 Back
BackDNA and the Gene: Synthesis and Repair – Study Notes
Study Guide - Smart Notes

DNA and the Gene: Synthesis and Repair
Overview
This chapter explores how DNA replication and repair preserve genetic information. It addresses the molecular nature of genes, the mechanisms of DNA synthesis, the challenges of replicating chromosome ends, and the cellular systems for correcting DNA errors and repairing damage.
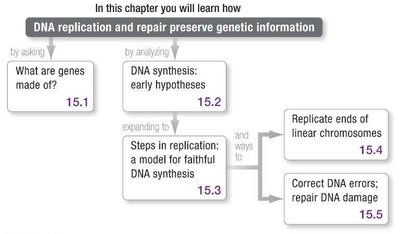
What Are Genes Made Of?
Chromosome Composition and the Hershey-Chase Experiment
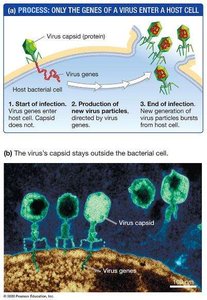
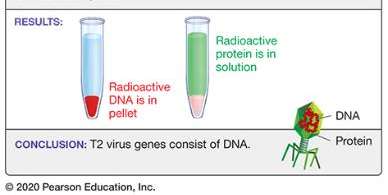
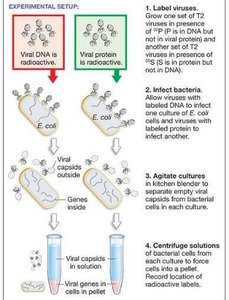
Chromosomes are composed of DNA and protein. The Hershey-Chase experiment (1952) used the T2 bacteriophage to determine whether genes are made of DNA or protein. By labeling viral DNA with radioactive phosphorus (32P) and protein with radioactive sulfur (35S), they showed that only DNA enters the bacterial cell during infection, proving that DNA is the genetic material.
Bacteriophage (phage): A virus that infects bacteria.
Capsid: The protein coat of a virus.
Key finding: Only radioactive DNA was found inside cells after infection.
Structure of DNA
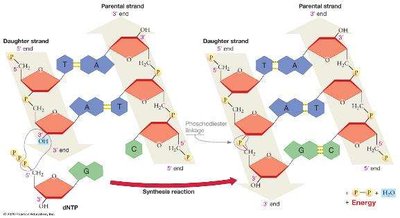
Deoxyribonucleotide Structure and DNA Polymerization
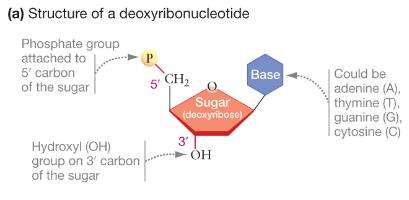
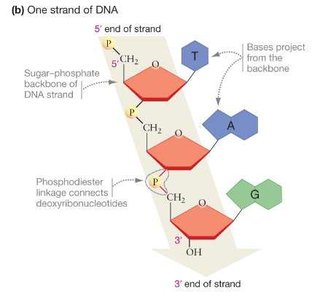
DNA is a double-stranded polymer of deoxyribonucleotides. Each nucleotide consists of a nitrogenous base (adenine, thymine, guanine, or cytosine), a deoxyribose sugar, and a phosphate group. Nucleotides are linked by phosphodiester bonds, forming a sugar-phosphate backbone.
Directionality: DNA strands have polarity, with a 5′ end (phosphate) and a 3′ end (hydroxyl).
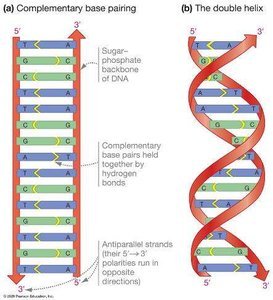
Antiparallel strands: The two DNA strands run in opposite directions.
Complementary base pairing: Adenine pairs with thymine (A-T), and guanine pairs with cytosine (G-C) via hydrogen bonds.
Testing DNA Synthesis Hypotheses
Models of DNA Replication
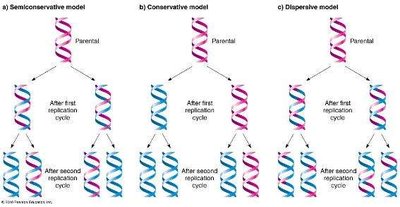
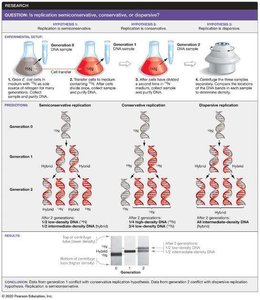
Three models were proposed for DNA replication:
Semiconservative replication: Each daughter DNA molecule consists of one parental and one newly synthesized strand.
Conservative replication: The parental molecule remains intact, and an entirely new molecule is synthesized.
Dispersive replication: Both strands of daughter molecules contain interspersed segments of parental and new DNA.
Meselson-Stahl Experiment
The Meselson-Stahl experiment (1958) used nitrogen isotopes (15N and 14N) to distinguish old and new DNA strands in Escherichia coli. Their results supported the semiconservative model of DNA replication.
Model for DNA Synthesis
DNA Polymerase and the Mechanism of Replication
DNA polymerase catalyzes DNA synthesis in the 5′ to 3′ direction, reading the template strand 3′ to 5′. Polymerization is endergonic, driven by the hydrolysis of dNTPs (deoxyribonucleoside triphosphates).
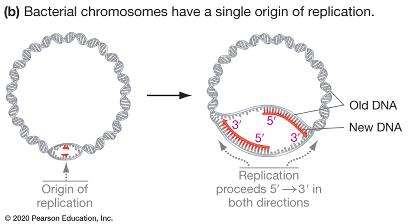
Origin of replication: Specific DNA sequence where replication begins.
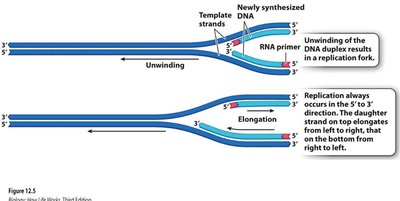
Replication bubble: Region where the DNA double helix is unwound for replication.
Replication fork: Y-shaped region where new DNA strands are synthesized.
Bidirectional replication: DNA synthesis proceeds in both directions from the origin.
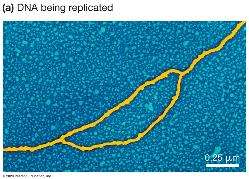


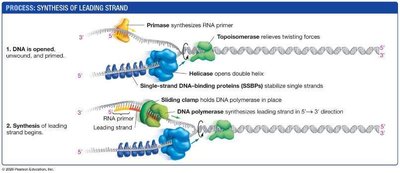
Opening and Stabilizing Single-Stranded DNA
DNA helicase: Unwinds the DNA double helix.
Single-strand DNA-binding proteins (SSBPs): Stabilize single-stranded DNA.
Topoisomerase: Relieves tension ahead of the replication fork by cutting and rejoining DNA.
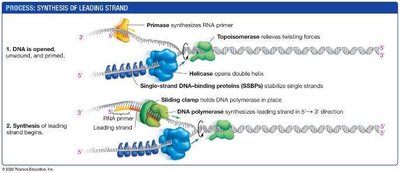
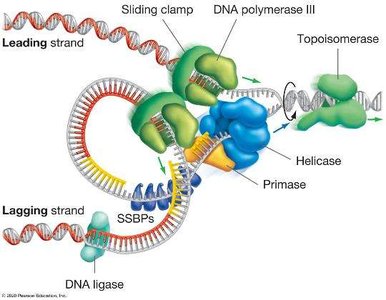
Leading and Lagging Strand Synthesis
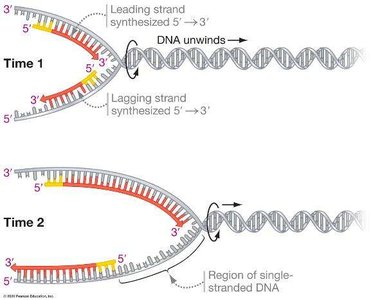
Because DNA polymerase can only synthesize in the 5′ to 3′ direction, replication is continuous on the leading strand and discontinuous on the lagging strand.
Leading strand: Synthesized continuously toward the replication fork.
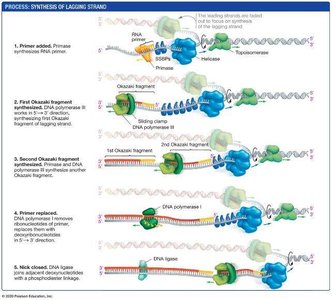
Lagging strand: Synthesized discontinuously away from the fork in short segments called Okazaki fragments.
Primase: Synthesizes short RNA primers to provide a starting point for DNA polymerase.
DNA ligase: Joins Okazaki fragments into a continuous strand.
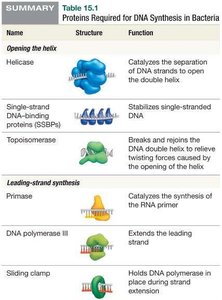
Key Proteins in DNA Replication
Name | Structure | Function |
|---|---|---|
Helicase | Catalyzes the separation of DNA strands to open the double helix | |
SSBPs | Stabilizes single-stranded DNA | |
Topoisomerase | Breaks and rejoins DNA to relieve twisting forces | |
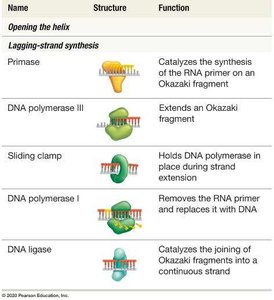
Primase | Synthesizes RNA primer | |
DNA polymerase III | Extends DNA strand | |
Sliding clamp | Holds DNA polymerase in place | |
DNA polymerase I | Removes RNA primer and replaces with DNA | |
DNA ligase | Joins Okazaki fragments |
Replication of Chromosome Ends
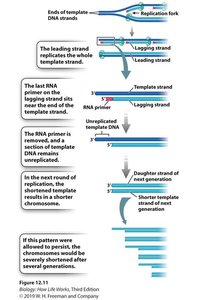
Telomeres and the End-Replication Problem
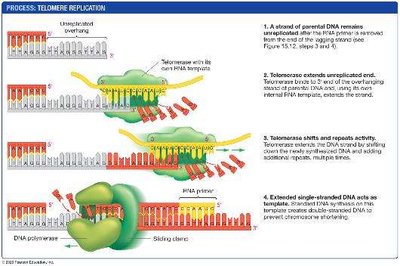
Linear chromosomes face an end-replication problem: after each round of replication, the lagging strand is shortened because the final RNA primer cannot be replaced with DNA. This leads to progressive shortening of chromosomes.
Telomeres: Repetitive DNA sequences at chromosome ends that protect coding regions.
Telomerase: An enzyme that extends telomeres using an internal RNA template. Active in germ cells, stem cells, and some cancer cells.
Somatic cells: Limited telomerase activity, leading to cellular aging.
Repairing Mistakes and DNA Damage
Proofreading and Mismatch Repair
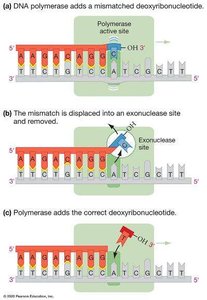
DNA polymerase is highly accurate due to its proofreading ability. If a mismatched base is incorporated, the enzyme's exonuclease site removes it. Additional mismatch repair systems correct errors missed during replication.
Proofreading: DNA polymerase removes mismatched nucleotides during synthesis.
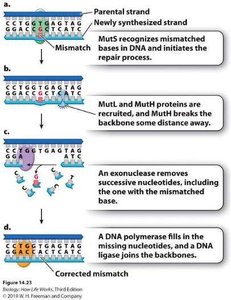
Mismatch repair: Enzymes recognize and repair mismatches after DNA synthesis is complete.
DNA Damage and Nucleotide Excision Repair
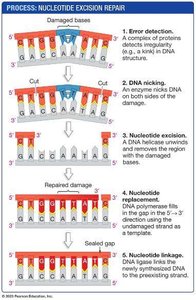
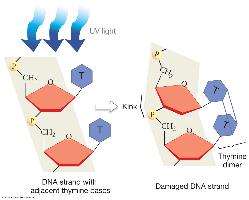
DNA can be damaged by UV light, X-rays, and chemicals. Nucleotide excision repair removes damaged regions, such as thymine dimers caused by UV light, and fills in the gap with new DNA.
Nucleotide excision repair: Detects, excises, and replaces damaged DNA segments.
Example: Xeroderma pigmentosum (XP) is a genetic disorder caused by defects in nucleotide excision repair, leading to extreme UV sensitivity and increased cancer risk.

Additional info: The accuracy and efficiency of DNA replication and repair are essential for maintaining genetic stability and preventing diseases such as cancer.
