 Back
BackGene Expression: From Gene to Protein – Transcription, Translation, and Mutations
Study Guide - Smart Notes

Gene Expression: From Gene to Protein
Overview of Gene Expression
Gene expression is the process by which information from a gene is used to synthesize functional gene products, typically proteins. This process involves two main stages: transcription and translation. The genotype refers to the genetic makeup, while the phenotype is the observable trait.
Transcription: Synthesis of RNA from a DNA template.
Translation: Synthesis of a polypeptide (protein) using the information in mRNA.
The central dogma of molecular biology:
Historical Foundations of Gene-Protein Relationships
Garrod and Inborn Errors of Metabolism
Sir Archibald Garrod was the first to propose a connection between genes, proteins, and metabolic diseases. He suggested that inherited disorders could result from the absence of specific enzymes in metabolic pathways, coining the term "inborn errors of metabolism." An example is alkaptonuria, a disorder causing black urine due to the lack of an enzyme to break down alkapton.

One Gene-One Enzyme/Protein Hypothesis
Beadle and Tatum's experiments with Neurospora (a type of mold) led to the "one gene-one enzyme" hypothesis, later refined to "one gene-one protein." They demonstrated that mutations in genes could inactivate enzymes required for synthesizing essential nutrients, linking genes directly to protein function.

Principles of Transcription and Translation
Transcription and Translation in Prokaryotes vs. Eukaryotes
Transcription is the synthesis of RNA using DNA as a template, producing messenger RNA (mRNA). Translation is the synthesis of a polypeptide at ribosomes using the mRNA sequence. In prokaryotes, translation can begin before transcription is complete, while in eukaryotes, the nuclear envelope separates these processes, and RNA transcripts undergo processing before translation.
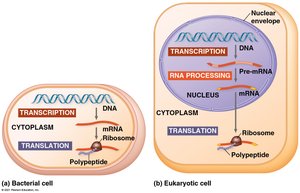
Transcription: DNA-Directed Synthesis of RNA
Transcription involves copying the DNA sequence of a gene into RNA. The template strand of DNA is used to order the sequence of RNA nucleotides. The coding strand is the non-template strand, which has the same sequence as the RNA (except T is replaced by U).
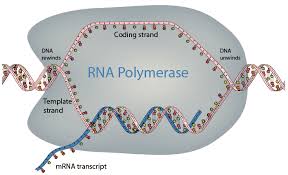


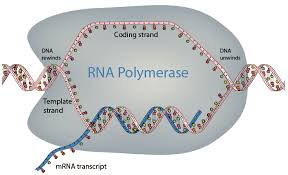
Mechanism of Transcription
RNA synthesis is catalyzed by RNA polymerase, which binds to a specific DNA sequence called the promoter. The enzyme pries apart the DNA strands and joins RNA nucleotides complementary to the DNA template. RNA polymerase does not require a primer, and uracil (U) replaces thymine (T) in RNA.
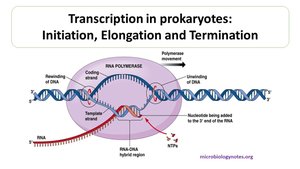
Stages of Transcription: Initiation, Elongation, Termination
Transcription proceeds in three stages:
Initiation: Promoters signal the start point; transcription factors help RNA polymerase bind and initiate transcription.
Elongation: RNA polymerase moves along the DNA, adding nucleotides to the 3' end of the growing RNA molecule. Multiple RNA polymerases can transcribe a gene simultaneously.
Termination: The polymerase stops transcription at the terminator sequence, releasing the mRNA.
Transcription in Eukaryotes: RNA Processing
In eukaryotes, the promoter often contains a TATA box. The initial RNA transcript (pre-mRNA) undergoes processing:
The 5' end receives a modified guanine nucleotide "cap".
The 3' end receives a poly-A tail (50–250 adenine nucleotides).
These modifications protect mRNA and aid in export and translation.
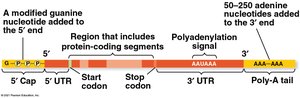
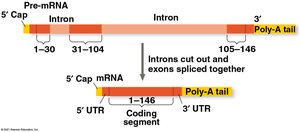
Split Genes and RNA Splicing
Eukaryotic genes contain introns (noncoding regions) and exons (coding regions). RNA splicing removes introns and joins exons to produce mature mRNA. This process is carried out by spliceosomes (complexes of proteins and small RNAs) or by ribozymes (catalytic RNAs).
Translation: RNA-Directed Synthesis of Polypeptides
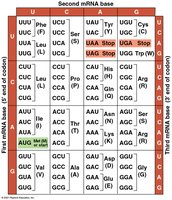
The Genetic Code
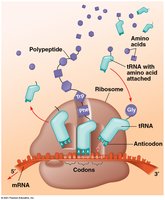
The genetic code consists of codons, sequences of three mRNA nucleotides that specify amino acids. The code is nearly universal among organisms, supporting the idea of a common evolutionary origin.
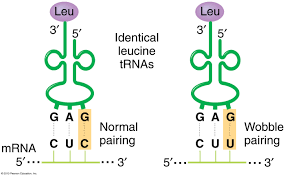
tRNA and the Role of Anticodons
Transfer RNA (tRNA) molecules carry specific amino acids and have anticodons that base-pair with mRNA codons. Accurate translation requires correct matching between tRNA anticodon and mRNA codon, and between tRNA and its amino acid (catalyzed by aminoacyl-tRNA synthetase). Wobble pairing at the third codon position allows some tRNAs to recognize multiple codons.
Ribosomes and Sites of Translation
Ribosomes facilitate the coupling of tRNA anticodons with mRNA codons. Eukaryotic ribosomes are larger and more complex than prokaryotic ones. Each ribosome has three binding sites for tRNA:
P site: Holds tRNA with the growing polypeptide chain.
A site: Holds tRNA with the next amino acid.
E site: Exit site for discharged tRNAs.
Initiation, Elongation, and Termination of Translation
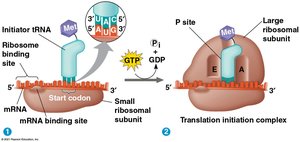
Translation proceeds in three stages:
Initiation: The small ribosomal subunit binds mRNA and initiator tRNA; initiation factors help assemble the complete ribosome.
Elongation: Amino acids are added one by one to the growing chain; empty tRNAs exit and are recharged.
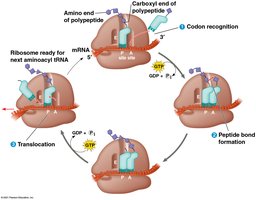
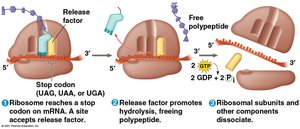
Termination: When a stop codon is reached, release factors bind, prompting the release of the polypeptide and dissociation of the ribosome.
Protein Folding and Post-Translational Modifications
After translation, polypeptides fold into their functional shapes and may undergo further modifications (e.g., cleavage, addition of chemical groups) before becoming active proteins. The primary structure (amino acid sequence) determines the final shape and function.
Mutations and Their Effects on Proteins
Types of Mutations
Mutations are changes in the genetic material. Point mutations affect a single nucleotide pair and can have various effects:
Silent mutations: No effect on the amino acid sequence.
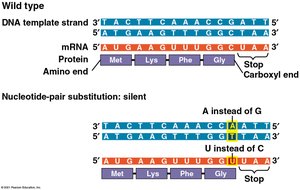
Missense mutations: Change one amino acid to another, possibly altering protein function.
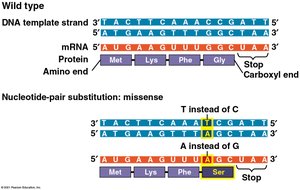
Nonsense mutations: Change an amino acid codon to a stop codon, leading to a truncated, usually nonfunctional protein.
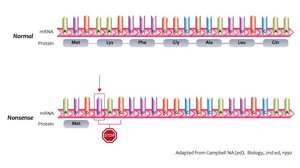
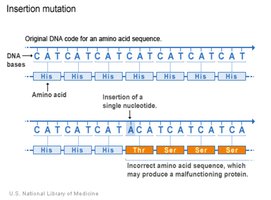
Insertions and Deletions
Insertions and deletions add or remove nucleotide pairs, often causing frameshift mutations that alter the reading frame and usually result in nonfunctional proteins. These mutations can also affect gene expression if they occur outside coding regions.

Mutagens and Spontaneous Mutations
Mutations can arise spontaneously during DNA replication or recombination. Mutagens are physical or chemical agents that increase mutation rates. Many carcinogens are mutagens, highlighting the importance of DNA repair mechanisms in maintaining genetic stability.
Mutation Type | Effect on Protein | Example |
|---|---|---|
Silent | No change in amino acid | GAA → GAG (both code for Glu) |
Missense | One amino acid replaced by another | Sickle cell anemia (Glu → Val) |
Nonsense | Premature stop codon | Duchenne muscular dystrophy |
Frameshift | Altered reading frame, usually nonfunctional protein | Cystic fibrosis (deletion) |
Additional info: The above table summarizes the main types of small-scale mutations and their typical effects on protein structure and function.
