 Back
BackL 6.3: Classification Systems: Morphological and Molecular Data in Modern Evolutionary Biology
Study Guide - Smart Notes

L 6.3
Modern Evolutionary Classification
Cladograms and Phylogenetic Trees
Modern evolutionary classification relies on interpreting phylogenetic trees (cladograms) to understand the relationships among organisms. The most recent common ancestor is the key criterion for determining relatedness, not the number of nodes separating taxa.
Cladogram: A diagram that shows evolutionary relationships among groups based on shared derived characteristics.
Most Recent Common Ancestor (MRCA): The closest shared ancestor between two taxa, indicating their evolutionary proximity.
Interpretation: For example, a frog is more closely related to a human than to a shark because the frog and human share a more recent common ancestor.
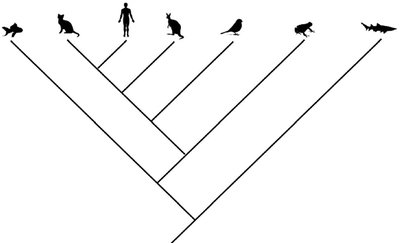
Note: The arrangement of branches (diagonal or bracketed) does not affect the interpretation; bracketed trees are often easier to read.
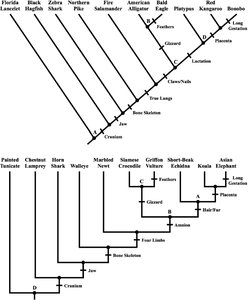
Genomic Tree of Life and DNA Barcoding
DNA Barcoding: Principles and Applications
DNA barcoding is a molecular technique for species identification using short, standardized regions of the genome. It is a cost-effective and efficient alternative to whole-genome sequencing for distinguishing species.
Animals: 648 base pair fragment of the mitochondrial cytochrome c oxidase 1 (COI) gene.
Plants: 600 bp fragment of rbcL gene and 800 bp segment of matK gene, often complemented by trnH-psbA and ITS regions.
Fungi: Internal transcribed spacer (ITS) of nuclear ribosomal DNA.
Protists: V4 region of 18S rDNA as a pre-barcode, with additional clade-specific barcodes.
Bacteria and Archaea: 16S rRNA gene is the standard barcode, despite some criticisms.
DNA barcoding has enabled global research programs and the creation of large databases, such as the Barcode of Life Data Systems (BOLD).
Microbial Barcoding and Community Fingerprinting
Techniques for Profiling Microbial Diversity
Molecular biology techniques, such as community fingerprinting, allow for rapid profiling of microbial diversity in various environments (e.g., soil, water, human microbiome) without the need for culturing individual species.
Community Fingerprinting: Uses nucleic acid techniques to detect gene variants and analyze microbial communities over time (time-series analysis).
Phylogenetic Analysis: Often based on the small subunit ribosomal RNA genes, especially the 16S rRNA gene for bacteria and archaea.
Phylotype: A classification based on genetic similarity, often used when species cannot be cultured or fully described.
The 16S rRNA gene is highly conserved and is used to determine taxonomy, phylogeny, and evolutionary divergence rates. Carl Woese used 16S rDNA to define the three domains of life: Bacteria, Archaea, and Eukarya.
Polymerase Chain Reaction (PCR)
Principles and Steps of PCR
The Polymerase Chain Reaction (PCR) is a rapid, automated method for amplifying specific DNA sequences. It is essential for molecular identification and barcoding.
Denaturation: Heating the DNA to 94–95°C to separate the double strands.
Annealing: Cooling to allow primers to bind to complementary sequences on the template DNA.
Extension: DNA polymerase extends the primers at 72°C, synthesizing new DNA strands.
Primers: Short nucleic acid sequences (18–24 bp) required for DNA polymerase to initiate synthesis.
These steps are repeated in cycles to exponentially amplify the target DNA region.
Microarray Technology: PhyloChip
High-Throughput Microbial Identification
The PhyloChip is a microarray-based technology for detecting and identifying thousands of microbial taxa simultaneously, using 16S rRNA gene probes.
Microarray: Contains hundreds of thousands of nucleotide probes for hybridization with known 16S rRNA gene sequences.
Application: Enables comprehensive analysis of microbial communities in environmental and clinical samples.

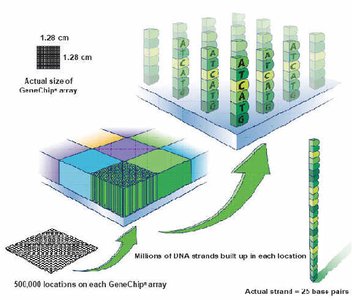
Analysis of Results: Sequence Search and BLAST
Gene Bank Inquiry and BLAST Algorithm
After sequencing, the Basic Local Alignment Search Tool (BLAST) is used to compare nucleotide sequences against databases such as GenBank to identify organisms based on sequence similarity.
BLAST: An algorithm that finds regions of local similarity between sequences, aiding in species identification and taxonomic classification.
16S rRNA Sequencing: Considered the gold standard for bacterial identification, capable of discovering novel species and reclassifying known taxa.
Hypervariable Regions: Specific regions within the 16S rRNA gene provide species-specific signatures for identification.
Applications: Used in medical microbiology for rapid, cost-effective bacterial identification, and in environmental studies to describe uncultured species.
Summary Table: Standard DNA Barcodes for Major Groups
Group | Standard Barcode Region | Notes |
|---|---|---|
Animals | COI (cytochrome c oxidase 1, 648 bp) | Mitochondrial gene, widely used |
Plants | rbcL (600 bp), matK (800 bp), trnH-psbA, ITS | Plastid and nuclear DNA regions |
Fungi | ITS (internal transcribed spacer) | Nuclear ribosomal DNA |
Protists | V4 region of 18S rDNA, others | Two-step strategy, clade-specific |
Bacteria & Archaea | 16S rRNA gene | Highly conserved, gold standard |
Key Equations and Concepts
PCR Amplification: The number of DNA copies after n cycles is given by: where is the initial number of DNA molecules and is the number of cycles.
Additional info: The notes above integrate both morphological (tree interpretation, cladistics) and molecular (DNA barcoding, PCR, microarrays, BLAST) approaches to classification, reflecting modern trends in systematics and taxonomy.
