Exterior | Vesicular transport | Golgi → Cell Membrane |
Conclusion
The endomembrane system is a highly organized network that ensures proper synthesis, modification, sorting, and quality control of proteins and lipids. The ER plays a central role in these processes, with specialized mechanisms for protein targeting, folding, and trafficking. Quality control checkpoints and chaperone proteins maintain cellular health by preventing the accumulation of misfolded proteins and triggering protective responses when necessary.
