 Back
BackDNA Damage, Mutation, and Repair Mechanisms in Cellular Metabolism
Study Guide - Smart Notes

DNA Damage and Cellular Metabolism
Overview of Cellular Metabolism
Cellular metabolism encompasses the chemical reactions that occur within cells to sustain life. Two central metabolic processes are photosynthesis and cellular respiration, which are tightly linked to the flow of energy and the cycling of matter in biological systems.
Photosynthesis: Occurs in chloroplasts, converting carbon dioxide and water into glucose and oxygen using light energy.
Cellular Respiration: Occurs in mitochondria, breaking down glucose in the presence of oxygen to produce carbon dioxide, water, and energy (ATP).
Key Equations:
(Photosynthesis)
(Cellular Respiration)
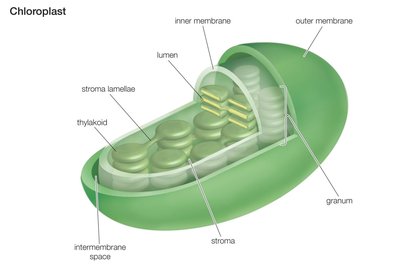
Electron Transport Chain and Final Electron Acceptors
The electron transport chain (ETC) is the final stage of aerobic respiration, where electrons are transferred through a series of protein complexes to a final electron acceptor, generating a proton gradient used to produce ATP.
Oxygen (O2) is the typical final electron acceptor in aerobic respiration, forming water.
Alternative acceptors (e.g., fluorine gas, F2) can alter the amount of ATP produced due to differences in electronegativity and energy yield.
Higher electronegativity of the final acceptor (like F2) could theoretically increase ATP yield per glucose molecule.
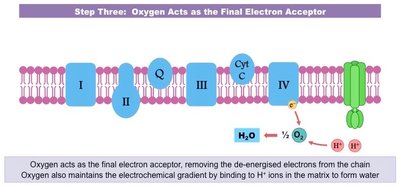
DNA Damage: Sources and Consequences
Types and Sources of DNA Damage
DNA damage can arise from endogenous metabolic processes or exogenous environmental factors. If unrepaired, DNA damage can lead to mutations, cellular dysfunction, or disease.
Reactive Oxygen Species (ROS): Byproducts of metabolism that can oxidize DNA bases, leading to mutations.
Environmental Factors: UV light, chemicals, and radiation can induce DNA lesions.
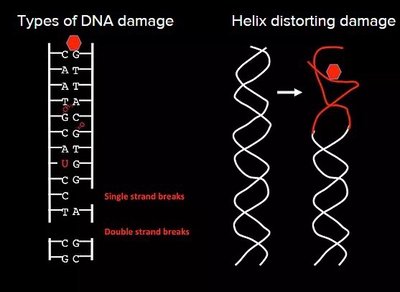
Types of DNA Damage: Single-strand breaks, double-strand breaks, base modifications, and helix-distorting lesions.
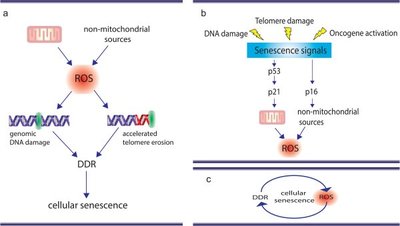
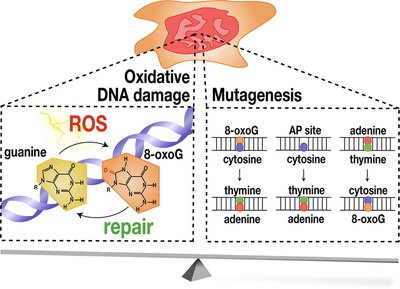
Consequences of DNA Damage
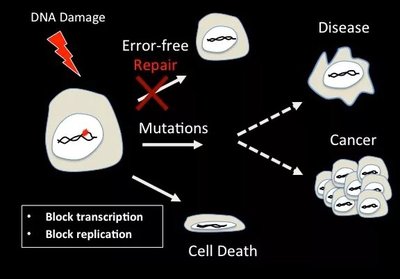
Unrepaired DNA damage can block essential cellular processes or lead to mutations, which may result in cell death, disease, or cancer.
Cell Cycle Arrest: Cells may halt division to attempt repair.
Mutagenesis: Permanent changes in DNA sequence can alter gene function.
Cell Death or Cancer: Extensive damage can trigger apoptosis or contribute to oncogenesis.
DNA Mutation Types and Their Effects
Point Mutations
Point mutations involve changes to a single nucleotide pair in DNA. They can be classified based on their effect on the encoded protein:
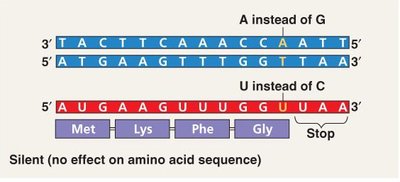
Silent Mutation: Alters a nucleotide without changing the amino acid sequence due to redundancy in the genetic code.
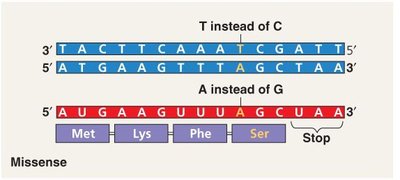
Missense Mutation: Results in a single amino acid change in the protein, which may affect function.
Nonsense Mutation: Introduces a premature stop codon, often resulting in a truncated, nonfunctional protein.
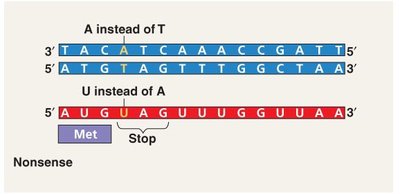
Insertions, Deletions, and Frameshift Mutations
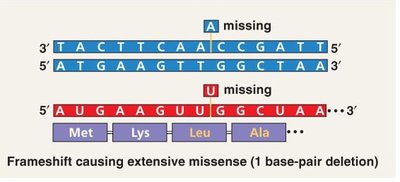
Insertions or deletions of nucleotides can shift the reading frame of the genetic code, leading to extensive changes in the amino acid sequence downstream of the mutation (frameshift mutation).
Frameshift Mutation: Caused by insertions or deletions not in multiples of three nucleotides, altering the reading frame.
Consequences: Often results in nonfunctional proteins due to widespread missense and premature stop codons.
DNA Repair Mechanisms
Overview of DNA Repair Pathways
Cells possess multiple mechanisms to detect and repair DNA damage, maintaining genomic integrity and preventing disease.
Base Excision Repair (BER): Repairs small, non-helix-distorting base lesions.
Nucleotide Excision Repair (NER): Removes bulky, helix-distorting lesions such as thymine dimers.
Mismatch Repair (MMR): Corrects errors introduced during DNA replication.
Double-Strand Break Repair: Includes homologous recombination and non-homologous end joining.
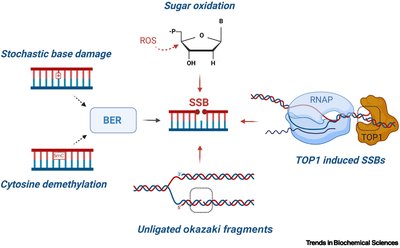
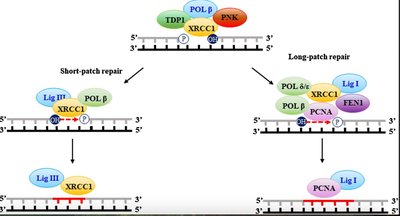
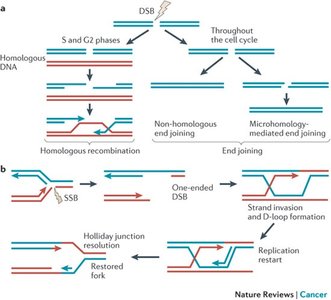
Homologous Recombination and Non-Homologous End Joining
Double-strand breaks are particularly dangerous and require specialized repair mechanisms:
Homologous Recombination (HR): Uses a homologous DNA template for accurate repair, typically active in S and G2 phases of the cell cycle.
Non-Homologous End Joining (NHEJ): Directly ligates broken DNA ends, often resulting in small insertions or deletions.
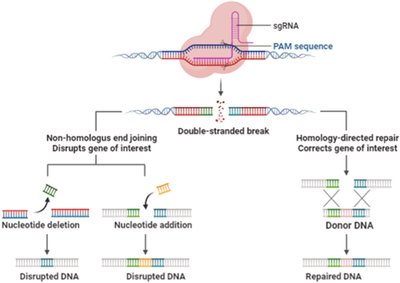
CRISPR-Cas9 and Genome Editing
The CRISPR-Cas9 system is a revolutionary genome editing tool that introduces double-strand breaks at specific genomic locations, enabling targeted gene disruption or correction via cellular repair pathways.
Non-Homologous End Joining: Can introduce insertions or deletions, disrupting gene function.
Homology-Directed Repair: Allows precise gene correction using a supplied DNA template.
Summary Table: Types of DNA Mutations
Mutation Type | Definition | Effect on Protein |
|---|---|---|
Silent | Base substitution with no amino acid change | No effect |
Missense | Base substitution causing amino acid change | Variable (may alter function) |
Nonsense | Base substitution introducing stop codon | Truncated, usually nonfunctional protein |
Frameshift | Insertion/deletion altering reading frame | Extensive missense, premature stop codons |
Further Reading and Assignments
Review Chapter 13 on meiosis for next week’s discussion on sexual reproduction.
Assignments on chapters 12 & 17 are due soon, covering cellular reproduction and gene expression.
