 Back
BackTranscription, mRNA Processing, and Translation in Prokaryotes and Eukaryotes
Study Guide - Smart Notes

Transcription: DNA to RNA
Overview of Transcription
Transcription is the process by which genetic information encoded in DNA is copied into messenger RNA (mRNA). This process is fundamental for gene expression and occurs in both prokaryotic and eukaryotic cells, though with important differences.
Template strand: The DNA strand used by RNA polymerase to synthesize RNA.
RNA polymerase: The enzyme responsible for synthesizing RNA from the DNA template.
Direction: RNA is synthesized in the 5' to 3' direction, using the 3' to 5' DNA template strand.
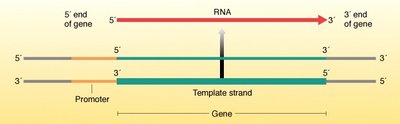
Mechanism of Transcription
Transcription involves several steps: initiation, elongation, and termination. RNA polymerase binds to the promoter region, unwinds the DNA, and synthesizes a complementary RNA strand.
Initiation: RNA polymerase binds to the promoter sequence upstream of the gene.
Elongation: RNA polymerase moves along the DNA, synthesizing RNA by adding ribonucleotides complementary to the DNA template.
Termination: Transcription ends when RNA polymerase encounters a termination signal.
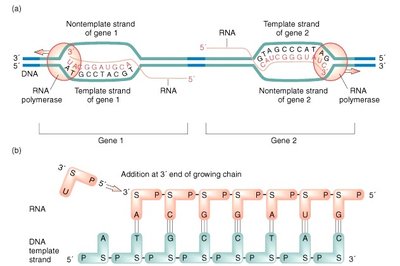
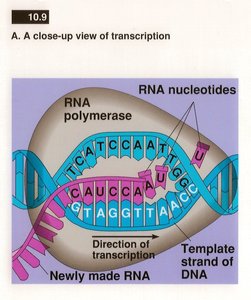
Transcription Products
The RNA produced is complementary to the DNA template strand and nearly identical to the non-template (coding) strand, except that uracil (U) replaces thymine (T).
mRNA: Messenger RNA, carries genetic information from DNA to ribosomes for protein synthesis.
tRNA and rRNA: Transfer and ribosomal RNAs are also transcribed but are not translated into proteins.

Promoters and Initiation of Transcription
Prokaryotic Promoters
Promoters are DNA sequences that define where transcription of a gene by RNA polymerase begins. In E. coli, strong promoters have conserved sequences at the -35 and -10 regions relative to the transcription start site.
-35 region: TTGACA
-10 region (Pribnow box): TATAAT
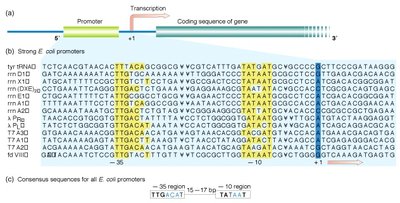
Eukaryotic Promoters
Eukaryotic promoters are more complex and differ depending on the type of RNA polymerase involved:
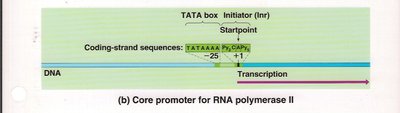
RNA polymerase II: Core promoter contains a TATA box and initiator (Inr) sequence.
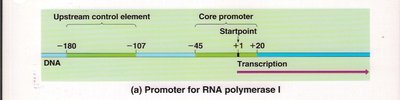
RNA polymerase I: Promoter includes upstream control elements and a core promoter.
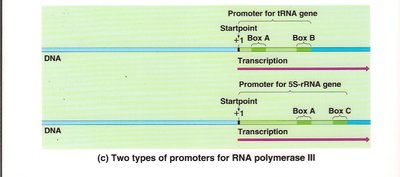
RNA polymerase III: Promoters for tRNA and 5S rRNA genes contain internal control elements (Box A, Box B, Box C).
Termination of Transcription
Rho-Independent Termination
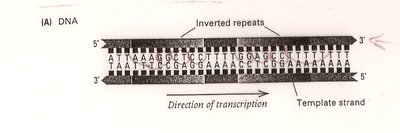
In prokaryotes, transcription can terminate via the formation of a hairpin loop in the RNA, caused by inverted repeat sequences in the DNA. This structure destabilizes the RNA-DNA hybrid and releases the RNA transcript.
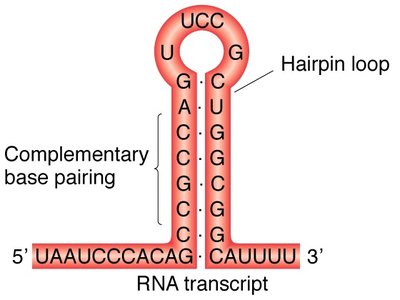
Rho-Dependent Termination
Rho-dependent termination requires the Rho protein, which binds to the rut site on the mRNA and moves toward the RNA polymerase, causing dissociation of the transcription complex.
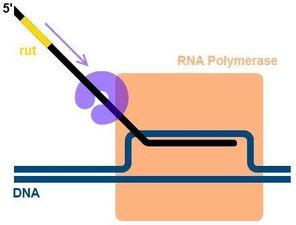
Prokaryotic vs. Eukaryotic mRNA
Key Differences
Prokaryotic mRNA: Often polycistronic (encodes multiple proteins), mature immediately after transcription, and can be translated while being transcribed.
Eukaryotic mRNA: Usually monocistronic (encodes one protein), requires processing (capping, polyadenylation, splicing), and transcription and translation are separated by the nuclear envelope.
mRNA Processing in Eukaryotes
Steps in mRNA Maturation
Eukaryotic pre-mRNA undergoes several modifications before becoming functional mRNA:
5' Capping: Addition of a methylated guanine cap to the 5' end.
3' Polyadenylation: Addition of a poly(A) tail to the 3' end.
Splicing: Removal of non-coding introns and joining of exons.
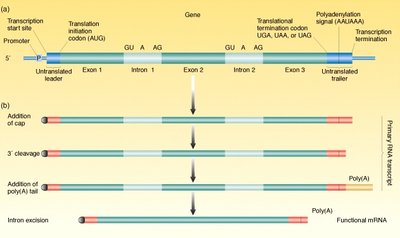
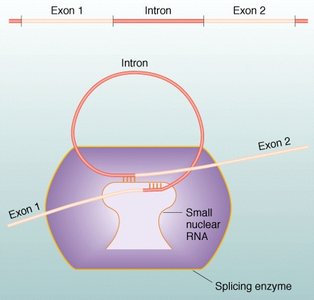
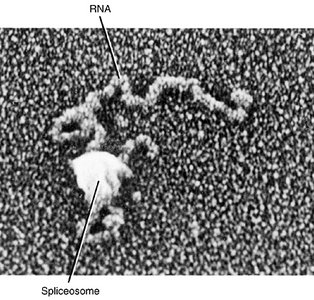
Splicing Mechanism
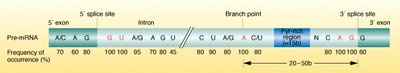
Splicing is catalyzed by the spliceosome, a complex of small nuclear RNAs and proteins. It recognizes specific sequences at the exon-intron boundaries and removes introns, joining exons together.
5' splice site: Typically GU at the intron start.
Branch point: An adenine residue within the intron.
3' splice site: Typically AG at the intron end.
The Genetic Code and Mutations
Structure of the Genetic Code
The genetic code is read in triplets (codons), with each codon specifying an amino acid. There are 64 possible codons, more than enough to encode the 20 amino acids, resulting in redundancy (degeneracy) of the code.
Codon: A sequence of three nucleotides in mRNA that specifies an amino acid.
Degeneracy: Multiple codons can code for the same amino acid.
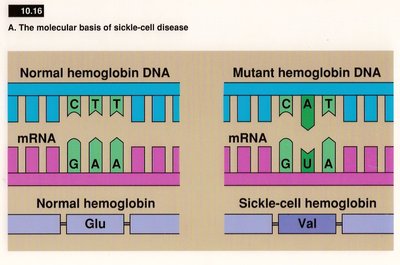
Types of Mutations
Missense mutation: Changes one amino acid to another.
Nonsense mutation: Converts a codon to a stop codon, truncating the protein.
Frameshift mutation: Insertion or deletion of nucleotides that shifts the reading frame.
Silent mutation: Alters a codon but does not change the amino acid.
Null mutation: Results in complete loss of function.
Translation: mRNA to Protein
Requirements for Translation
Translation is the process by which ribosomes synthesize proteins using mRNA as a template. It requires:
mRNA transcript
Ribosome (rRNA and proteins)
Charged tRNA (tRNA with attached amino acid)
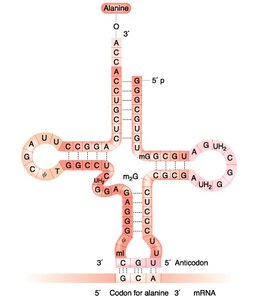
tRNA Structure and Function
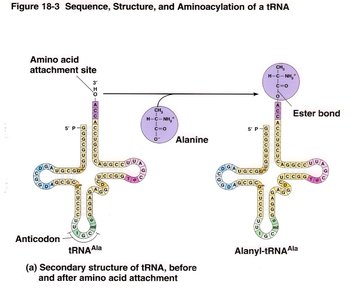
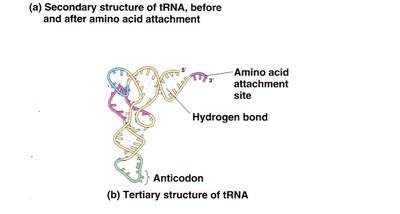
Transfer RNAs (tRNAs) have a cloverleaf secondary structure and carry specific amino acids to the ribosome. The anticodon loop pairs with the codon on mRNA.
Acceptor arm: Site of amino acid attachment (always CCA at the 3' end).
Anticodon arm: Contains the anticodon sequence complementary to the mRNA codon.
Other arms: D arm, TΨC arm, and variable arm.
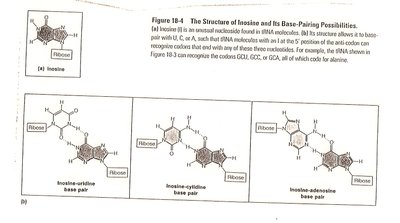
Wobble Hypothesis
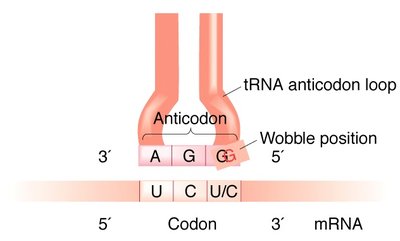
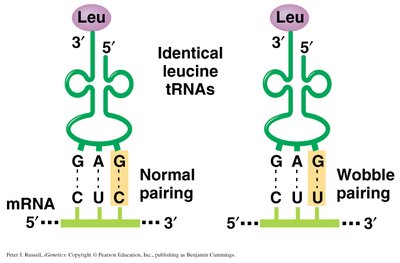
The wobble hypothesis explains how some tRNAs can recognize more than one codon due to flexible base pairing at the third codon position. Inosine (I), a modified base, can pair with A, U, or C.
Wobble position: 5' end of the tRNA anticodon, 3' end of the mRNA codon.
Result: Fewer tRNAs are needed to read all codons.
Ribosome Structure and Function
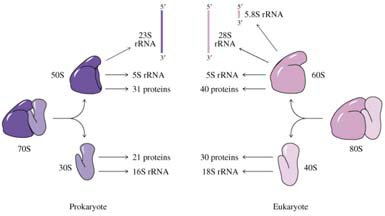
Prokaryotic and Eukaryotic Ribosomes
Ribosomes are composed of rRNA and proteins and consist of large and small subunits. Prokaryotic ribosomes are 70S (50S + 30S), while eukaryotic ribosomes are 80S (60S + 40S).
16S rRNA: Involved in initiation and termination in prokaryotes.
23S rRNA: Peptidyl transferase activity in prokaryotes.
5S rRNA: Structural role.

Initiation of Translation
Prokaryotic Initiation
In prokaryotes, the ribosome binds to the Shine-Dalgarno sequence in the 5' untranslated region (UTR) of mRNA to initiate translation. Initiation factors (IF2, IF3) assist in the assembly of the initiation complex.
Shine-Dalgarno sequence: Ribosome binding site in prokaryotic mRNA.
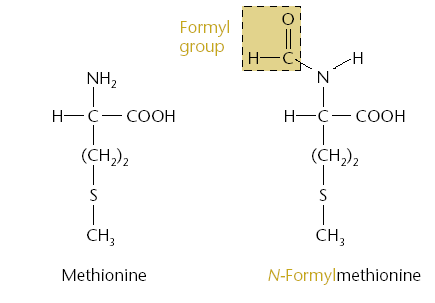
Initiator tRNA: Carries N-formylmethionine (fMet).
Eukaryotic Initiation
In eukaryotes, the small ribosomal subunit binds to the 5' cap of mRNA and scans for the start codon (AUG), which is recognized in the context of the Kozak sequence.
5' Cap: Recognized by the small ribosomal subunit.
Kozak sequence: Consensus sequence surrounding the start codon in eukaryotic mRNA.
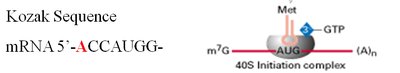
Summary Table: Prokaryotic vs. Eukaryotic mRNA and Translation
Feature | Prokaryotes | Eukaryotes |
|---|---|---|
mRNA Structure | Polycistronic, no cap or tail | Monocistronic, 5' cap, 3' poly(A) tail |
Transcription & Translation | Coupled (simultaneous) | Separated by nuclear envelope |
Initiation Site | Shine-Dalgarno sequence | 5' cap and Kozak sequence |
Initiator tRNA | fMet-tRNAfMet | Met-tRNAiMet |
Ribosome Size | 70S (50S + 30S) | 80S (60S + 40S) |
