 Back
Back23: Plant Cell Walls and the Eukaryotic Cell Cycle: Structure, Function, and Regulation
Study Guide - Smart Notes

Plant Cell Walls
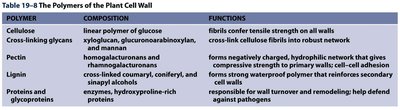
The Polymers of the Plant Cell Wall
The plant cell wall is a complex structure composed of various polymers that provide mechanical support, protection, and regulate cell growth. The main polymers include cellulose, cross-linking glycans, pectin, lignin, and proteins/glycoproteins.
Polymer | Composition | Functions |
|---|---|---|
Cellulose | Linear polymer of glucose | Fibrils confer tensile strength on all walls |
Cross-linking glycans | Xyloglucan, glucuronoarabinoxylan, and mannan | Cross-link cellulose fibrils into robust network |
Pectin | Homogalacturonans and rhamnogalacturonans | Forms negatively charged, hydrophilic network that gives compressive strength to primary walls; cell-cell adhesion |
Lignin | Cross-linked coniferyl, coniferyl, and sinapyl alcohols | Forms strong waterproof polymer that reinforces secondary walls |
Proteins and glycoproteins | Enzymes, hydroxyproline-rich proteins | Responsible for wall turnover and remodeling; help defend against pathogens |
Specialized Plant Cell Types and Cell Wall Structure
Plant cells exhibit specialized cell wall structures adapted for different functions, such as mechanical support, protection, and water transport. The arrangement and composition of cell wall polymers vary among cell types.
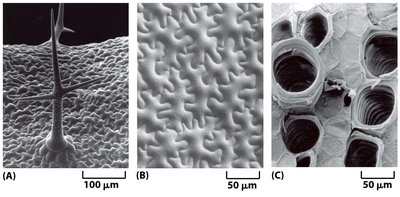

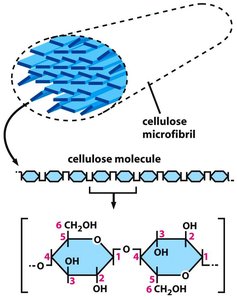
Cellulose: Structure and Function
Cellulose is the primary structural component of the plant cell wall. It consists of long, unbranched chains of β-1,4-linked glucose units, which aggregate to form microfibrils. These microfibrils provide tensile strength to the cell wall, allowing it to resist internal turgor pressure.
Structure: Cellulose molecules align parallel to each other, forming crystalline microfibrils.
Function: Provides rigidity and strength to the cell wall, essential for maintaining cell shape and withstanding osmotic pressure.
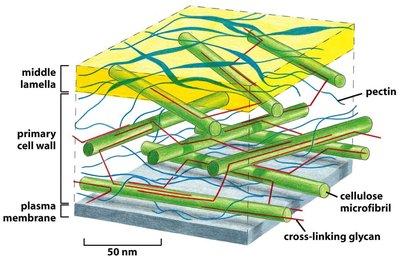
Organization of the Primary Plant Cell Wall
The primary cell wall is a composite structure consisting of cellulose microfibrils embedded in a matrix of cross-linking glycans and pectin. The middle lamella, rich in pectin, glues adjacent cells together.
Cellulose microfibrils: Provide tensile strength.
Cross-linking glycans: Connect cellulose microfibrils, forming a network.
Pectin: Fills spaces between microfibrils, providing compressive strength and hydration.
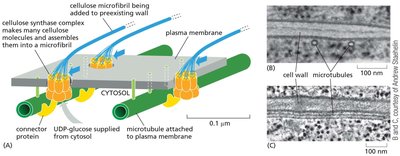
Cellulose Synthesis and Deposition
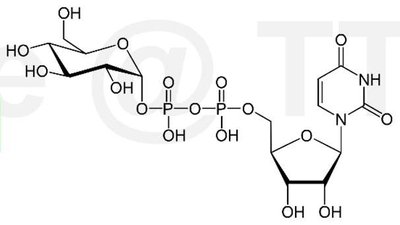
Cellulose is synthesized at the plasma membrane by cellulose synthase complexes, which use UDP-glucose as a substrate. The orientation of cellulose microfibrils is guided by cortical microtubules, influencing the direction of cell expansion.
Cellulose synthase: A membrane-bound enzyme complex that polymerizes glucose into cellulose chains.
UDP-glucose: The activated glucose donor for cellulose synthesis.
Microtubules: Guide the movement of cellulose synthase complexes, determining the orientation of microfibril deposition.
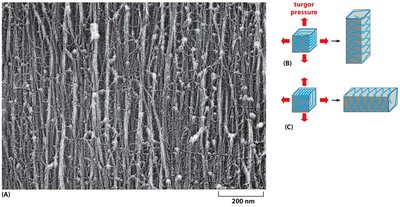
Cellulose Microfibril Orientation and Cell Elongation
The orientation of cellulose microfibrils in the cell wall determines the direction of cell elongation. Microfibrils are typically arranged perpendicular to the axis of elongation, restricting expansion in that direction and allowing elongation along the axis.
Parallel microfibrils: Restrict expansion perpendicular to their orientation.
Cell elongation: Occurs in the direction perpendicular to the microfibril orientation.

The Eukaryotic Cell Cycle and Its Control
Overview of the Cell Cycle
The cell cycle is the series of events that lead to cell division and duplication. It consists of four main phases: G1 (gap 1), S (DNA synthesis), G2 (gap 2), and M (mitosis). The cycle ensures accurate replication and segregation of genetic material.
G1 phase: Cell growth and preparation for DNA replication.
S phase: DNA replication occurs.
G2 phase: Preparation for mitosis.
M phase: Mitosis and cytokinesis (cell division).
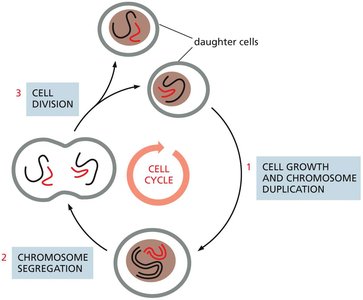
Major Events of the Cell Cycle
During the cell cycle, cells undergo nuclear division (mitosis) and cytoplasmic division (cytokinesis). These processes ensure that each daughter cell receives a complete set of chromosomes and sufficient cellular components.
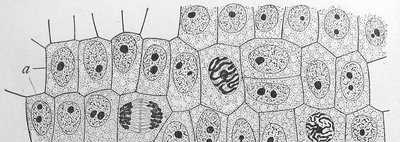
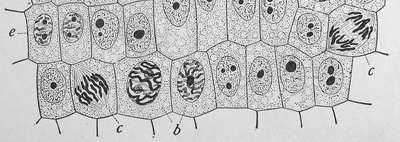
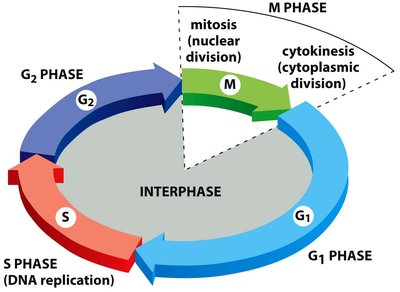
Phases of the Cell Cycle
The cell cycle is divided into interphase (G1, S, G2) and M phase (mitosis and cytokinesis). Interphase is the period of cell growth and DNA replication, while M phase is when the cell divides.
Methods for Studying Cell Cycle Progression
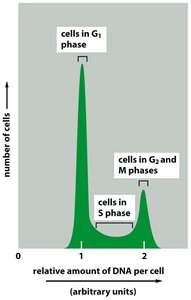
Several experimental techniques are used to study cell cycle progression, including EdU labeling for S-phase detection, live-cell imaging with fluorescent markers, and flow cytometry for DNA content analysis.
EdU labeling: Incorporates into newly synthesized DNA, detected by fluorescent azide.
Flow cytometry: Measures DNA content to distinguish cell cycle phases.
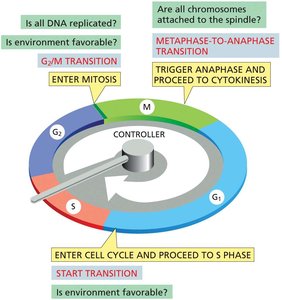
Cell Cycle Control System
The cell cycle is regulated by a control system that ensures the correct order and timing of events. Key checkpoints monitor the completion of critical processes and the cell's environment before allowing progression to the next phase.
Checkpoints: G1/S, G2/M, and metaphase-to-anaphase transitions.
Restriction point: Commitment to cell division, regulated by growth factors and nutrients.
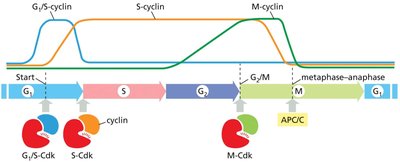
Key Regulators: Cyclins and Cyclin-Dependent Kinases (Cdks)
Progression through the cell cycle is driven by cyclin-dependent kinases (Cdks), which are activated by binding to regulatory proteins called cyclins. The levels of cyclins fluctuate during the cell cycle, while Cdk levels remain constant.
Cyclins: Regulatory proteins whose concentrations rise and fall cyclically.
Cdks: Protein kinases activated by cyclin binding; phosphorylate target proteins to drive cell cycle events.

Major Cyclins and Cdks in Vertebrates and Yeast
Different cyclin-Cdk complexes function at specific stages of the cell cycle. The table below summarizes the major cyclins and Cdks in vertebrates and budding yeast.
Cyclin–Cdk complex | Vertebrates (Cyclin) | Vertebrates (Cdk partner) | Budding yeast (Cyclin) | Budding yeast (Cdk partner) |
|---|---|---|---|---|
G1-Cdk | Cyclin D* | Cdk4, Cdk6 | Cln3 | Cdk1** |
G1/S-Cdk | Cyclin E | Cdk2 | Cln1, 2 | Cdk1 |
S-Cdk | Cyclin A | Cdk2, Cdk1** | Clb5, 6 | Cdk1 |
M-Cdk | Cyclin B | Cdk1 | Clb1, 2, 3, 4 | Cdk1 |
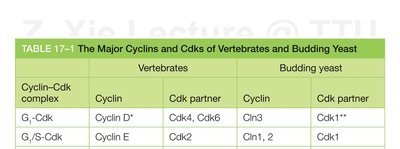

Additional info: *There are three D cyclins in mammals (D1, D2, D3). **The original name of Cdk1 was Cdc2 in vertebrates and fission yeast, and Cdc28 in budding yeast.
Regulation of Cdk Activity
Cdk activity is regulated by cyclin binding, phosphorylation, and Cdk inhibitor proteins (CKIs). Cyclin binding induces a conformational change in Cdk, activating its kinase activity. Inhibitory phosphorylation and CKIs can suppress Cdk activity, ensuring proper timing of cell cycle events.
Activation: Cyclin binding and activating phosphorylation.
Inhibition: Inhibitory phosphorylation and CKIs.
Additional info: The cell cycle control system functions as a network of biochemical switches, ensuring unidirectional and irreversible transitions between phases.
Restriction Point and Rb Protein
The restriction point in late G1 is a critical checkpoint where the cell commits to division. The retinoblastoma protein (Rb) regulates this transition by inhibiting E2F transcription factors. Phosphorylation of Rb by G1-Cdk and G1/S-Cdk releases E2F, allowing the expression of genes required for DNA replication.
Rb protein: Tumor suppressor that prevents excessive cell growth by inhibiting cell cycle progression.
E2F: Transcription factor required for S-phase gene expression.
Additional info: Loss of Rb function can lead to uncontrolled cell proliferation and cancer.
Control of Chromosome Duplication
DNA replication is tightly regulated to ensure that each chromosome is duplicated only once per cell cycle. S-Cdk initiates DNA replication by activating pre-replicative complexes (preRC) at replication origins. After initiation, origins cannot be reused until the next cell cycle.
PreRC: Pre-replicative complex loaded onto replication origins during G1.
S-Cdk: Triggers the firing of replication origins at the onset of S phase.
