 Back
BackGene Expression: From Gene to Protein & Regulation of Gene Expression (Chapters 14 & 15)
Study Guide - Smart Notes

Gene Expression: From Gene to Protein
One Gene, One Polypeptide Hypothesis
The "one gene, one polypeptide" hypothesis states that each gene within DNA codes for a single polypeptide chain, which may function independently or as part of a larger protein complex. This concept evolved from the earlier "one gene, one enzyme" hypothesis, reflecting the discovery that not all proteins are enzymes and that some proteins are composed of multiple polypeptide chains.
Gene: A segment of DNA that encodes instructions for making a specific polypeptide or RNA molecule.
Polypeptide: A chain of amino acids linked by peptide bonds; may function as a protein or as a subunit of a protein.
Example: Hemoglobin consists of four polypeptide subunits, each encoded by a separate gene.
Key Terminology in Gene Expression
Transcription: The process by which a DNA sequence is copied into messenger RNA (mRNA).
Translation: The process by which the sequence of an mRNA molecule is used to assemble amino acids into a polypeptide.
mRNA (messenger RNA): The RNA copy of a gene that carries genetic information from the nucleus to the ribosome.
Ribosome: The molecular machine that reads mRNA and synthesizes polypeptides.
Primary transcript: The initial RNA product synthesized from a gene, which may undergo processing to become mature mRNA.
Codon: A sequence of three nucleotides in mRNA that specifies a particular amino acid.
Template strand: The DNA strand that is read by RNA polymerase to synthesize RNA.
Coding strand: The DNA strand whose sequence matches the mRNA (except T is replaced by U).
Promoter: A DNA sequence where RNA polymerase binds to initiate transcription.
RNA polymerase: The enzyme that synthesizes RNA from a DNA template.
5’ cap: A modified guanine nucleotide added to the 5’ end of eukaryotic mRNA for stability and ribosome binding.
Poly-A tail: A string of adenine nucleotides added to the 3’ end of eukaryotic mRNA for stability and export from the nucleus.
RNA splicing: The removal of introns and joining of exons in eukaryotic mRNA.
Exons: Coding regions of a gene that remain in mature mRNA.
Introns: Non-coding regions removed during RNA splicing.
Mutagen: An agent that causes changes (mutations) in DNA.
Steps of Transcription and Translation
Transcription:
Initiation: RNA polymerase binds to the promoter region (often containing a TATA box in eukaryotes).
Elongation: RNA polymerase synthesizes the RNA strand by adding nucleotides complementary to the DNA template strand.
Termination: Transcription ends when RNA polymerase reaches a terminator sequence.
Translation:
Initiation: The small ribosomal subunit binds to the mRNA, and the initiator tRNA pairs with the start codon (AUG).
Elongation: tRNAs bring amino acids to the ribosome, matching codons in the mRNA with anticodons in the tRNA.
Termination: When a stop codon is reached, the polypeptide is released from the ribosome.
Locations of Transcription and Translation
Transcription: Occurs in the nucleus of eukaryotic cells.
Translation: Occurs in the cytoplasm at ribosomes.
The Central Dogma of Molecular Biology
The central dogma describes the flow of genetic information: DNA → RNA → Protein. This concept is foundational, though exceptions exist (e.g., some genes code for functional RNAs, and alternative splicing allows one gene to produce multiple polypeptides).
Example: Retroviruses can reverse transcribe RNA into DNA.
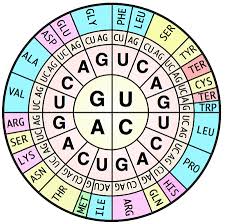
Codons and the Genetic Code
Codons are triplets of nucleotides in mRNA that specify amino acids. The genetic code is redundant (more than one codon can specify the same amino acid) but not ambiguous (each codon specifies only one amino acid).
Start codon: AUG (codes for methionine, Met)
Stop codons: UAA, UAG, UGA (signal termination of translation)
Redundancy: Multiple codons can code for the same amino acid (e.g., GGU, GGC, GGA, GGG all code for glycine).
Promoters and RNA Polymerase
Promoter: Essential for initiating transcription; contains specific sequences (like the TATA box in eukaryotes) recognized by transcription factors and RNA polymerase.
RNA polymerase: Synthesizes RNA from a DNA template; unlike DNA polymerase, it does not require a primer and can initiate synthesis de novo.
RNA Splicing and Its Importance
Splicing: Removes introns and joins exons, allowing for the production of mature mRNA. Alternative splicing can generate multiple proteins from a single gene.
Role of Ribosome and tRNA in Translation
Ribosome: Facilitates the decoding of mRNA into a polypeptide chain.
tRNA: Brings the correct amino acid to the ribosome by matching its anticodon with the mRNA codon.
Targeting Polypeptides to the ER
Some polypeptides have signal sequences that direct them to the endoplasmic reticulum (ER) for further processing or secretion.
Types of Mutations
Point mutations: Change a single nucleotide (e.g., silent, missense, nonsense mutations).
Insertions/Deletions: Add or remove nucleotides, potentially causing frameshifts.
Effect: Mutations can alter gene expression and protein function, sometimes causing disease.
Regulation of Gene Expression
Importance of Gene Control
Gene regulation allows cells to respond to environmental changes and differentiate into specialized cell types. Both prokaryotes and eukaryotes tightly control gene expression to conserve resources and ensure proper function.
Operons in Prokaryotes
Operon: A cluster of genes under the control of a single promoter and operator, allowing coordinated regulation (e.g., lac operon in E. coli).
Benefit: Efficient regulation of genes with related functions.
Repression vs. Induction
Repression: Gene expression is turned off by a repressor protein (e.g., trp operon).
Induction: Gene expression is turned on in response to a specific molecule (e.g., lac operon induced by lactose).
The lac Operon
Operator: DNA segment where a repressor binds to block transcription.
Promoter: Site where RNA polymerase binds to initiate transcription.
Repressor gene: Encodes a protein that can bind to the operator to inhibit transcription.
RNA polymerase: Enzyme that transcribes the operon's genes.
Effect of glucose: High glucose inhibits lac operon expression even if lactose is present, conserving energy (catabolite repression).
Points of Gene Regulation
Gene expression can be regulated at multiple stages: chromatin structure, transcription, RNA processing, mRNA stability, translation, and protein modification/degradation.
Nucleosomes and Chromatin Structure
Nucleosome: DNA wrapped around histone proteins; basic unit of chromatin.
Methylation: Addition of methyl groups to DNA or histones, often repressing gene expression by tightening chromatin structure.
Eukaryotic Transcription Factors
General transcription factors: Required for transcription of all genes; include the TATA-binding protein (TBP), which recognizes the TATA box in promoters.
Specific transcription factors: Regulate expression of particular genes in response to signals.
Alternative Splicing
Allows a single gene to produce multiple mRNA variants and thus different proteins, increasing proteomic diversity.
Control of Translation and mRNA Longevity
Translation can be regulated by proteins or small RNAs that bind mRNA, and by the stability (half-life) of the mRNA itself.
Ubiquitination and Proteasome Function
Ubiquitination: The attachment of ubiquitin molecules to a protein, marking it for degradation.
Proteasome: A large protein complex that degrades and recycles ubiquitinated proteins, regulating protein levels and quality in the cell.
