 Back
BackLEC 19 :DNA Replication and Repair Mechanisms
Study Guide - Smart Notes

DNA Replication and Repair
Overview of Genetic Material
Genetic material in living organisms is primarily double-stranded DNA (dsDNA), but some viruses use single-stranded DNA (ssDNA) or RNA as their genetic material. The structure and function of DNA and RNA are foundational to understanding molecular biology.
dsDNA: The genetic material for all known cellular organisms.
ssDNA: Found in some viruses.
RNA: Serves as genetic material in certain viruses (e.g., SARS-CoV-2).
RNA differs from DNA in three main ways:
Usually single-stranded (though double-stranded RNA can form).
Contains ribose sugar instead of deoxyribose.
Uses uracil as a nitrogenous base instead of thymine.
Requirements for DNA Replication
DNA replication is a highly regulated process requiring several components and enzymes to ensure accurate duplication of genetic material.
DNA Polymerase: Enzyme that catalyzes DNA synthesis from deoxyribonucleotides. Multiple types exist with specialized roles in replication and repair.
dNTPs (Deoxynucleoside Triphosphates): Building blocks for DNA synthesis (dATP, dGTP, dCTP, dTTP).
Mg2+ ions: Essential cofactors for enzyme function.
Template Strand: Provides the sequence for complementary base pairing.
Primer: A free 3'-OH group is required to initiate DNA synthesis.
Mechanism of DNA Polymerization
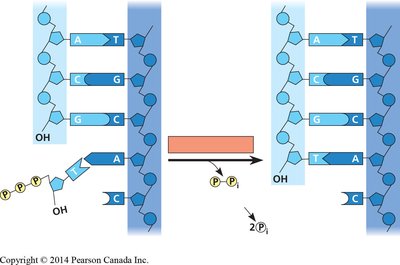
DNA polymerase adds nucleotides to the 3'-OH end of a growing DNA strand via a dehydration reaction, forming a phosphodiester bond. The energy for this reaction comes from the hydrolysis of pyrophosphate released from the incoming dNTP.
Directionality: DNA synthesis occurs in the 5' to 3' direction.
Base Pairing: The choice of incoming nucleotide is determined by the template strand, ensuring complementary base pairing (A-T, G-C).
Exergonic Reaction: The loss of pyrophosphate (PPi) drives the polymerization process.
Initiation of DNA Replication
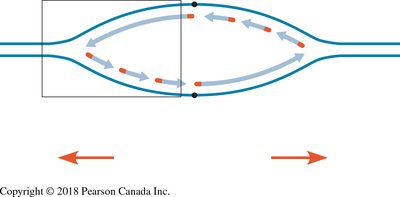
Replication begins at specific sites called origins of replication. In eukaryotes, multiple origins exist per chromosome, allowing replication to proceed in both directions, forming replication bubbles.
Helicase: Unwinds the DNA helix at the origin.
Single-Strand Binding Proteins: Stabilize unwound DNA, preventing reannealing.
Topoisomerase: Relieves strain ahead of the replication fork by cutting and rejoining DNA strands.
Elongation: Leading and Lagging Strand Synthesis
DNA polymerase III synthesizes new DNA in the 5' to 3' direction. The process differs between the leading and lagging strands due to the antiparallel nature of DNA.
Primase: Synthesizes short RNA primers to provide a starting point for DNA polymerase.
Leading Strand: Synthesized continuously toward the replication fork.
Lagging Strand: Synthesized discontinuously away from the fork in short segments called Okazaki fragments.
Sliding Clamp: Holds DNA polymerase in place during strand extension.
Okazaki fragments are joined to form a continuous strand:
DNA Polymerase I: Removes RNA primers and replaces them with DNA.
DNA Ligase: Seals nicks between adjacent DNA fragments, forming a continuous strand.
Proofreading and DNA Repair Mechanisms
High fidelity in DNA replication is maintained by proofreading and repair systems that correct errors during and after DNA synthesis.
Proofreading by DNA Polymerase III
DNA polymerase III has 3'→5' exonuclease activity, allowing it to remove incorrectly paired nucleotides immediately after incorporation.
This reduces the error rate from ~1 in 105 to ~1 in 108 nucleotides.

Mismatch Repair
Corrects errors missed by proofreading, such as mispaired bases and small insertions or deletions.
In E. coli, the parental DNA strand is methylated, allowing the repair system to distinguish the new strand for correction.
Endonucleases cut the new strand on both sides of the mismatch, DNA polymerase fills the gap, and DNA ligase seals the nick.
This process further reduces the error rate to ~1 in 1010 nucleotides.
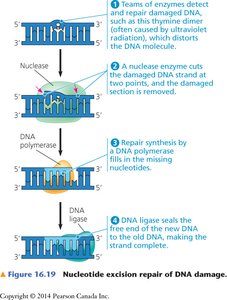
Nucleotide Excision Repair
This mechanism removes bulky DNA lesions, such as pyrimidine dimers caused by UV radiation, which distort the DNA helix.
Teams of enzymes detect the distortion.
An endonuclease cuts the DNA on both sides of the lesion, removing the damaged section.
DNA polymerase fills in the missing nucleotides.
DNA ligase seals the final nick in the backbone.
Clinical Relevance: Xeroderma Pigmentosum (XP)
XP is a genetic disorder caused by defects in nucleotide excision repair enzymes. Individuals with XP are highly sensitive to UV light and have a greatly increased risk of skin cancer due to the inability to repair UV-induced DNA damage.
DNA and Protein Synthesis
DNA encodes the information for protein synthesis, but it is the proteins themselves that carry out most cellular functions, including structural, enzymatic, and regulatory roles. The sequence of amino acids in a protein determines its structure and function, and this sequence is ultimately dictated by the DNA sequence.
