 Back
BackControl of Gene Expression in Bacteria: The lac and trp Operons
Study Guide - Smart Notes

Control of Gene Expression in Bacteria
Overview of Gene Regulation
Gene expression in bacteria is tightly regulated to ensure that proteins are produced only when needed, conserving energy and resources. The primary mechanisms of regulation include transcriptional, translational, and post-translational control, with transcriptional control being the most common in bacteria.
Transcriptional control: Regulates whether or not an mRNA is synthesized from a gene.
Translational control: Regulates whether or not a protein is synthesized from an mRNA.
Post-translational control: Regulates the activity of proteins after they have been made, often through chemical modifications.
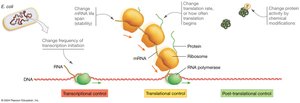
Why do bacteria primarily utilize transcriptional control? Transcriptional control is energy efficient because it prevents the synthesis of unnecessary mRNAs and proteins. This is especially important for single-celled organisms like bacteria that must rapidly adapt to changing environments.
Advantage of post-translational control: It allows for the fastest response to environmental changes because proteins can be activated or deactivated almost instantly.
Lactose Metabolism in E. coli: A Model System
Key Enzymes and Transporters
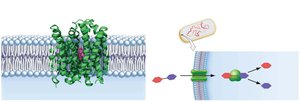
Escherichia coli can metabolize lactose when glucose is unavailable. Two main proteins are involved:
Galactoside permease: Transports lactose into the cell.
β-galactosidase: Breaks down lactose into glucose and galactose.
These proteins are produced in high amounts only when lactose is present and glucose is absent, demonstrating regulated gene expression.
Mechanisms of Transcriptional Regulation
Negative and Positive Control
Transcription in bacteria can be regulated by two main mechanisms:
Negative control: A repressor protein binds to DNA and prevents RNA polymerase from transcribing the gene.
Positive control: An activator protein binds to DNA and helps RNA polymerase initiate transcription.
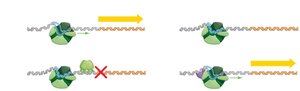
Negative control acts like an on/off switch, while positive control can increase the rate of transcription, acting more like a volume knob.
The lac Operon: Structure and Function
Components of the lac Operon
The lac operon is a group of genes involved in lactose metabolism, regulated together as a single unit. Its main components are:
lacI: Encodes the repressor protein.
Promoter: Site where RNA polymerase binds to initiate transcription.
Operator: DNA sequence where the repressor binds.
lacZ: Encodes β-galactosidase.
lacY: Encodes galactoside permease.
lacA: Encodes transacetylase (function less critical for lactose metabolism).

Negative Control of the lac Operon
In the absence of lactose, the lac repressor (encoded by lacI) binds to the operator, blocking RNA polymerase and preventing transcription. When lactose is present, it acts as an inducer by binding to the repressor, causing it to release from the operator, allowing transcription to proceed.
Without lactose: No transcription of lac genes.
With lactose: Transcription of lac genes occurs.
Mutations in the lac operon: Mutations in the operator, repressor, or structural genes can alter the regulation and expression of the operon, leading to constitutive expression or inability to metabolize lactose.
Regulation of the lac Operon by Glucose
Positive Control via CAP and cAMP
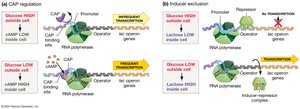
When glucose is scarce, cyclic AMP (cAMP) levels rise. cAMP binds to the catabolite activator protein (CAP), which then binds to the promoter region of the lac operon, enhancing RNA polymerase binding and increasing transcription.
Inducer Exclusion
When glucose is present, it inhibits the activity of galactoside permease, preventing lactose from entering the cell. This is known as inducer exclusion and ensures that the lac operon is not expressed when glucose is available.
Comparison: lac Operon vs. trp Operon
The trp Operon: Alternative Negative Control
The trp operon encodes enzymes for tryptophan biosynthesis. It is regulated by a repressor that is active only when bound to tryptophan (the corepressor). When tryptophan is abundant, it binds to the repressor, which then blocks transcription. When tryptophan is scarce, the repressor is inactive, and the operon is transcribed.
Similarity: Both operons use negative control via a repressor protein.
Difference: The lac operon is induced by the presence of lactose (substrate induction), while the trp operon is repressed by the presence of tryptophan (end-product repression).

Summary Table: Comparison of lac and trp Operons
Feature | lac Operon | trp Operon |
|---|---|---|
Regulatory Mechanism | Inducible (substrate induction) | Repressible (end-product repression) |
Default State | Off (repressor bound) | On (repressor inactive) |
Effector Molecule | Lactose (inducer) | Tryptophan (corepressor) |
Function | Lactose metabolism | Tryptophan biosynthesis |
