 Back
BackMicroscopy and Membrane Structure: Principles and Applications
Study Guide - Smart Notes

Microscopy: Visualizing Cells
Factors Affecting Resolution
The resolution of a microscope is its ability to distinguish two closely spaced objects as separate entities. It is fundamentally limited by the wavelength of light and the optical properties of the system.
Wavelength (λ): Shorter wavelengths provide better resolution.
Numerical Aperture (NA): Defined as n sinθ, where n is the refractive index of the medium and θ is the half-angle of the maximum cone of light that can enter the lens.
Refractive Index (n): Higher values (e.g., using oil instead of air) improve resolution.
The limit of resolution is the minimum distance at which two points can be distinguished. The relationship is given by:
Smaller resolution value = better resolving power.
Higher numerical aperture and refractive index = better resolution.
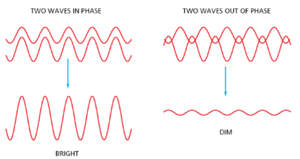

Types of Light Microscopy
Different light microscopy techniques enhance contrast and reveal cellular details in various ways:
Bright-field: White light passes through the sample; contrast is generated by stains that absorb light.
Dark-field: Only scattered light enters the objective, making unstained specimens visible against a dark background.
Phase-contrast/Differential Interference Contrast (DIC): Utilize differences in refractive index and interference of light waves to visualize transparent specimens without staining.
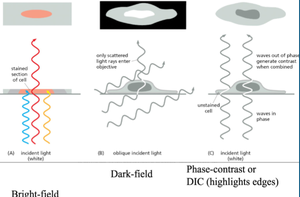
Sample Preparation: Fixing, Staining, and Sectioning
To observe cells and tissues under a microscope, samples are typically fixed, sectioned, and stained:
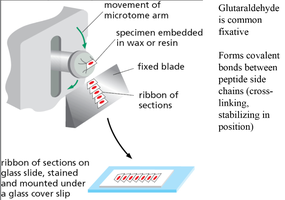
Fixation: Chemicals like glutaraldehyde cross-link proteins, preserving cellular structure.
Sectioning: Samples are embedded in wax or resin and sliced into thin sections using a microtome.
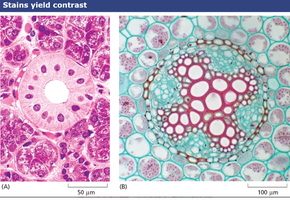
Staining: Dyes bind to specific cell components, enhancing contrast.
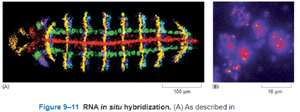
In-Situ Hybridization
In-situ hybridization uses labeled DNA or RNA probes to detect specific RNA sequences within cells or tissues, revealing gene expression patterns.
Allows localization of gene activity in situ (in the original place).
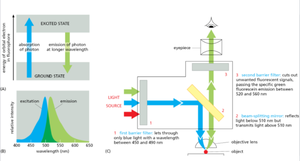
Fluorescence Microscopy
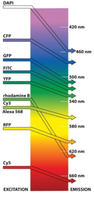
Fluorescence microscopy uses fluorophores that absorb light at one wavelength (excitation) and emit at a longer wavelength (emission). This technique enables visualization of specific molecules within cells.
Different fluorophores have distinct excitation and emission spectra.
Filters and dichroic mirrors ensure only the desired wavelengths reach the detector.

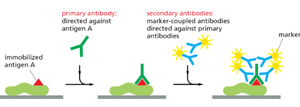
Immunolocalization and Fluorescent Proteins
Immunohistochemistry and immunocytochemistry use antibodies to detect specific proteins. Secondary antibodies conjugated to fluorophores amplify the signal.
Fluorescent proteins (e.g., GFP) can be genetically fused to proteins of interest, allowing live-cell imaging of protein localization and dynamics.
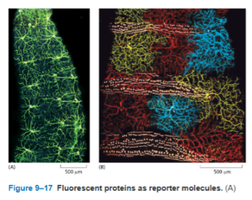
Membrane Structure and Dynamics
Properties of Amphipathic Lipids
Amphipathic lipids contain both hydrophilic (polar) and hydrophobic (nonpolar) regions, driving their self-assembly into structures such as bilayers, micelles, and liposomes.
Lipid bilayer: Most stable arrangement, forms the basis of biological membranes.
Micelles: Spherical structures with hydrophobic cores.
Liposomes: Spherical vesicles with a lipid bilayer enclosing an aqueous compartment.
Membrane Fluidity
Membrane fluidity is influenced by lipid composition and temperature:
Cholesterol: Reduces fluidity at high temperatures, prevents solidification at low temperatures.
Saturated fatty acids: Pack tightly, decreasing fluidity.
Unsaturated fatty acids: Kinks from double bonds increase fluidity.
Shorter chains: Increase fluidity.
Movement of Membrane Lipids
Lateral diffusion: Lipids move within the same leaflet, contributing to membrane fluidity.
Rotational movement: Lipids rotate around their long axis.
Flip-flop: Rare spontaneous movement between leaflets; requires enzymes (flippases, scramblases).
Lipid Aggregation and Rafts
Lipid aggregation: Spontaneous formation of bilayers, micelles, and liposomes due to amphipathic nature.
Lipid rafts: Microdomains enriched in cholesterol, saturated lipids, and glycolipids; more ordered and tightly packed, often involved in signaling.
Lipid Droplet Formation
Originates from the cytosolic leaflet of the ER membrane.
Requires proteins such as seipin for proper formation.
Stores neutral lipids like triacylglycerol and cholesterol esters.
Asymmetrical Features of Biological Membranes
Lipids are distributed asymmetrically between the two leaflets.
Negatively charged lipids (e.g., phosphatidylserine) face the cytosol.
Glycolipids and carbohydrates are exposed on the extracellular surface, important for cell recognition and protection.
