 Back
BackDNA Structure, Replication, and Chromosome Organization: Study Notes
Study Guide - Smart Notes

DNA: The Genetic Material
Discovery and Significance
DNA (deoxyribonucleic acid) is the molecule that encodes hereditary information and directs the development of biochemical, anatomical, physiological, and behavioral traits in organisms. The double-helical model of DNA was introduced by James Watson and Francis Crick in 1953, revolutionizing our understanding of genetics.
Hereditary Information: DNA is reproduced in all cells and directs cellular functions.
Gene Expression: DNA directs synthesis of messenger RNA (mRNA), which controls protein synthesis.
Nucleic Acids: Structure and Components
Types and Monomers
Nucleic acids are polymers called polynucleotides, made of monomers called nucleotides. There are two main types: DNA and RNA.
DNA: Contains deoxyribose sugar.
RNA: Contains ribose sugar.
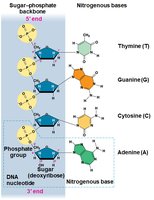
Nucleotide Structure: Each nucleotide consists of a nitrogenous base, a pentose sugar, and a phosphate group.
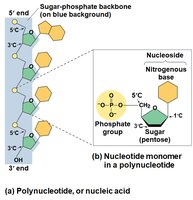
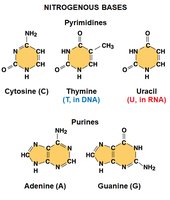
Nitrogenous Bases
Nitrogenous bases are classified into two families:
Pyrimidines: Single ring (cytosine, thymine, uracil)
Purines: Double ring (adenine, guanine)
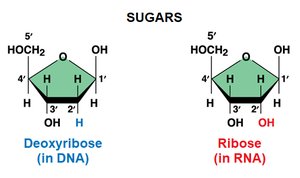
Sugar Component
The pentose sugar differs between DNA and RNA:
Deoxyribose: Found in DNA, lacks an oxygen atom at the 2' position.
Ribose: Found in RNA, has an OH group at the 2' position.
Evidence for DNA as Genetic Material
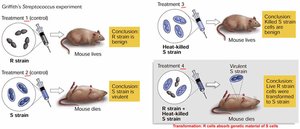
Griffith's Transformation Experiment
Frederick Griffith demonstrated that DNA can transform bacteria, showing that genetic material can be transferred between cells.
Transformation: Living R cells became pathogenic when mixed with heat-killed S cells.
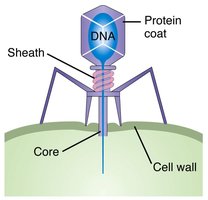
Hershey-Chase Experiment
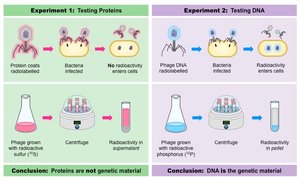
Hershey and Chase used bacteriophages to show that DNA, not protein, is the genetic material that enters cells and directs viral reproduction.
Bacteriophage: Virus that infects bacteria, composed of DNA and protein coat.
Conclusion: Only DNA enters the bacterial cell and provides genetic information.
DNA Structure and Base Pairing
Chargaff's Rules
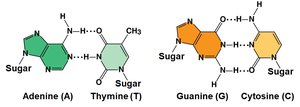
Erwin Chargaff found that the base composition of DNA varies between species, but in any species, the amount of adenine (A) equals thymine (T), and guanine (G) equals cytosine (C).
Base Pairing: A pairs with T, G pairs with C.
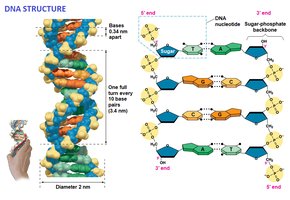
Double Helix Model
Rosalind Franklin's X-ray crystallography images enabled Watson and Crick to deduce the helical structure of DNA, with two antiparallel strands and paired nitrogenous bases in the interior.
Antiparallel: Strands run in opposite directions (5' to 3' and 3' to 5').
Hydrogen Bonds: Hold base pairs together.

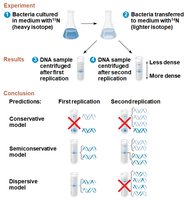
DNA Replication: Mechanisms and Models
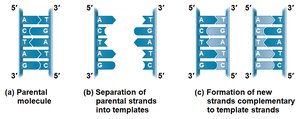
Base Pairing to a Template Strand
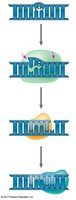
During DNA replication, each strand serves as a template for building a new complementary strand. The process is highly accurate and involves many proteins and enzymes.
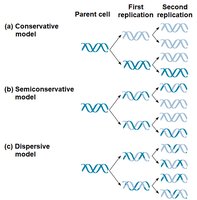
Semiconservative Model: Each new DNA molecule has one old strand and one new strand.
Competing Models: Conservative (parent strands rejoin), dispersive (mix of old and new).
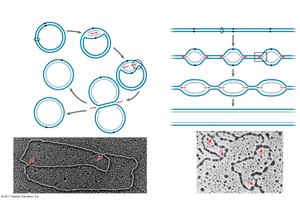
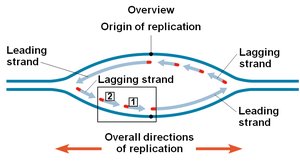
Origins of Replication
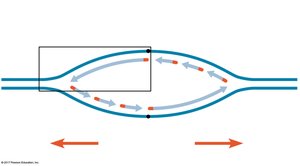
Replication begins at specific sites called origins of replication, forming replication bubbles and forks. Eukaryotic chromosomes have multiple origins, while prokaryotes typically have one.
Replication Fork: Y-shaped region where new DNA strands are synthesized.
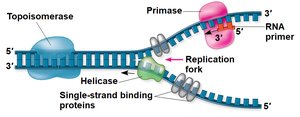
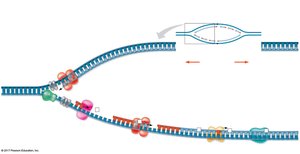
Key Enzymes and Proteins
Several enzymes and proteins are essential for DNA replication:
Helicase: Unwinds the DNA double helix.
Single-Strand Binding Proteins: Stabilize single-stranded DNA.
Topoisomerase: Relieves strain ahead of the replication fork.
Primase: Synthesizes RNA primers.
DNA Polymerase: Catalyzes synthesis of new DNA.
DNA Ligase: Joins Okazaki fragments on the lagging strand.
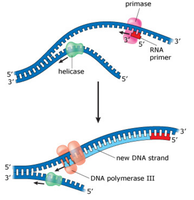
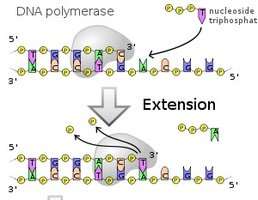
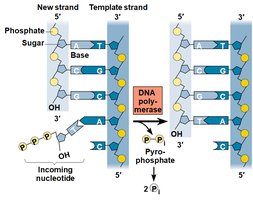
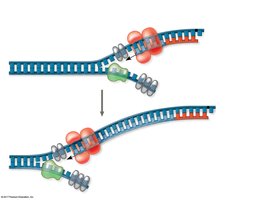
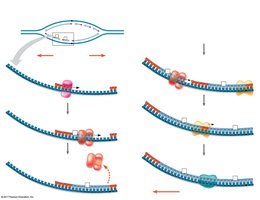
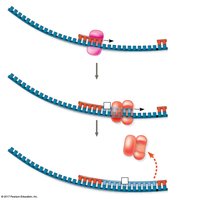
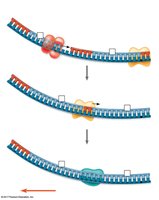
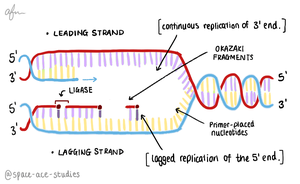
Antiparallel Elongation
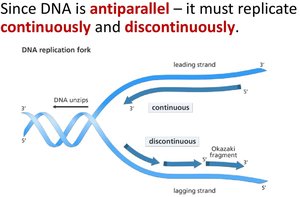
DNA polymerases add nucleotides only to the free 3' end, so new strands elongate in the 5' to 3' direction. This results in continuous synthesis on the leading strand and discontinuous synthesis on the lagging strand (Okazaki fragments).
Leading Strand: Synthesized continuously toward the replication fork.
Lagging Strand: Synthesized discontinuously away from the fork.
Replication Proteins Table
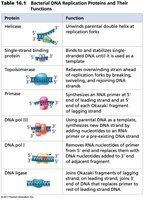
The following table summarizes the main proteins involved in bacterial DNA replication and their functions:
Protein | Function |
|---|---|
Helicase | Unwinds parental double helix at replication forks |
Single-strand binding protein | Stabilizes unwound DNA |
Topoisomerase | Relieves strain ahead of replication fork |
Primase | Synthesizes RNA primer at 5' end of leading strand and each Okazaki fragment |
DNA pol III | Uses parental DNA as template, synthesizes new DNA strand |
DNA pol I | Removes RNA primer and replaces with DNA |
DNA ligase | Joins Okazaki fragments and replaces primer with DNA |
Proofreading, Repair, and Mutation
DNA Proofreading and Repair
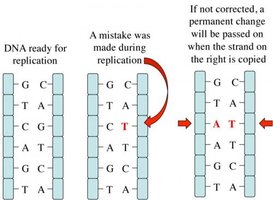
DNA polymerases proofread newly synthesized DNA, correcting errors. Additional repair mechanisms include mismatch repair and nucleotide excision repair, which remove and replace damaged DNA segments.
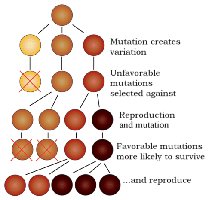
Mutation: Sequence changes that can be passed to the next generation, providing genetic variation for evolution.
Replicating the Ends of DNA Molecules
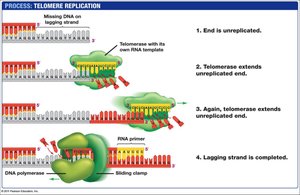
Telomeres and Telomerase
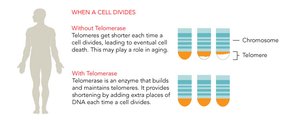
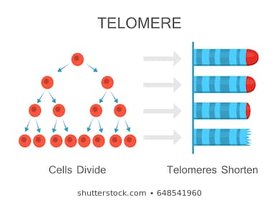
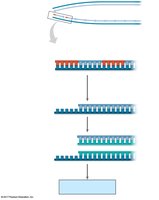
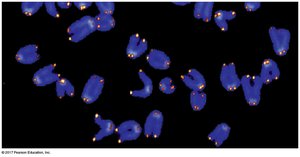
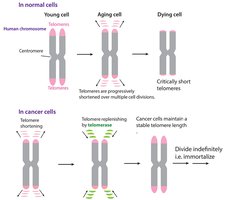
Linear DNA in eukaryotic chromosomes faces the problem of incomplete replication at the ends (telomeres). Telomeres are repetitive nucleotide sequences that protect genes from erosion. Telomerase is an enzyme that extends telomeres in germ cells, preventing loss of essential genetic information.
Telomere Shortening: Associated with aging and cell death.
Telomerase Activity: Found in germ cells and cancer cells, allowing indefinite cell division.

Chromosome Structure and DNA Packing
Chromatin Organization
Eukaryotic chromosomes consist of linear DNA molecules packed with proteins, forming chromatin. Chromatin undergoes multilevel packing to fit into the nucleus, with histones responsible for the first level of packing.
Nucleosome: Basic unit of DNA packaging, composed of histone proteins and DNA.
Histone Tails: Involved in regulation of gene expression.
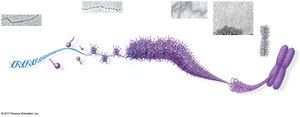
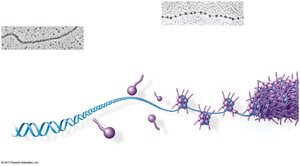
Chromatin States
Chromatin can be loosely packed (euchromatin) or highly condensed (heterochromatin). Dense packing makes gene expression difficult in those regions.
Euchromatin: Loosely packed, accessible for transcription.
Heterochromatin: Highly condensed, less accessible.
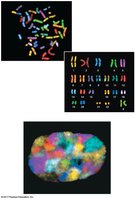
Size Hierarchy of Chromosome Structures
The following order describes the size hierarchy from smallest to largest:
Histone
Nucleosome
30-nm fiber
Looped domain
Metaphase chromosome
*Additional info: These notes expand on the original content by providing definitions, context, and examples for each topic, ensuring completeness and academic quality for exam preparation.*
