 Back
BackDNA and the Gene: Synthesis and Repair – Study Notes
Study Guide - Smart Notes

DNA and the Gene: Synthesis and Repair
Testing Early Hypotheses about DNA Synthesis
Understanding how DNA replicates was a foundational question in molecular biology. Three main models were proposed:
Semiconservative replication: Parental DNA strands separate and each serves as a template for a new daughter strand. Each daughter DNA molecule consists of one old and one new strand.
Conservative replication: The parental DNA molecule serves as a template for an entirely new molecule, so one daughter molecule is all old DNA and the other is all new DNA.
Dispersive replication: The parent DNA is cut into small pieces, and each daughter molecule contains interspersed segments of old and new DNA.
Example: The Meselson–Stahl experiment provided evidence for the semiconservative model by using isotopic labeling to distinguish old and new DNA strands.
A Model for DNA Synthesis
DNA Polymerase and Directionality
DNA polymerase is the enzyme responsible for catalyzing DNA synthesis. Several types exist, but all share key properties:
They work only in one direction: new nucleotides are added to the 3' end of a growing DNA chain, so synthesis proceeds in the 5' → 3' direction.
DNA polymerization is endergonic, but the energy comes from the high-energy bonds in deoxyribonucleoside triphosphates (dNTPs).
Formation of phosphodiester bonds during DNA synthesis is exergonic due to the hydrolysis of dNTPs.
Equation:
Origin of Replication and Replication Bubbles
DNA replication begins at specific sequences called origins of replication:
Bacteria typically have a single origin per chromosome, forming one replication bubble.
Eukaryotic chromosomes have multiple origins, forming many replication bubbles.
Each bubble has two replication forks where DNA is actively synthesized in both directions (bidirectional replication).
Opening and Stabilizing the DNA Helix
Several proteins are required to open and stabilize the DNA double helix:
DNA helicase: Breaks hydrogen bonds between DNA strands, separating them.
Single-strand DNA-binding proteins (SSBPs): Bind to separated strands to prevent them from re-annealing.
Topoisomerase: Relieves tension created by unwinding the helix by cutting and rejoining DNA downstream of the replication fork.
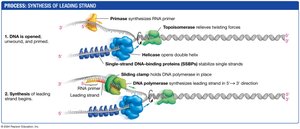
Leading Strand Synthesis
The antiparallel structure of DNA means that synthesis occurs differently on each strand. The leading strand is synthesized continuously toward the replication fork:
DNA polymerase cannot start synthesis de novo; it requires a free 3' hydroxyl group.
A short RNA primer, synthesized by primase (an RNA polymerase), provides this 3' end.
DNA polymerase then adds dNTPs to the primer’s 3' end, synthesizing the new DNA strand in the 5' → 3' direction.
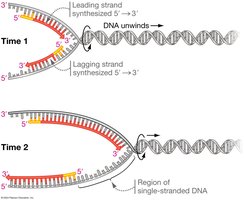
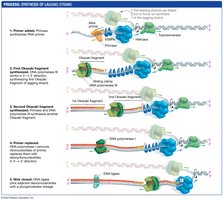
Lagging Strand Synthesis
The lagging strand is synthesized discontinuously, away from the replication fork, as short fragments called Okazaki fragments:
Primase synthesizes new RNA primers as the replication fork opens.
DNA polymerase synthesizes short DNA fragments from these primers.
Fragments are later joined into a continuous strand by DNA ligase.
Proteins Required for DNA Synthesis in Bacteria
The following table summarizes the main proteins involved in bacterial DNA replication:
Enzyme/Protein | Function |
|---|---|
Helicase | Catalyzes the separation of DNA strands to open the double helix |
Single-strand DNA-binding proteins (SSBPs) | Stabilize single-stranded DNA |
Topoisomerase | Relieves twisting forces caused by the opening of the helix |
Primase | Catalyzes the synthesis of RNA primer |
DNA polymerase III | Extends the leading strand and Okazaki fragments |
Sliding clamp | Holds DNA polymerase in place during strand extension |
DNA polymerase I | Removes RNA primer and replaces it with DNA |
DNA ligase | Joins Okazaki fragments into a continuous strand |

Replicating the Ends of Linear Chromosomes
The End Replication Problem and Telomeres
Replication of linear eukaryotic chromosomes presents a unique challenge at the ends, called telomeres:
Leading strand synthesis can proceed to the very end, but lagging strand synthesis leaves a single-stranded overhang after primer removal.
This overhang is eventually degraded, causing chromosomes to shorten with each cell division.
Telomeres consist of short, repeating, non-coding DNA sequences that protect genes from erosion.
Telomerase and Chromosome Maintenance
Telomerase is an enzyme that extends telomeres, solving the end replication problem:
Telomerase carries its own RNA template and adds DNA repeats to the overhang.
Once the overhang is long enough, normal DNA polymerase can complete lagging strand synthesis.
Telomerase is active in gametes and stem cells, but not in most somatic cells.
Example: Most cancer cells reactivate telomerase, allowing unlimited cell division.
Correcting Mistakes in DNA Synthesis
DNA Polymerase Proofreading
DNA polymerase has a proofreading function to ensure high fidelity during replication:
Incorrectly paired bases are detected by their abnormal shape.
The enzyme’s exonuclease site removes the mismatched nucleotide.
Proofreading reduces the error rate to about one mistake per 107 nucleotides.
Mismatch Repair
Sometimes, mismatches escape proofreading. Mismatch repair enzymes correct these errors after DNA synthesis:
They recognize the mismatch, remove a segment of the newly synthesized strand, and fill in the correct bases using the template strand.
Repairing Damaged DNA
DNA can be damaged by environmental factors such as UV light, X-rays, and chemicals:
UV light can cause thymine dimers, which create kinks in the DNA and block replication.
The nucleotide excision repair system recognizes distortions, removes the damaged DNA, and fills in the gap using the undamaged strand as a template.
DNA ligase seals the repaired strand into the backbone.
Summary Table: Key Steps and Enzymes in DNA Replication and Repair
Process | Key Enzyme(s) | Function |
|---|---|---|
Initiation | Helicase, SSBPs, Topoisomerase | Unwind and stabilize DNA |
Primer synthesis | Primase | Synthesize RNA primer |
Elongation | DNA polymerase III, Sliding clamp | Synthesize new DNA strand |
Primer removal | DNA polymerase I | Replace RNA primer with DNA |
Ligation | DNA ligase | Join DNA fragments |
Proofreading | DNA polymerase | Remove mismatched bases |
Mismatch repair | Mismatch repair enzymes | Correct errors post-replication |
Damage repair | Nucleotide excision repair system | Remove and replace damaged DNA |
Telomere extension | Telomerase | Extend chromosome ends |
Additional info: The accuracy and efficiency of DNA replication and repair are essential for genetic stability and prevention of diseases such as cancer.
