 Back
BackDNA Structure and Replication: Foundations and Mechanisms
Study Guide - Smart Notes

DNA Structure and Replication
DNA as a Nucleic Acid
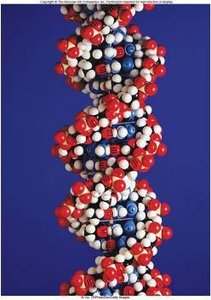
Deoxyribonucleic acid (DNA) is the hereditary material in almost all living organisms. It is a type of nucleic acid, composed of long chains of nucleotides.
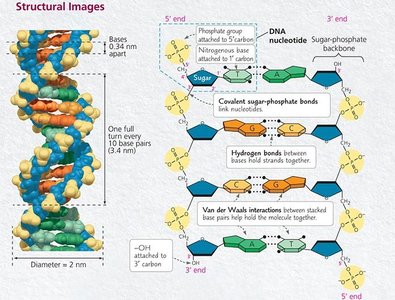
Nucleotides are the building blocks of DNA, each consisting of a phosphate group, a deoxyribose sugar, and a nitrogenous base (adenine [A], cytosine [C], guanine [G], or thymine [T]).
Uracil (U) is found only in RNA, replacing thymine.
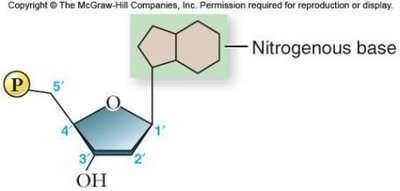
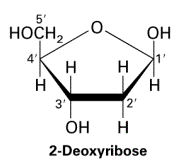
Phosphodiester Bonds and DNA Backbone
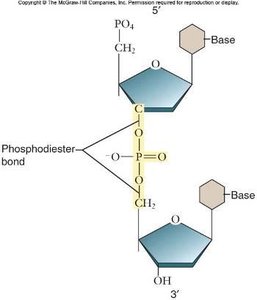
Nucleotides are joined together by phosphodiester bonds, forming the sugar-phosphate backbone of DNA. These covalent bonds link the 5' phosphate group of one nucleotide to the 3' hydroxyl group of the next.
The backbone provides structural stability and directionality (5' to 3').
Double Helix and Base Pairing
DNA consists of two antiparallel strands twisted into a double helix. The strands are held together by hydrogen bonds between complementary bases and by base stacking interactions.
Base pairing: Adenine pairs with thymine (A-T) via two hydrogen bonds; guanine pairs with cytosine (G-C) via three hydrogen bonds.
Base stacking: Van der Waals interactions between stacked bases add further stability.
The double helix has a diameter of about 2 nm, with bases spaced 0.34 nm apart.
Simplified Representations and DNA Sequences
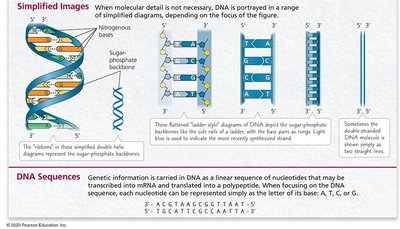
DNA can be depicted in various ways, from detailed atomic models to simplified ladder or ribbon diagrams. The genetic information is encoded in the linear sequence of bases.
DNA sequences are often written as strings of letters (A, T, C, G).
DNA Replication: Models and Mechanisms
Historical Models of DNA Replication
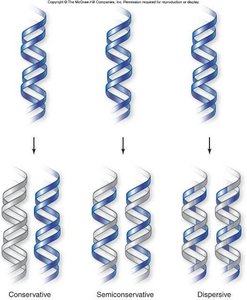
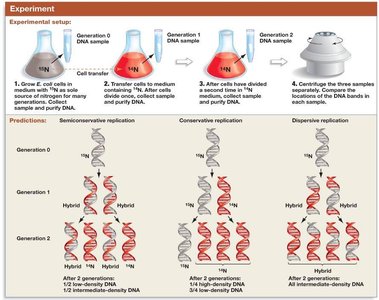
Three models were proposed for DNA replication: conservative, semiconservative, and dispersive. The semiconservative model was confirmed by the Meselson-Stahl experiment.
Conservative: Parental DNA remains intact; new molecule is entirely new DNA.
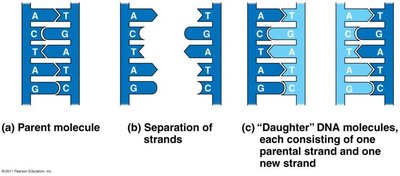
Semiconservative: Each daughter DNA has one parental and one new strand.
Dispersive: Each strand is a mix of old and new DNA segments.
The Meselson-Stahl Experiment
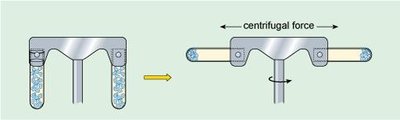
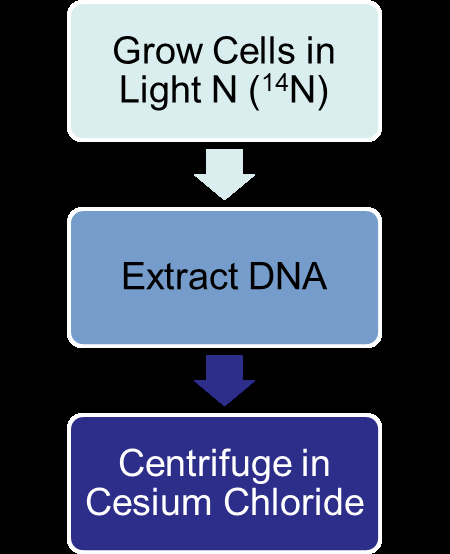
Matthew Meselson and Franklin Stahl used isotopic labeling and density gradient centrifugation to distinguish between the models of DNA replication.
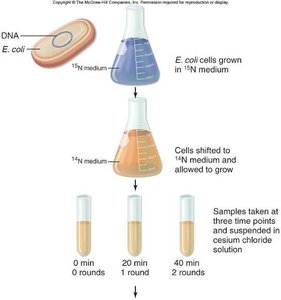
Cells were grown in heavy nitrogen (15N), then shifted to light nitrogen (14N).
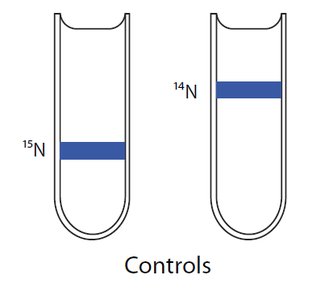
DNA was extracted and centrifuged in cesium chloride to separate by density.
Results showed hybrid DNA after one generation and both hybrid and light DNA after two generations, supporting the semiconservative model.
Mechanism of DNA Replication
Origins of Replication and Replication Forks
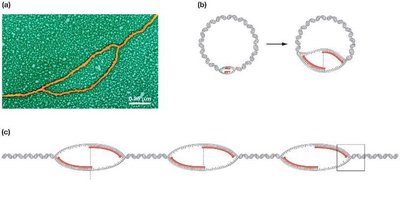
Replication begins at specific sites called origins of replication. Prokaryotes typically have a single origin, while eukaryotes have multiple origins per chromosome. Replication proceeds bidirectionally, forming replication bubbles and forks.
At each fork, parental DNA is unwound and new strands are synthesized.
Initiation: Unwinding the DNA
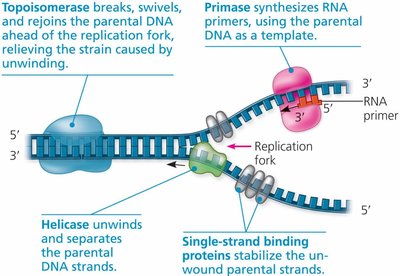
Several proteins are involved in unwinding DNA and preparing it for replication:
Helicase: Unwinds and separates the DNA strands.
Single-strand binding proteins: Stabilize unwound DNA.
Topoisomerase: Relieves strain ahead of the replication fork by breaking and rejoining DNA strands.
Elongation: Synthesizing New DNA Strands
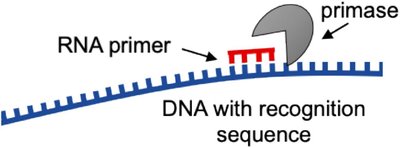
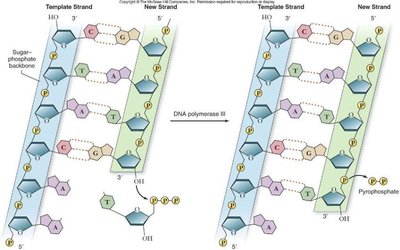
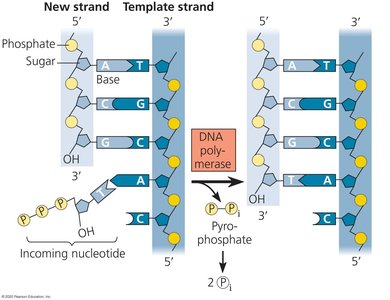
DNA polymerases synthesize new DNA by adding nucleotides to the 3' end of an existing chain. A short RNA primer, synthesized by primase, is required to start the process.
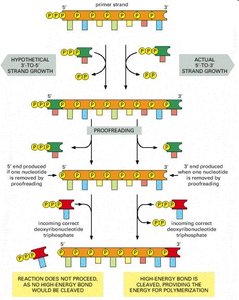
DNA polymerase adds nucleotides in the 5' to 3' direction only.
As each nucleotide is added, two phosphate groups are released as pyrophosphate.
Leading and Lagging Strand Synthesis
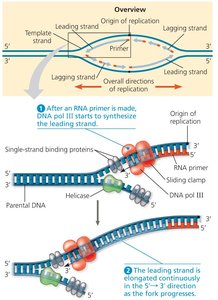
Because DNA polymerase can only synthesize in the 5' to 3' direction, replication is continuous on the leading strand and discontinuous on the lagging strand.
Leading strand: Synthesized continuously toward the replication fork.
Lagging strand: Synthesized away from the fork in short segments called Okazaki fragments, which are later joined by DNA ligase.
Proofreading and Error Correction
DNA polymerases have proofreading activity, correcting errors during replication. If a mismatch occurs, the enzyme removes the incorrect nucleotide and replaces it with the correct one. This high fidelity results in an error rate of about one in 10 billion nucleotides.
Proofreading is more efficient in the 5' to 3' direction due to the energy released by pyrophosphate cleavage.
DNA Repair Mechanisms
Additional repair systems correct errors and damage after replication:
Mismatch repair: Enzymes correct errors missed by DNA polymerase.
Base excision repair: Removes and replaces damaged bases.
Nucleotide excision repair: Removes and replaces damaged nucleotides, such as those caused by UV light.
Defects in repair enzymes can lead to diseases such as hereditary colon cancer.
Termination of DNA Replication
In prokaryotes, replication ends when the forks meet around the circular chromosome. In eukaryotes, replication ends at the convergence of forks or at chromosome ends (telomeres).
Linear chromosomes shorten with each round of replication due to incomplete lagging strand synthesis at the ends.
Telomeres are repetitive sequences at chromosome ends that protect genes from erosion.
Telomerase extends telomeres in germ cells, some stem cells, and cancer cells.
Telomere shortening is associated with cellular aging.
