 Back
BackDNA Structure and Replication: A Comprehensive Study Guide
Study Guide - Smart Notes

DNA Structure and Function
Basic Components of DNA
DNA (deoxyribonucleic acid) is the hereditary material in almost all living organisms. Its structure and function are fundamental to understanding genetics and cell biology.
Nucleotide: The basic building block of DNA, consisting of a phosphate group, a deoxyribose sugar, and a nitrogenous base.
Four Bases: Adenine (A), Thymine (T), Guanine (G), and Cytosine (C).
Base Pairing: Adenine pairs with Thymine (A-T), and Guanine pairs with Cytosine (G-C).
Hydrogen Bonds: A-T pairs have 2 hydrogen bonds; G-C pairs have 3 hydrogen bonds, making G-C pairs slightly stronger.
Antiparallel: The two strands of DNA run in opposite directions (5' to 3' and 3' to 5').
3' and 5' Ends: Refer to the carbon numbers in the DNA's sugar backbone. The 5' end has a phosphate group attached to the 5' carbon, and the 3' end has a hydroxyl group on the 3' carbon.
Bonds in DNA: Covalent phosphodiester bonds form the backbone (between sugar and phosphate), while hydrogen bonds connect the bases.
DNA vs. RNA: DNA contains deoxyribose sugar and thymine; RNA contains ribose sugar and uracil instead of thymine.
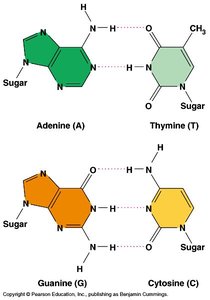
Base Pairing and Purine vs. Pyrimidine
Base pairing in DNA is highly specific and essential for accurate replication and function.
Pyrimidines: Cytosine (C) and Thymine (T) – single-ring structures.
Purines: Adenine (A) and Guanine (G) – double-ring structures.
Pairing Rule: Purines always pair with pyrimidines (A with T, G with C) to maintain a uniform width of the DNA double helix.
Bond Strength: Hydrogen bonds between bases are relatively weak, allowing the strands to separate during replication, but the covalent bonds in the backbone are strong.
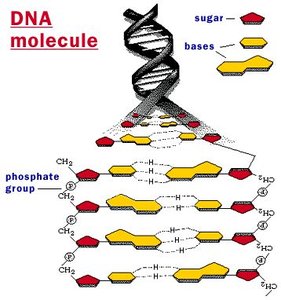
Directionality and DNA Backbone
The directionality of DNA is crucial for its replication and function.
Numbering Carbons: The carbons in the sugar are numbered 1' to 5'. The phosphate group attaches to the 5' carbon, and the next nucleotide attaches at the 3' carbon.
Phosphodiester Bonds: These strong covalent bonds link the 3' carbon of one sugar to the 5' phosphate of the next nucleotide.
Antiparallel Strands: The two DNA strands run in opposite directions, which is essential for replication and base pairing.
DNA Replication
Overview and Purpose
DNA replication is the process by which a cell copies its DNA before cell division, ensuring that each daughter cell receives an identical set of genetic information.
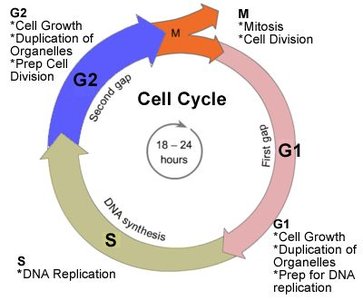
Occurs During: S (synthesis) phase of interphase in the cell cycle.
Location: Nucleus of eukaryotic cells.
Purpose: To provide each new cell with a complete set of DNA after cell division.
Semi-Conservative Replication
DNA replication follows the semi-conservative model, where each new DNA molecule consists of one old (parental) strand and one newly synthesized strand.
Template Mechanism: Each parental strand serves as a template for a new complementary strand.
Result: Two DNA molecules, each with one old and one new strand.

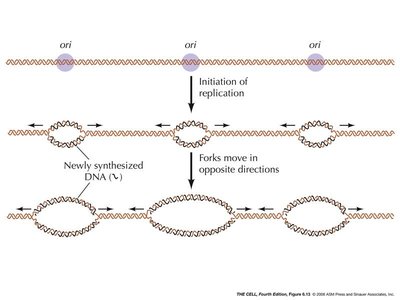
Origins of Replication and Replication Forks
Replication begins at specific locations called origins of replication, where the DNA strands are separated to form replication forks.
Multiple Origins: Eukaryotic chromosomes have multiple origins to speed up replication.
Replication Fork: The Y-shaped region where the DNA is split into two separate strands for copying.

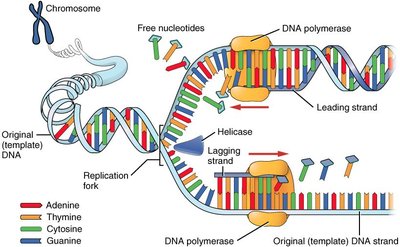
Enzymes and Proteins in DNA Replication
Several enzymes and proteins coordinate the unwinding and synthesis of new DNA strands.
Topoisomerase: Relieves strain ahead of the replication fork by cutting and rejoining DNA.
Helicase: Unwinds and separates the two DNA strands by breaking hydrogen bonds.
Single-Strand Binding Proteins (SSBP): Stabilize and protect single-stranded DNA.
Primase: Synthesizes short RNA primers to provide a starting point for DNA polymerase.
DNA Polymerase: Adds nucleotides to the 3' end of the new DNA strand, synthesizing in a 5' to 3' direction.
Ligase: Joins Okazaki fragments on the lagging strand.
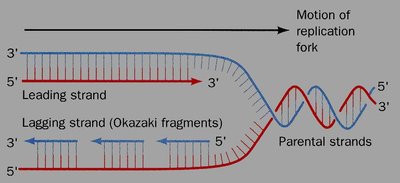
Leading and Lagging Strands
DNA polymerase can only synthesize DNA in the 5' to 3' direction, resulting in continuous and discontinuous synthesis on the two template strands.
Leading Strand: Synthesized continuously toward the replication fork.
Lagging Strand: Synthesized discontinuously away from the fork in short segments called Okazaki fragments, which are later joined by ligase.

Proofreading and DNA Repair
DNA replication is highly accurate, but errors can occur. Cells have mechanisms to correct these mistakes and repair DNA damage.
Proofreading: DNA polymerase checks and corrects mismatched bases during replication.
Exonuclease Activity: Removes incorrectly paired nucleotides.
Repair Enzymes: Additional enzymes, such as DNA ligase and other repair proteins, fix errors and damage (e.g., from UV light).
Continuous Repair: DNA is constantly monitored and repaired to maintain genetic integrity.

Summary Table: DNA vs. RNA
Feature | DNA | RNA |
|---|---|---|
Sugar | Deoxyribose | Ribose |
Bases | A, T, G, C | A, U, G, C |
Strands | Double-stranded | Single-stranded |
Location | Nucleus | Nucleus & Cytoplasm |
Function | Genetic storage | Protein synthesis, gene regulation |
Practice Question Example
Question: If a DNA strand is 30% Adenine, how much Cytosine is present?
Answer: 20% Cytosine. Since A = T and G = C, if A = 30%, T = 30%, leaving 40% for G and C (20% each).
Key Equations and Concepts
Chargaff's Rule: %A = %T and %G = %C in double-stranded DNA.
Direction of Synthesis: DNA polymerase synthesizes new DNA in the 5' to 3' direction.
Phosphodiester Bond Formation:
