 Back
BackProtein Synthesis and Sorting: Cellular Mechanisms and Regulation
Study Guide - Smart Notes

Protein Synthesis and Sorting
Overview of Protein Synthesis
Protein synthesis is a fundamental cellular process governed by the central dogma of biology, which describes the flow of genetic information from DNA to RNA to protein. This process is tightly regulated and involves multiple cellular compartments and molecular machinery.
Central Dogma: DNA is transcribed into RNA, which is then translated into protein.
Regulation: Protein synthesis is not constitutive; it is regulated by nutrient availability, environmental signals, and quality control mechanisms.
Key Cellular Compartments: Nucleus (transcription), cytoplasm (translation), endoplasmic reticulum (sorting).
Cellular Compartments Involved in Protein Synthesis
Each step of protein synthesis and sorting occurs in specific cellular compartments, ensuring proper regulation and localization of proteins.
Nucleus: Site of DNA storage, transcription, and ribosome assembly (in the nucleolus).
Cytoplasm: Location of translation and initial protein folding.
Endoplasmic Reticulum (ER): Site for synthesis and sorting of secretory and membrane proteins.
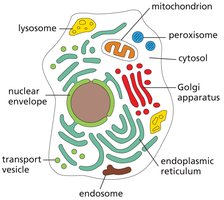
Nuclear Pore Complexes and Protein Targeting
Nuclear pore complexes (NPCs) are large structures that regulate the movement of molecules between the nucleus and cytoplasm. Proteins are targeted to the nucleus or exported out using specific signal sequences.
Nuclear Localization Signal (NLS): Directs proteins into the nucleus.
Nuclear Export Signal (NES): Directs proteins out of the nucleus.
mRNA Export: mRNA is exported from the nucleus for translation in the cytoplasm.
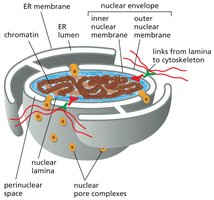
Translation: Mechanism and Regulation
Ribosomes: Structure and Function
Ribosomes are macromolecular complexes responsible for translating mRNA into polypeptides. Eukaryotic and prokaryotic ribosomes differ in size, composition, and rRNA content.
Subunits: Small subunit matches tRNA to mRNA codons; large subunit catalyzes peptide bond formation.
Composition: Eukaryotic ribosomes (80S) have more proteins and larger rRNAs than prokaryotic ribosomes (70S).
Experimental Identification: 16S rRNA (prokaryotes), 18S rRNA (eukaryotes).
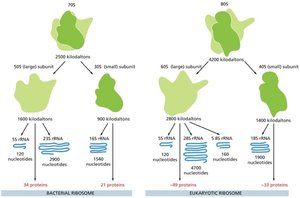
Stages of mRNA Translation
Translation occurs in three distinct stages: initiation, elongation, and termination. Each stage is mediated by specific factors and is subject to regulation.
Initiation: Formation of the 43S pre-initiation complex, recognition of the start codon (AUG), and assembly of the 80S initiation complex. Key factors include eIF2 (GTPase) and eIF4 (mRNA delivery).
Elongation: Addition of amino acids to the growing polypeptide chain, mediated by elongation factors (eEFs) and quality control mechanisms.
Termination: Release of the completed polypeptide and dissociation of the ribosome from mRNA.
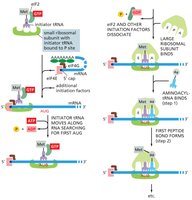
Translation Elongation: Ribosome Dynamics
During elongation, the ribosome moves along the mRNA, recruiting tRNAs and catalyzing peptide bond formation. The ribosome contains three sites: A (aminoacyl), P (peptidyl), and E (exit).
A Site: Entry point for aminoacyl-tRNA.
P Site: Holds the growing polypeptide chain.
E Site: Exit site for deacylated tRNA.

Nucleic Acids Required for Translation
Three types of nucleic acids are essential for translation:
mRNA: Provides the template for protein synthesis.
tRNA: Matches codons to amino acids.
rRNA: Structural and catalytic component of the ribosome.
Quality Control and Protein Folding
Regulation of Translation and Protein Folding
Quality control mechanisms ensure the accuracy and structural integrity of newly synthesized proteins. Chaperones, especially heat shock proteins (hsp), assist in proper folding and prevent aggregation.
Heat Shock Proteins: Hsp60, Hsp70, Hsp90 cooperate to fold proteins correctly; upregulated under stress conditions.
Chaperone Function: Maintain protein structure and prevent misfolding.
Energetic Cost: Protein folding is energetically expensive but essential for cell viability.
Summary Table: Comparison of Ribosome Types
This table summarizes the main differences between bacterial and eukaryotic ribosomes, including subunit size, protein and rRNA composition.
Type | Large Subunit | Small Subunit | rRNA | Proteins |
|---|---|---|---|---|
Bacterial (70S) | 50S | 30S | 23S, 5S, 16S | 54 |
Eukaryotic (80S) | 60S | 40S | 28S, 5.8S, 5S, 18S | ~80 |

Additional info:
Heat shock proteins are named according to their molecular weight (e.g., Hsp70 is 70 kDa).
Translation initiation is a key regulatory step, often targeted by cellular signaling pathways.
Protein sorting in the ER involves signal sequences that direct nascent proteins to their correct cellular locations.
