 Back
BackDNA Replication: Structure, Mechanism, and Enzymatic Function
Study Guide - Smart Notes

DNA Structure and Historical Discoveries
Discovery of DNA Structure
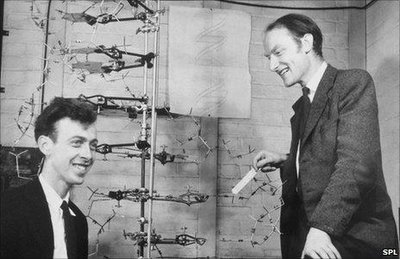
The structure of DNA was elucidated in the early 1950s through the combined efforts of several scientists. Chargaff's rules, Rosalind Franklin's X-ray diffraction images, and the model-building of Watson and Crick were pivotal in revealing the double helix structure of DNA.
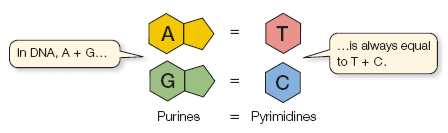
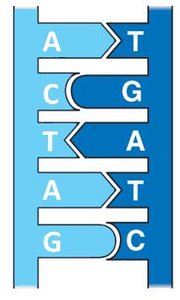
Chargaff's Rules: The amount of adenine (A) equals thymine (T), and the amount of guanine (G) equals cytosine (C) in DNA. This base pairing is essential for the double helix structure.
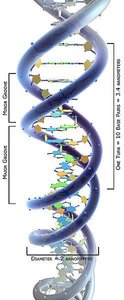
X-ray Diffraction: Provided evidence for the helical structure of DNA.
Watson and Crick Model: Proposed the double helix structure, with complementary base pairing and antiparallel strands.

Components of DNA and RNA Nucleotides
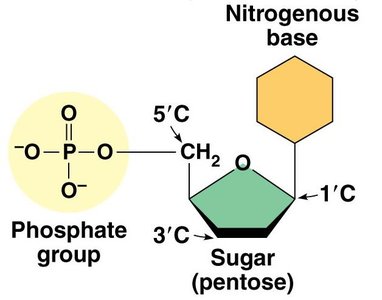
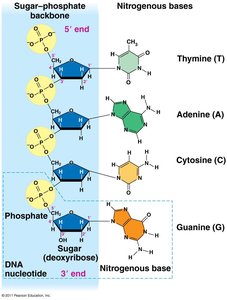
Both DNA and RNA nucleotides are composed of three main components:
Pentose Sugar: Deoxyribose in DNA, ribose in RNA.
Phosphate Group: Links nucleotides together via phosphodiester bonds.
Nitrogenous Base: Adenine (A), Thymine (T), Guanine (G), Cytosine (C) in DNA; Uracil (U) replaces Thymine in RNA.
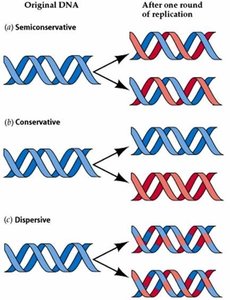
Semiconservative Model of DNA Replication
Overview of DNA Replication
DNA replication is the process by which a cell copies its DNA, ensuring that each daughter cell receives an identical set of genetic information. This process occurs during the S phase of interphase in the cell cycle.
Semiconservative Replication: Each new DNA molecule consists of one original (parental) strand and one newly synthesized strand.
Template Mechanism: Parental strands serve as templates for the synthesis of complementary new strands.
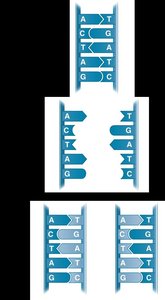
Steps of DNA Replication
The parent DNA molecule unwinds, exposing two template strands.
Each template strand is used to synthesize a new complementary strand.
The result is two DNA molecules, each with one old and one new strand.
Ingredients and Enzymes Required for DNA Replication
Key Ingredients
Template DNA: The original DNA strands to be copied.
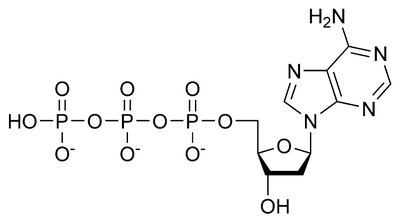
Deoxyribonucleoside Triphosphates (dNTPs): Building blocks for new DNA strands (dATP, dTTP, dCTP, dGTP).
Enzymes and Proteins: Catalyze and regulate the replication process.
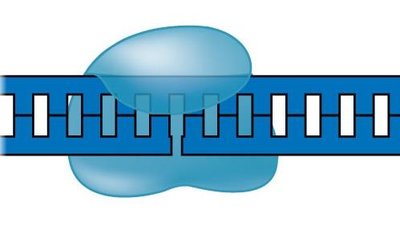
Major Enzymes and Their Functions
Helicase: Unwinds the DNA double helix at the replication fork.
Single-Strand Binding Proteins (SSBs): Stabilize unwound DNA strands.
Topoisomerase: Relieves supercoiling ahead of the replication fork.
Primase: Synthesizes short RNA primers to provide a starting point for DNA synthesis.
DNA Polymerase III: Main enzyme that adds nucleotides to the growing DNA strand in the 5' to 3' direction.
DNA Polymerase I: Removes RNA primers and replaces them with DNA.
DNA Ligase: Joins Okazaki fragments on the lagging strand, forming a continuous DNA strand.
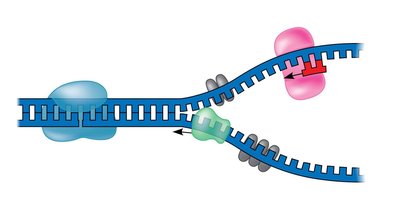
Mechanism of DNA Replication
Initiation of Replication
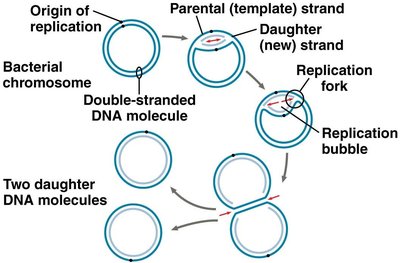
Replication begins at specific sequences called origins of replication (ori). Prokaryotes typically have a single origin, while eukaryotes have multiple origins to facilitate rapid replication of large genomes.
Replication Fork: The Y-shaped region where DNA is unwound and new strands are synthesized.
Replication Bubble: The region of active DNA synthesis formed by the opening of the double helix.
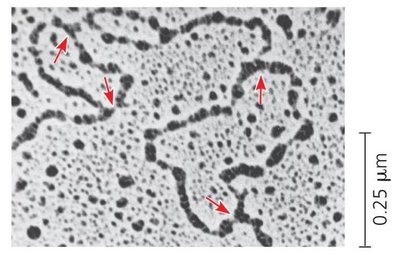
Elongation: Leading and Lagging Strands
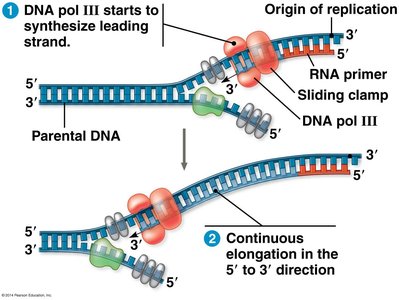
DNA polymerase can only synthesize DNA in the 5' to 3' direction. This leads to continuous synthesis on one strand (leading) and discontinuous synthesis on the other (lagging).
Leading Strand: Synthesized continuously toward the replication fork, requiring only one primer.
Lagging Strand: Synthesized discontinuously away from the fork in short segments called Okazaki fragments, each requiring a new primer.
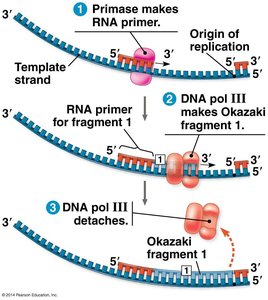
Joining Okazaki Fragments
On the lagging strand, RNA primers are removed and replaced with DNA by DNA polymerase I. DNA ligase then seals the nicks between fragments, creating a continuous strand.
Okazaki Fragments: Short DNA segments synthesized on the lagging strand.
DNA Ligase: Forms covalent bonds between DNA fragments.
Proofreading and DNA Repair
Error Correction During Replication
DNA polymerase III has proofreading activity, correcting most errors during replication. Additional repair mechanisms, such as nucleotide excision repair, correct errors missed during synthesis.
Proofreading: DNA polymerase III removes incorrectly paired nucleotides.
Nucleotide Excision Repair: Nuclease removes damaged DNA, DNA polymerase I fills the gap, and DNA ligase seals the strand.
Replication of Chromosome Ends: Telomeres
Telomere Structure and Function
Telomeres are repetitive DNA sequences at the ends of eukaryotic chromosomes. They protect genes from erosion during replication but shorten with each cell division in somatic cells.
Protective Role: Prevent chromosome ends from being recognized as DNA damage and provide a buffer against gene loss.
Telomerase: An enzyme that extends telomeres in germ cells, maintaining chromosome length across generations.
Aging: Telomere shortening is associated with cellular aging and limits the number of cell divisions.
Key Terms and Enzymes to Know
Semiconservative Replication
Origin of Replication
Replication Fork
Single-Strand Binding Protein
Leading Strand, Lagging Strand
Okazaki Fragment
Telomeres
Nucleotide Excision Repair
dNTP
Topoisomerase
Helicase
Primase
DNA Polymerases
Ligase
Telomerase
Summary Table: Major Enzymes in DNA Replication
Enzyme/Protein | Function |
|---|---|
Helicase | Unwinds the DNA double helix |
Single-Strand Binding Protein | Stabilizes unwound DNA strands |
Topoisomerase | Relieves supercoiling ahead of the fork |
Primase | Synthesizes RNA primers |
DNA Polymerase III | Main enzyme for DNA synthesis |
DNA Polymerase I | Removes RNA primers, replaces with DNA |
DNA Ligase | Joins Okazaki fragments |
Telomerase | Extends telomeres in germ cells |
Key Equations and Concepts
Base Pairing: ,
Direction of Synthesis: DNA is synthesized in the 5' to 3' direction.
Energy for Polymerization: Hydrolysis of dNTPs provides energy for DNA synthesis.
Additional info: The above equation shows the addition of a nucleotide to a growing DNA chain, with pyrophosphate (PPi) released as a byproduct.
