 Back
BackDNA Replication, Transcription, and Translation: From Genes to Proteins
Study Guide - Smart Notes

DNA Replication
Overview of DNA Replication
DNA replication is the process by which a cell duplicates its DNA, ensuring that each daughter cell receives an identical copy during cell division. This process is semiconservative, meaning each new DNA molecule consists of one parental and one newly synthesized strand.
Key Enzymes: Helicase, DNA polymerase, primase, ligase, topoisomerase, and single-strand DNA-binding proteins (SSBPs).
Directionality: DNA polymerase synthesizes new DNA in the 5' to 3' direction.
Initiation of Replication
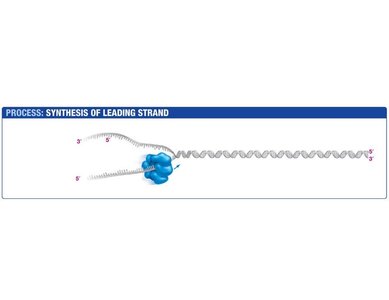
Replication begins at specific locations called origins of replication. Helicase unwinds the DNA double helix, creating a replication fork. SSBPs stabilize the unwound strands, and topoisomerase relieves the tension caused by unwinding.
Helicase: Unwinds the DNA double helix.
SSBPs: Prevent reannealing of single strands.
Topoisomerase: Relieves supercoiling ahead of the fork.
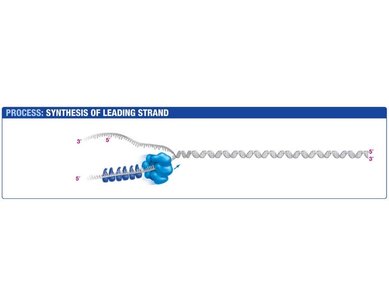
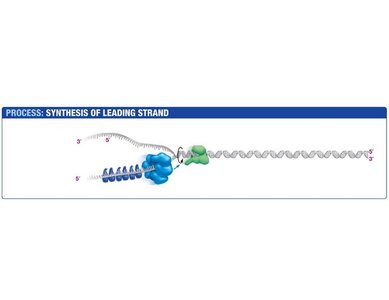
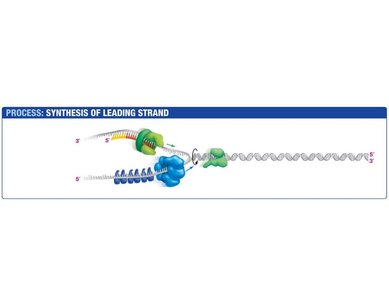
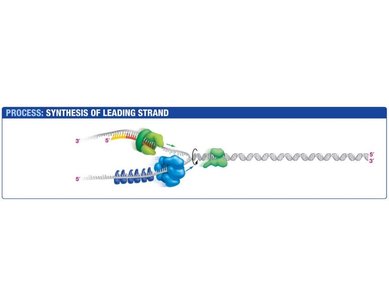
Leading Strand Synthesis
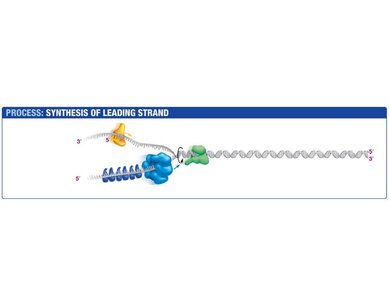
The leading strand is synthesized continuously in the direction of the replication fork. Primase synthesizes a short RNA primer, which is extended by DNA polymerase III.
Primase: Synthesizes RNA primer.
DNA Polymerase III: Extends the primer, synthesizing DNA in the 5' to 3' direction.
Sliding Clamp: Holds DNA polymerase in place.
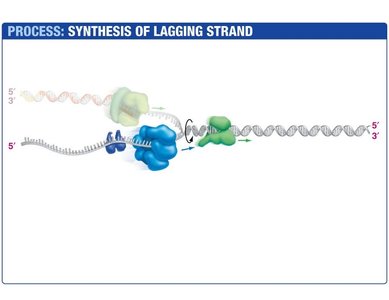
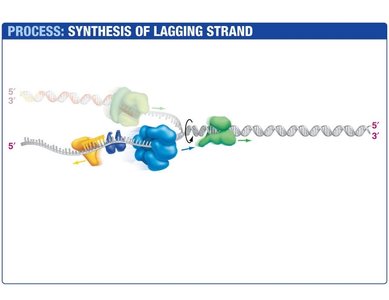
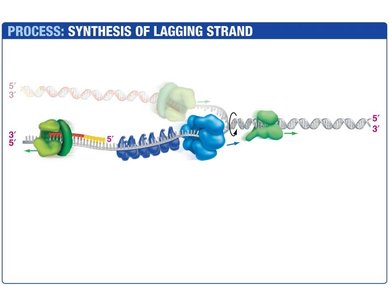
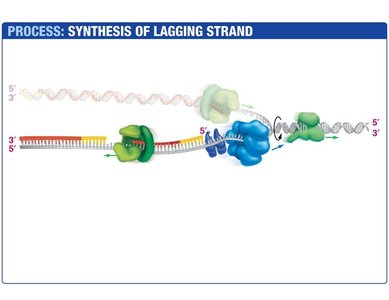
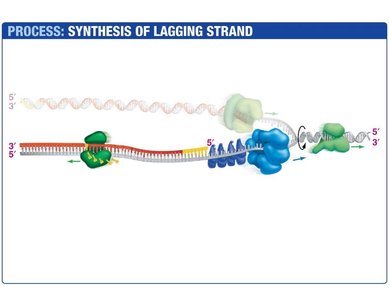
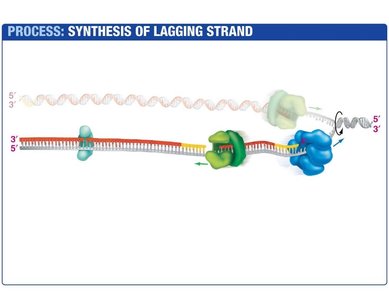
Lagging Strand Synthesis
The lagging strand is synthesized discontinuously, forming short DNA fragments called Okazaki fragments. Each fragment begins with an RNA primer synthesized by primase. DNA polymerase III extends the primer, and DNA polymerase I replaces the RNA with DNA. DNA ligase seals the nicks between fragments.
Okazaki Fragments: Short DNA segments synthesized on the lagging strand.
DNA Ligase: Joins Okazaki fragments by forming phosphodiester bonds.
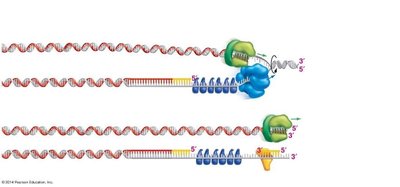
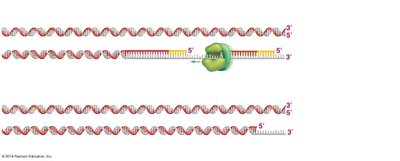
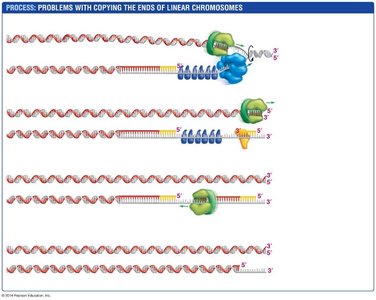
Problems with Replicating Chromosome Ends
Linear chromosomes pose a challenge for replication at their ends (telomeres). The lagging strand cannot be fully replicated, leading to progressive shortening of chromosomes with each cell division.
Telomerase: An enzyme that extends telomeres in certain cell types, preventing loss of genetic information.
Proofreading and DNA Repair
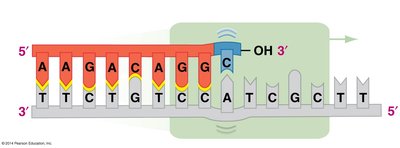
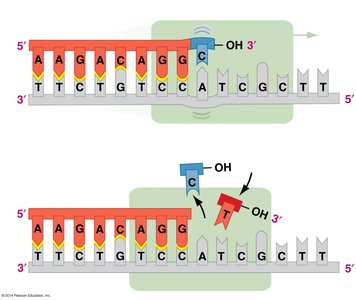
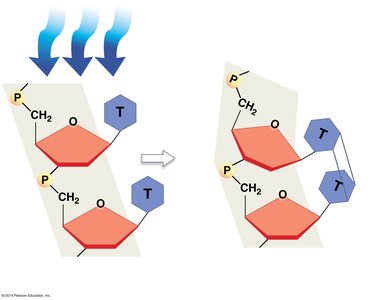
DNA polymerases have proofreading activity, correcting mismatched bases during replication. Additional repair mechanisms, such as nucleotide excision repair, fix DNA damage caused by environmental factors like UV light.
Proofreading: DNA polymerase removes and replaces incorrect nucleotides.
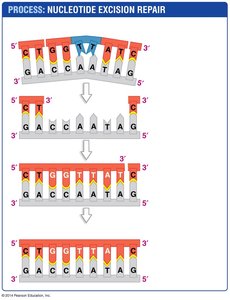
Nucleotide Excision Repair: Damaged DNA is recognized, excised, and replaced with correct nucleotides.
Gene Expression: From DNA to Protein
The Central Dogma of Molecular Biology
The central dogma describes the flow of genetic information: DNA is transcribed into RNA, which is then translated into protein. This process explains how genes determine phenotypes.
Transcription: Synthesis of RNA from a DNA template.
Translation: Synthesis of protein from an mRNA template.
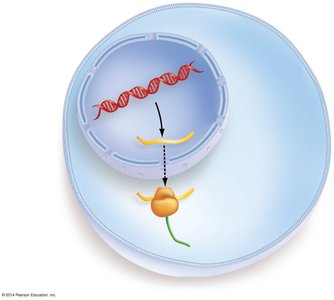
Genotype to Phenotype
Genetic information stored in DNA (genotype) is expressed as proteins, which determine an organism's traits (phenotype). Mutations in DNA can alter protein structure and function, leading to phenotypic changes.
Genotype: The genetic makeup of an organism.
Phenotype: Observable traits resulting from protein activity.
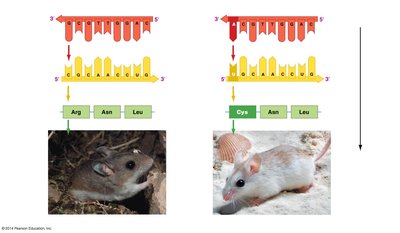
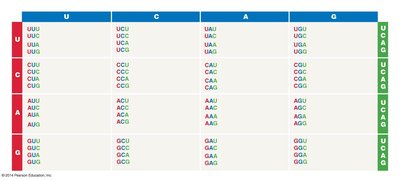
The Genetic Code
The genetic code is a set of rules by which information encoded in mRNA is translated into proteins. Each amino acid is specified by a sequence of three nucleotides (codon).
Codon: A sequence of three RNA bases that codes for a specific amino acid.
Start Codon: AUG (methionine) signals the start of translation.
Stop Codons: UAA, UAG, UGA signal the end of translation.
Transcription: Synthesis of RNA
Transcription is the process by which RNA is synthesized from a DNA template. RNA polymerase binds to the promoter region, unwinds the DNA, and synthesizes a complementary RNA strand.
Promoter: DNA sequence where RNA polymerase binds to initiate transcription.
RNA Polymerase: Enzyme that synthesizes RNA in the 5' to 3' direction.
Sigma Factor: Protein that helps RNA polymerase recognize the promoter in prokaryotes.
RNA Processing in Eukaryotes
In eukaryotes, the primary RNA transcript (pre-mRNA) undergoes several modifications before becoming mature mRNA:
Splicing: Removal of non-coding sequences (introns) and joining of coding sequences (exons).
5' Cap: Addition of a modified guanine nucleotide to the 5' end.
Poly(A) Tail: Addition of a string of adenine nucleotides to the 3' end.
Translation: Synthesis of Protein
Translation is the process by which ribosomes synthesize proteins using mRNA as a template. Transfer RNA (tRNA) molecules bring amino acids to the ribosome, matching their anticodon with codons on the mRNA.
Ribosome: Molecular machine that facilitates the decoding of mRNA into protein.
tRNA: Adapter molecule that carries amino acids and matches them to the mRNA codon via its anticodon.
Peptide Bond: Covalent bond formed between amino acids during protein synthesis.
Summary Table: Key Steps in Gene Expression
Step | Location | Key Enzymes/Structures | Product |
|---|---|---|---|
DNA Replication | Nucleus (eukaryotes) | DNA polymerase, helicase, primase, ligase | Identical DNA molecules |
Transcription | Nucleus (eukaryotes), cytoplasm (prokaryotes) | RNA polymerase, sigma factor (prokaryotes) | mRNA |
RNA Processing | Nucleus (eukaryotes) | Spliceosome, capping and polyadenylation enzymes | Mature mRNA |
Translation | Cytoplasm | Ribosome, tRNA | Protein |
Key Equations and Concepts
Base Pairing Rule:
Number of Possible Codons: possible codons (since there are 4 RNA bases and codons are triplets)
Additional info: The notes above integrate and expand upon the provided slides and images, ensuring a comprehensive, self-contained study guide for college-level biology students.
