 Back
BackGene Expression and Transcriptional Regulation in Prokaryotes and Eukaryotes
Study Guide - Smart Notes

Transcriptional Regulation in Prokaryotes
Overview of Gene Regulation in Bacteria
Gene expression in bacteria is tightly regulated to adapt to changing environmental conditions. Bacteria can turn genes on or off in response to environmental signals, ensuring efficient use of resources.
Gene regulation involves mechanisms that increase or decrease the production of specific gene products (proteins or RNAs).
Regulation can occur at multiple stages: transcription, translation, and post-translation.
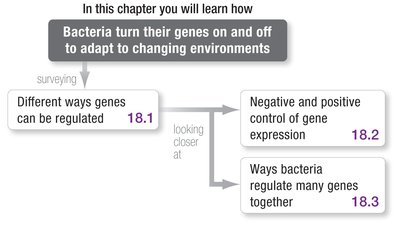
Negative and Positive Control of Gene Expression
Bacteria use both negative and positive regulatory systems to control gene expression:
Negative control: A repressor protein binds to DNA and blocks transcription.
Positive control: An activator protein increases the rate of transcription.
Example: The lac operon in Escherichia coli is a classic model of negative regulation, where the presence or absence of lactose determines whether the operon is transcribed.
Coordinated Regulation of Multiple Genes
Bacteria often organize genes with related functions into operons, allowing coordinated regulation by a single promoter and regulatory region.
Operon: A cluster of genes under the control of a single promoter and operator, transcribed as a single mRNA.
Example: The lac operon includes lacZ, lacY, and lacA genes, all involved in lactose metabolism.
Transcriptional Regulation in Eukaryotes
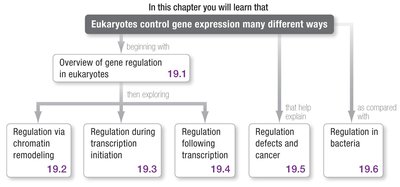
Overview of Gene Regulation in Eukaryotes
Eukaryotic gene expression is more complex than in prokaryotes, involving multiple levels of regulation from chromatin structure to post-translational modifications.
Gene regulation occurs at the levels of chromatin remodeling, transcription initiation, RNA processing, mRNA stability, translation, and post-translational modification.
Chromatin Structure and Remodeling
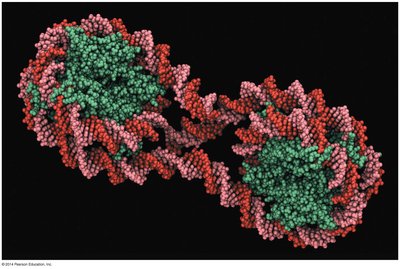
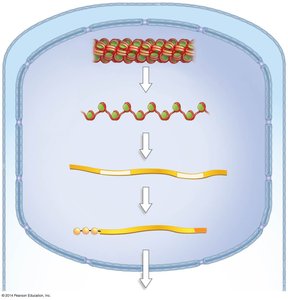
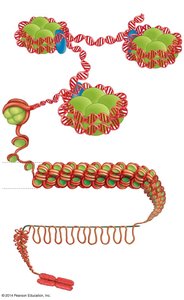
In eukaryotes, DNA is packaged with proteins into chromatin, which can be modified to regulate gene accessibility.
Nucleosome: The basic unit of chromatin, consisting of DNA wrapped around histone proteins.
Chromatin remodeling: Chemical modifications (e.g., acetylation, methylation) alter chromatin structure, making DNA more or less accessible for transcription.
Regulation During Transcription Initiation
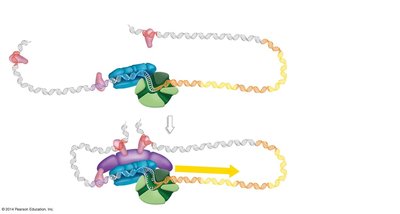
Transcription initiation in eukaryotes involves the assembly of multiple proteins at the promoter, including transcription factors and RNA polymerase II.
Enhancers and promoter-proximal elements are DNA sequences that increase the likelihood of transcription initiation.
Basal transcription factors are required for the assembly of the transcription initiation complex.
Mediator is a protein complex that facilitates interactions between regulatory proteins and RNA polymerase II.
Epigenetic Regulation
Epigenetic modifications, such as DNA methylation and histone modification, can be inherited and affect gene expression without altering the DNA sequence.
Environmental factors, such as nutrition, can influence epigenetic marks and gene expression in offspring.
Example: Offspring of protein-deprived mothers may have abnormal histone modifications and altered gene expression.
DNA Replication and Gene Expression
DNA Replication
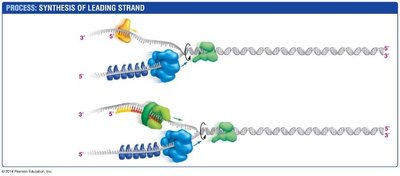
DNA replication is the process by which a cell copies its DNA before cell division. It involves the synthesis of a new strand complementary to each original strand.
Leading strand: Synthesized continuously in the 5' to 3' direction.
Lagging strand: Synthesized discontinuously as Okazaki fragments.
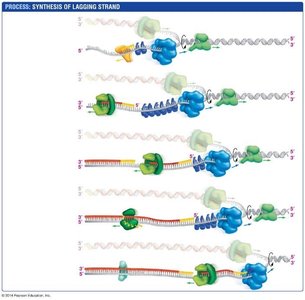
Transcription: Making RNA from DNA
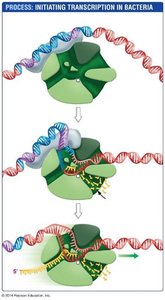
Transcription is the process by which RNA is synthesized from a DNA template. In bacteria, RNA polymerase binds to the promoter and synthesizes mRNA.
Promoter: DNA sequence where RNA polymerase binds to initiate transcription.
mRNA: Messenger RNA, which carries the genetic code from DNA to the ribosome for protein synthesis.
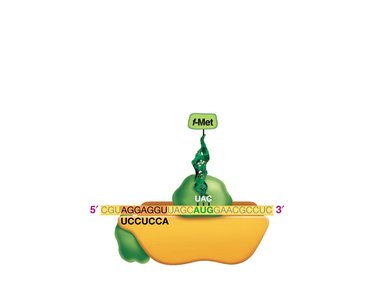
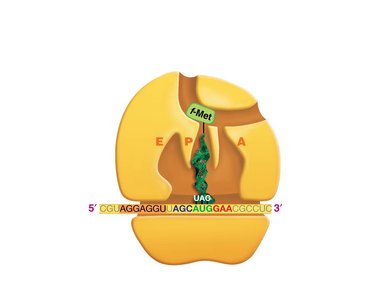
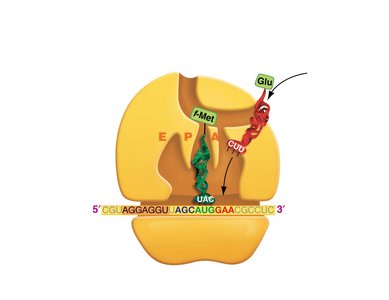
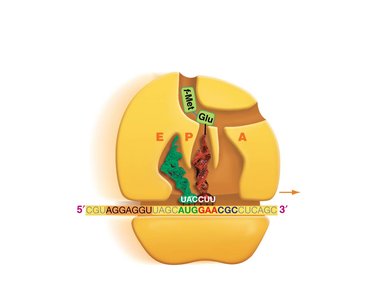
Translation: Protein Synthesis
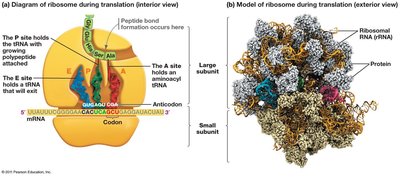
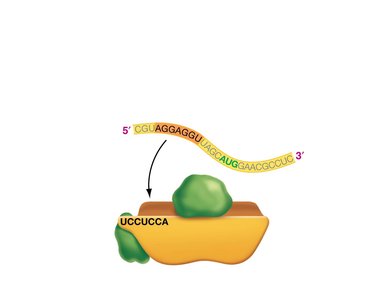
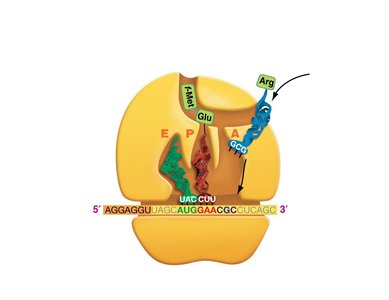
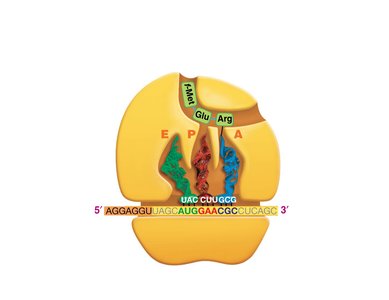
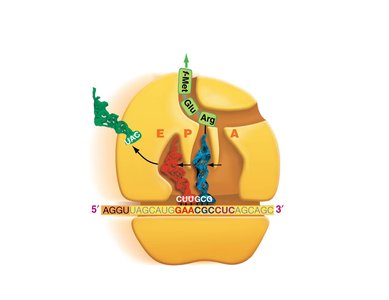
Translation is the process by which ribosomes synthesize proteins using mRNA as a template. tRNAs bring amino acids to the ribosome, matching codons in the mRNA with their anticodons.
Codon: A sequence of three nucleotides in mRNA that specifies an amino acid.
tRNA: Transfer RNA, which carries amino acids to the ribosome and matches them to the mRNA codon via its anticodon.
Ribosome: The molecular machine that facilitates the assembly of amino acids into a polypeptide chain.

Summary Table: Key Differences in Gene Regulation
Feature | Prokaryotes | Eukaryotes |
|---|---|---|
DNA Packaging | Not complexed with histones | Packaged into chromatin with histones |
Regulatory Elements | Promoters, operators | Promoters, enhancers, silencers |
Gene Organization | Operons (polycistronic mRNA) | Mostly monocistronic mRNA |
Transcription & Translation | Coupled (occur simultaneously) | Separated by nuclear envelope |
Epigenetic Regulation | Rare | Common (DNA/histone modifications) |
