 Back
BackNucleus: Gene Expression and Chromatin Structure
Study Guide - Smart Notes

Chromosome Structure and Organization
Levels of Chromatin Organization
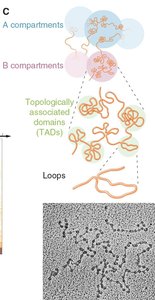
Chromatin is organized into hierarchical structures that facilitate compaction and regulation of the genome within the nucleus.
Nucleosomes: The basic unit of chromatin, consisting of DNA wrapped around histone proteins.
Loops: Chromatin forms loops that bring distant regions into proximity, influencing gene regulation.
Topologically Associated Domains (TADs): Self-interacting genomic regions that help organize the genome into functional compartments.
Compartments: Larger domains (A and B compartments) correspond to euchromatin (active) and heterochromatin (inactive) regions.
Chromosomes: The highest level of organization, visible during mitosis.
Chromosome Anatomy
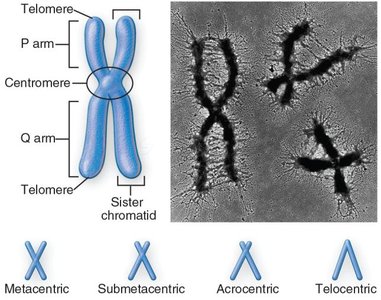
Chromosomes have distinct structural features that are essential for their function and segregation during cell division.
Centromere: The constricted region where sister chromatids are joined and where kinetochores assemble.
Telomeres: Repetitive DNA sequences at chromosome ends that protect against degradation and fusion.
P and Q arms: The short (p) and long (q) arms of a chromosome, separated by the centromere.
Chromosome Morphology: Chromosomes can be classified as metacentric, submetacentric, acrocentric, or telocentric based on centromere position.
Centromeres and Kinetochores
Centromere Identity and Function
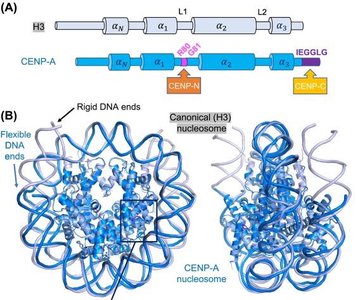
Centromeres are specialized chromosomal regions essential for accurate chromosome segregation during mitosis and meiosis.
CENP-A: A histone H3 variant that replaces H3 in centromeric nucleosomes, conferring unique structural properties.
CENP-B: Binds specific DNA sequences (CENP-B boxes) within centromeres.
Centromere-specific nucleosomes: Induce distinct DNA wrapping and are recognized by additional CENP proteins.
Kinetochore Structure and Function
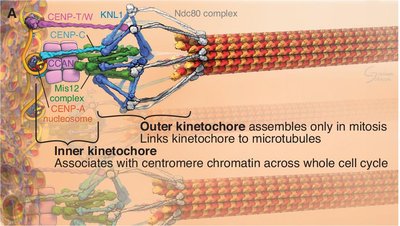
The kinetochore is a multiprotein complex that assembles on centromeres and mediates chromosome attachment to spindle microtubules.
Inner Kinetochore: Associates with centromeric chromatin throughout the cell cycle.
Outer Kinetochore: Assembles during mitosis to link chromosomes to microtubules.
Microtubule Attachment: Ensures proper chromosome segregation.
Mitotic Chromosome Structure
Chromosome Condensation and SMC Proteins
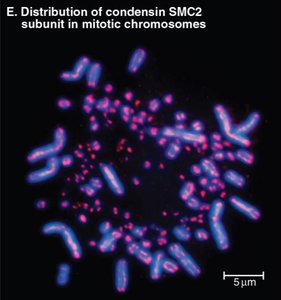
During mitosis, chromosomes condense into highly organized structures, a process dependent on Structural Maintenance of Chromosomes (SMC) proteins such as condensin.
Condensin: A protein complex that organizes and stabilizes mitotic chromosome architecture by forming loops of characteristic sizes.
Chromosome Fragility: Loss of condensin leads to fragile chromosomes and segregation errors.
Telomeres and Telomerase
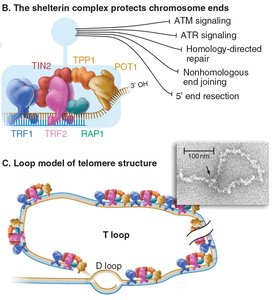
Telomere Structure and Function
Telomeres are repetitive DNA sequences (TTAGGG in humans) and associated proteins that protect chromosome ends from degradation and inappropriate repair.
Shelterin Complex: A protein complex that binds telomeric DNA and prevents activation of DNA damage responses.
T-loop Formation: The 3' overhang of telomeric DNA can invade upstream repeats, forming a protective loop structure.
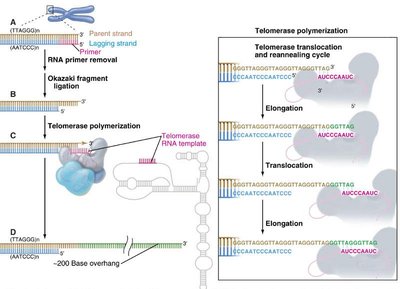
The End Replication Problem and Telomerase
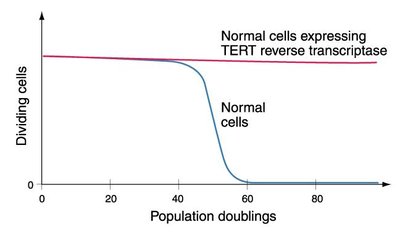
Conventional DNA replication cannot fully replicate chromosome ends, leading to progressive telomere shortening with each cell division.
Lagging Strand Synthesis: Removal of RNA primers leaves a 3' overhang.
Telomerase: A ribonucleoprotein enzyme with reverse transcriptase activity that extends telomeres using an RNA template.
Somatic Cells: Typically lack telomerase activity, resulting in replicative senescence after a finite number of divisions.
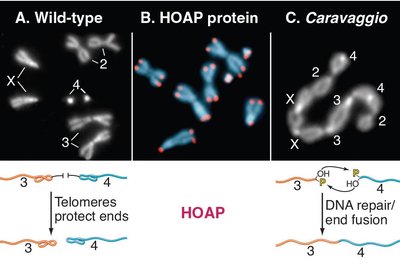
Telomere Dysfunction and Cellular Consequences
Disruption of telomere protection leads to chromosome end-to-end fusions and genome instability.
Chromosome Fusions: Occur when telomeres are not properly protected, mimicking DNA breaks.
Role in Aging and Cancer: Telomerase reactivation can bypass replicative senescence, contributing to cellular immortalization in cancer.
Chromatin States: Euchromatin and Heterochromatin
Functional Chromatin Domains
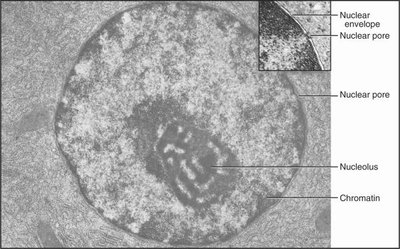
Chromatin exists in two major states that influence gene expression and nuclear organization.
Euchromatin: Loosely packed, transcriptionally active regions accessible to regulatory proteins.
Heterochromatin: Densely packed, transcriptionally silent regions, often found near the nuclear envelope or in nuclear bodies.
Constitutive Heterochromatin: Always compacted (e.g., centromeres, repetitive DNA).
Facultative Heterochromatin: Compact in some cells or conditions (e.g., Barr body/X-inactivation).
Epigenetic Regulation of Chromatin
Histone Modifications and the Histone Code
Post-translational modifications of histone tails regulate chromatin structure and gene expression, forming the basis of the "histone code." These modifications can be inherited through cell division.
Acetylation: Generally associated with open chromatin and active transcription (e.g., H3K9Ac, H3K4me3).
Methylation: Can be associated with either activation or repression, depending on the residue (e.g., H3K9me3, H3K27me3 mark heterochromatin).
Enzymes: HATs (histone acetyltransferases) add acetyl groups; HDACs (histone deacetylases) remove them.
Modification | Associated Chromatin State | Key Enzymes |
|---|---|---|
H3K9Ac, H3K4me3 | Euchromatin (active) | HATs |
H3K9me3, H3K27me3 | Heterochromatin (repressed) | Histone methyltransferases |
Transcription and RNA Polymerases
Transcription Units and RNA Classes
Transcription is the process by which RNA is synthesized from a DNA template. Eukaryotic cells contain several classes of RNA, each with distinct functions.
mRNA: Encodes proteins; processed from pre-mRNA by splicing.
rRNA: Forms the core of ribosomes; most abundant RNA.
tRNA: Transfers amino acids during translation.
snRNA: Involved in splicing and RNA processing.
Other small RNAs: Include miRNA and others with regulatory roles.
RNA Polymerases
Eukaryotes have three main RNA polymerases, each responsible for transcribing different classes of genes.
RNA Pol I: Transcribes rRNA genes.
RNA Pol II: Transcribes mRNA and some snRNA genes.
RNA Pol III: Transcribes tRNA, 5S rRNA, and other small RNAs.
Transcription Initiation and Regulation
Transcription initiation involves the assembly of a preinitiation complex at the promoter, recruitment of RNA polymerase II, and melting of DNA to form a transcription bubble.
General Transcription Factors (GTFs): Required for Pol II recruitment and initiation.
TATA Box: A promoter element recognized by TBP, present in some genes.
Elongation: RNA is synthesized in the 5' to 3' direction; Pol II can pause and edit errors.
Termination: Coupled to 3' end processing; involves cleavage and polyadenylation signals.
Summary Table: Chromatin States and Key Features
Feature | Euchromatin | Heterochromatin |
|---|---|---|
Structure | Loosely packed | Densely packed |
Transcriptional Activity | Active | Inactive |
Location | Internal nucleus | Periphery/nuclear bodies |
Histone Marks | H3K4me3, acetylation | H3K9me3, H3K27me3 |
