 Back
BackEukaryotic Gene Expression and Mutations: Structure, Function, and Analysis
Study Guide - Smart Notes

Eukaryotic Gene Expression
Structure of a Eukaryotic Gene
Eukaryotic genes contain several distinct regions that regulate and encode protein synthesis. Understanding these regions is essential for interpreting gene function and mutation effects.
Promoter Region: The promoter is a DNA sequence where RNA polymerase binds to initiate transcription. In the Gene Explorer simulation, the promoter sequence is 5'-TATAA-3'.
Terminator Region: The terminator signals the end of transcription. In the simulation, the terminator sequence is 5'-GGGGG-3'.
Exons: Exons are coding regions that remain in the mature mRNA after splicing and are translated into protein.
Introns: Introns are non-coding regions removed during mRNA splicing. Intron boundaries are marked by 5'-GTGCG-3' (start) and 5'-CAAAG-3' (end).
Start Codon: The start codon (ATG) marks the beginning of translation.
Stop Codons: Stop codons (TGA, TAA, TAG) signal the end of translation.
Leader and Trailing Sequences: Upstream leader sequences precede the start codon, while downstream trailing sequences follow the stop codon and are transcribed but not translated.
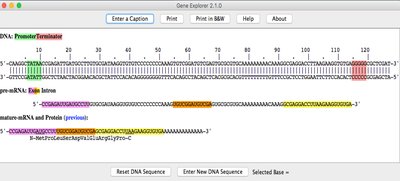
Example: The Gene Explorer output visually maps these regions, showing how DNA is transcribed to pre-mRNA, processed to mature mRNA, and translated to protein.
Transcription and Translation
Transcription and translation are the two main steps in gene expression, converting genetic information into functional proteins.
Transcription: The process by which RNA polymerase synthesizes a complementary RNA strand from the DNA template. Transcription starts at the promoter and ends at the terminator.
Pre-mRNA Processing: Eukaryotic pre-mRNA undergoes splicing to remove introns and join exons. A 5' cap and a poly-A tail are added to enhance stability and translation efficiency.
Translation: The mature mRNA is decoded by ribosomes to synthesize a polypeptide chain, starting at the start codon and ending at the stop codon.
Key Terms:
mRNA Splicing: Removal of introns and joining of exons.
5' Cap: Modified guanine nucleotide added to the 5' end of mRNA.
Poly-A Tail: Sequence of adenines added to the 3' end of mRNA.
Gene Mapping and Coding Regions
Gene mapping involves identifying the positions of promoters, exons, introns, start and stop codons, and terminators within a gene sequence. Coding regions are the portions of exons that are translated into protein.
Promoter: Typically located upstream of the coding region.
Exons: Labeled boxes in gene maps; coding regions are often indicated by hashed areas.
Introns: Bland areas between exons; not translated.
Start Codon: Marks the beginning of the coding region within an exon.
Stop Codon: Marks the end of the coding region within an exon.


Example: Gene maps visually represent the structure of a gene, highlighting the positions of exons, introns, and coding regions.
Mutations and Their Effects
Types of Mutations
Mutations are changes in the DNA sequence that can affect gene expression and protein function. The effects depend on the mutation type and its location.
Base Substitution (Point Mutation): A single nucleotide pair is changed (e.g., AT to GC).
Missense Mutation: Results in a different amino acid in the polypeptide chain.
Silent Mutation: Results in the same amino acid; no effect on protein function.
Nonsense Mutation: Results in a stop codon, prematurely ending translation.
Frameshift Mutation: Addition or deletion of bases alters the reading frame, potentially changing many amino acids.
Effects of Mutations:
Mutations in introns are often silent.
Mutations in exons can alter the amino acid sequence and protein function.
Mutations affecting start or stop codons can disrupt translation initiation or termination.
Example: Changing a base in the coding region may result in a missense or nonsense mutation, while changes in introns may have no effect.
Mutation Analysis Using Gene Explorer
Gene Explorer allows simulation of mutations and analysis of their effects on mRNA and protein sequences. By editing DNA bases, students can observe how mutations alter gene expression.
Mutation 1: Change C/G at position 58 to T/A. Analyze the effect on the amino acid sequence.
Mutation 2: Delete T/A at position 26. Observe changes in the reading frame and protein.
Mutation 3: Change A/T at position 51 to T/A. Determine if the mutation is silent, missense, or nonsense.
Mutation 4: Change T/A at position 21 to G/C. Assess the impact on gene structure and expression.
Mutation Table: (Example format for analysis)
Mutation | Region | Type | Effect on Amino Acid Sequence |
|---|---|---|---|
1 | Exon | Missense | Changes one amino acid |
2 | Exon | Frameshift | Alters reading frame, changes multiple amino acids |
3 | Exon | Silent/Missense | May change or not change amino acid |
4 | Promoter/Exon | Missense | May affect transcription or protein sequence |
Additional info: The exact effects depend on the position and nature of the mutation. Mutations in regulatory regions (promoter, terminator) can affect transcription, while those in coding regions affect translation.
Bioinformatics and Gene Analysis
Bioinformatics Tools
Bioinformatics combines computational tools and biological data to analyze gene sequences, mutations, and protein functions. The National Center for Biotechnology Information (NCBI) provides databases and resources for gene analysis.
NCBI: Offers DNA and protein sequence databases, visualization tools, and the Online Mendelian Inheritance in Man (OMIM) catalog.
Gene Explorer: Simulates gene structure, expression, and mutation effects for educational purposes.
Example: Scientists use bioinformatics to compare gene sequences, identify mutations, and study genetic disorders.
Key Equations and Concepts
Transcription and Translation Equations
Transcription and translation follow the genetic code, converting nucleotide sequences to amino acids.
Transcription: DNA (template strand) → mRNA (complementary sequence)
Translation: mRNA codons → Amino acids (using the genetic code)
Example Equation:
Additional info: The genetic code is universal, but eukaryotic genes are more complex due to introns, exons, and regulatory sequences.
