 Back
BackNucleus: Gene Expression and Chromatin Structure
Study Guide - Smart Notes

Chromosome and Chromatin Organization
Levels of Chromatin Structure
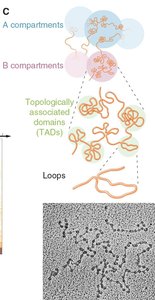
Chromatin is organized into hierarchical structures within the nucleus, which are essential for gene regulation and genome stability.
Nucleosomes: The basic unit of chromatin, consisting of DNA wrapped around histone proteins.
Loops: Chromatin forms loops that bring distant regions of DNA into proximity, facilitating regulatory interactions.
Topologically Associated Domains (TADs): Self-interacting genomic regions that help organize the genome into functional compartments.
Compartments: Larger domains, such as A (active/euchromatin) and B (inactive/heterochromatin) compartments, reflect transcriptional activity.
Chromosome Structure and Types
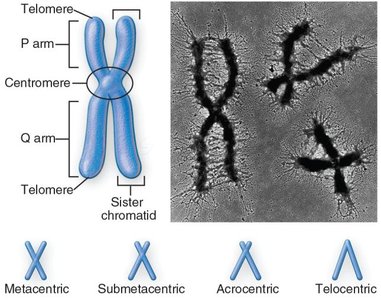
Chromosomes are highly organized structures composed of chromatin. They are classified based on the position of the centromere:
Metacentric: Centromere in the middle, arms of equal length.
Submetacentric: Centromere off-center, arms of unequal length.
Acrocentric: Centromere near one end.
Telocentric: Centromere at the very end.
Centromeres and Kinetochores
Centromere Identity and Function
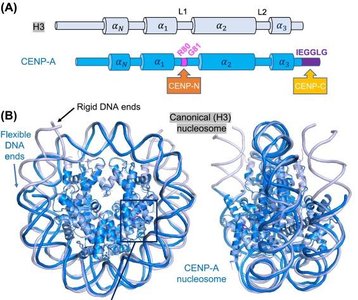
Centromeres are specialized chromosomal regions essential for proper chromosome segregation during cell division.
CENP-A: A histone H3 variant that replaces H3 in centromeric nucleosomes, altering DNA wrapping and serving as a foundation for kinetochore assembly.
CENP-B: Binds specific DNA sequences (CENP-B boxes) within centromeres.
Additional CENP proteins recognize CENP-A nucleosomes, ensuring centromere identity.
Kinetochore Structure and Function
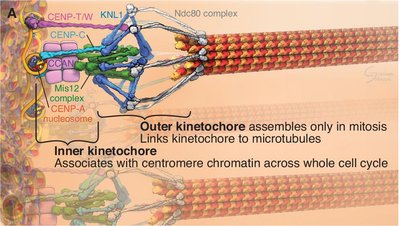
The kinetochore is a multiprotein complex that assembles on centromeres and mediates chromosome attachment to spindle microtubules during mitosis.
Inner Kinetochore: Associates with centromere chromatin throughout the cell cycle.
Outer Kinetochore: Assembles only during mitosis and links chromosomes to microtubules.
Mitotic Chromosome Structure
Chromosome Condensation and SMC Proteins
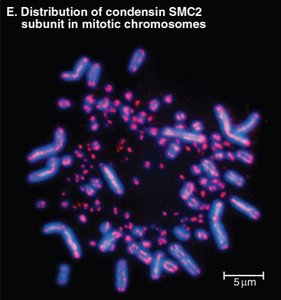
During mitosis, chromosomes condense into compact structures, a process dependent on Structural Maintenance of Chromosomes (SMC) proteins such as condensin.
Condensin: Forms ring-like complexes that organize and stabilize chromatin loops, ensuring proper chromosome segregation.
Without condensin, chromosomes are fragile and may lag during segregation.
Telomeres and Genome Stability
Telomere Structure and Function
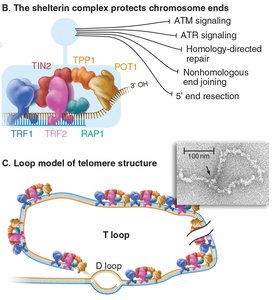
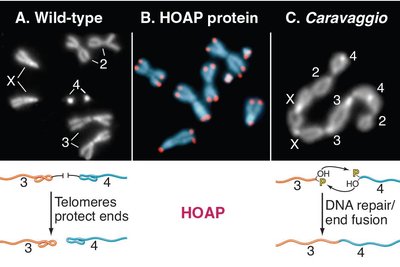
Telomeres are repetitive DNA sequences (TTAGGG in humans) and associated proteins at chromosome ends, forming a protective complex called shelterin.
Permit complete replication of chromosomal DNA.
Prevent chromosome ends from being recognized as DNA breaks.
Form specialized structures such as the T-loop, where the 3' overhang invades upstream telomeric repeats.
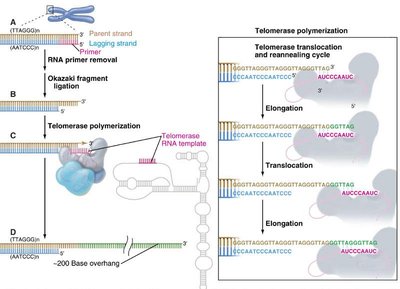
The End Replication Problem and Telomerase
During DNA replication, the lagging strand cannot be fully replicated, leading to progressive telomere shortening. Telomerase, a reverse transcriptase, extends telomeres by adding repeats using its RNA template.
Most somatic cells do not express telomerase, resulting in limited cell divisions (Hayflick limit).
Telomerase expression in cell lines allows indefinite proliferation, useful for research.
Chromatin States: Euchromatin and Heterochromatin
Definitions and Properties
Chromatin exists in two major states that influence gene expression:
Euchromatin: Loosely packed, transcriptionally active, accessible to transcription factors.
Heterochromatin: Densely packed, transcriptionally silent, often found near the nuclear envelope.
Constitutive heterochromatin is always compact (e.g., centromeres), while facultative heterochromatin can switch states (e.g., Barr body in X-inactivation).
Epigenetic Modifications of Histones
Histone tails undergo post-translational modifications (PTMs) that regulate chromatin structure and gene expression. These modifications can be inherited and include acetylation, methylation, and phosphorylation.
Acetylation (H3K9Ac, H3K4me3): Associated with open chromatin and active transcription.
Methylation (H3K9me3, H3K27me3): Associated with closed chromatin and gene silencing.
Enzymes: HATs (histone acetyltransferases) add acetyl groups; HDACs (histone deacetylases) remove them.
Histone modifications can recruit specific binding proteins (e.g., bromodomains bind acetylated lysines, chromodomains bind methylated lysines), propagating chromatin states.
Transcription and Gene Expression
Transcription Units and RNA Types
Transcription is the process by which RNA is synthesized from a DNA template. In eukaryotes, each gene typically has its own transcription unit, and the initial transcript (pre-mRNA) contains both coding and non-coding regions (introns and exons).
RNA Types: rRNA (75%), tRNA, snRNA, 5S rRNA, mRNA (10%), and small noncoding RNAs (e.g., miRNA).
RNA Polymerases: Pol I (rRNA), Pol II (mRNA, snRNA), Pol III (tRNA, 5S rRNA).
Transcription Initiation and Regulation
Transcription initiation involves the assembly of the preinitiation complex at promoter regions, recruitment of RNA polymerase II, and melting of DNA to form a transcription bubble. General transcription factors (GTFs), such as TATA-binding protein (TBP), facilitate this process.
Productive recruitment of Pol II is gene-specific and highly regulated.
Elongation involves addition of NTPs to the 3' end; Pol II can pause and edit errors.
Termination is coupled to 3' end processing, including cleavage and polyadenylation.
Summary Table: Chromatin States and Modifications
Chromatin State | Histone Modification | Transcriptional Activity | Location |
|---|---|---|---|
Euchromatin | H3K4me3, Acetylation | Active | Internal nuclear regions |
Heterochromatin | H3K9me3, H3K27me3 | Inactive | Periphery, centromeres |
