 Back
BackGene Expression: Transcription, RNA Processing, and Translation
Study Guide - Smart Notes

Gene Expression: From Gene to Protein
Overview of the Central Dogma
The central dogma of molecular biology describes the flow of genetic information within a biological system. It states that genetic information is transcribed from DNA to RNA and then translated from RNA to protein. This process involves several key steps: transcription, RNA processing (in eukaryotes), and translation.
Transcription: Synthesis of RNA from DNA
Mechanism of Transcription
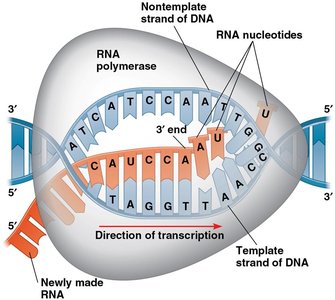
Transcription is the process by which RNA is synthesized from a DNA template. RNA polymerase reads the DNA template strand and synthesizes a complementary RNA strand in the 5' to 3' direction. Only one DNA strand (the template strand) is used for RNA synthesis, while the other (the coding strand) matches the RNA sequence except for uracil (U) replacing thymine (T).
Template strand: The DNA strand used by RNA polymerase to synthesize RNA.
Coding (non-template) strand: The DNA strand with the same sequence as the RNA (except T is replaced by U).
RNA polymerase: The enzyme that catalyzes RNA synthesis, does not require a primer.
Directionality: RNA is synthesized 5' to 3', reading the template 3' to 5'.
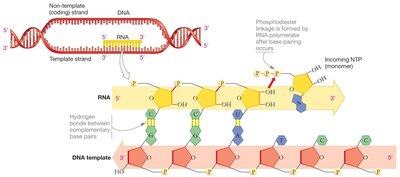
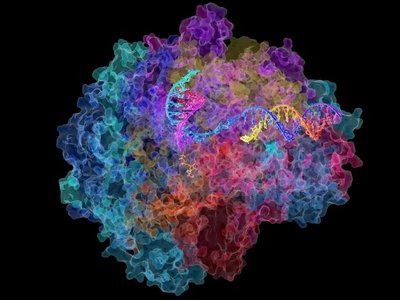
Structure of RNA Polymerase
RNA polymerase is a large, multi-subunit enzyme that binds to DNA and catalyzes the formation of phosphodiester bonds between ribonucleotides, forming the RNA strand.
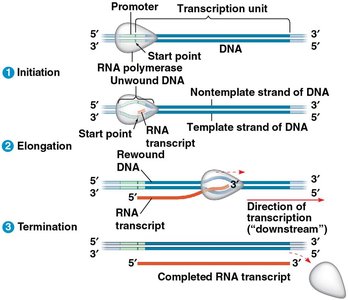
Stages of Transcription
Transcription occurs in three main stages: initiation, elongation, and termination.
Initiation: RNA polymerase binds to the promoter region of DNA, unwinds the DNA, and begins RNA synthesis.
Elongation: RNA polymerase moves along the DNA, adding ribonucleotides to the growing RNA strand.
Termination: RNA polymerase encounters a termination signal, releasing the RNA transcript.
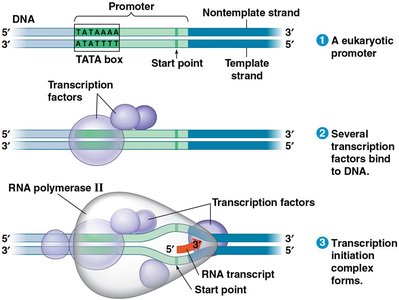
Transcription in Eukaryotes
In eukaryotes, transcription is more complex and involves multiple RNA polymerases and a variety of promoter sequences, such as the TATA box. General transcription factors help RNA polymerase recognize and bind to promoters, forming the transcription initiation complex.
Transcription Elongation and Termination
During elongation, RNA polymerase synthesizes RNA by adding nucleotides complementary to the DNA template. In eukaryotes, transcription continues until a polyadenylation signal (AAUAAA) is transcribed, after which the RNA transcript is cleaved and released as pre-mRNA.
RNA Processing in Eukaryotes
Splicing: Removal of Introns
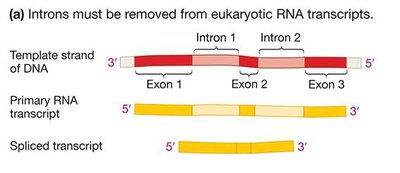
In eukaryotes, the initial RNA transcript (pre-mRNA) contains both exons (coding regions) and introns (non-coding regions). Introns are removed by RNA splicing, which is catalyzed by the spliceosome, a complex of small nuclear ribonucleoproteins (snRNPs) and other proteins. Splicing allows for the production of multiple proteins from a single gene through alternative splicing.
Exons: Sequences that remain in the mature mRNA.
Introns: Sequences removed during RNA processing.
Spliceosome: The complex responsible for splicing.
5' Capping and 3' Polyadenylation
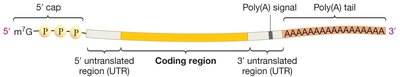
Pre-mRNAs are further processed by the addition of a 5' cap (a modified guanine nucleotide) and a 3' poly(A) tail (a stretch of adenine nucleotides). These modifications protect mRNA from degradation, aid in export from the nucleus, and are important for translation initiation.
5' cap: Facilitates ribosome binding and protects mRNA.
Poly(A) tail: Enhances stability and translation efficiency.
Untranslated regions (UTRs): Non-coding regions at both ends of mature mRNA.
Translation: Synthesis of Proteins from mRNA
Overview of Translation
Translation is the process by which the sequence of bases in mRNA is converted into an amino acid sequence, forming a polypeptide. This process requires ribosomes, mRNA, and transfer RNAs (tRNAs).
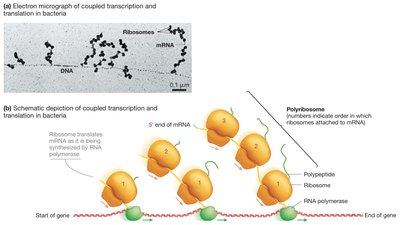
Coupling of Transcription and Translation in Prokaryotes
In bacteria, translation can begin before transcription is complete, as both processes occur in the cytoplasm. Multiple ribosomes can simultaneously translate a single mRNA, forming a polyribosome (polysome).
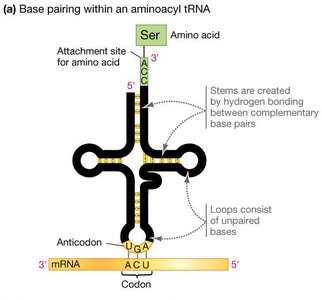
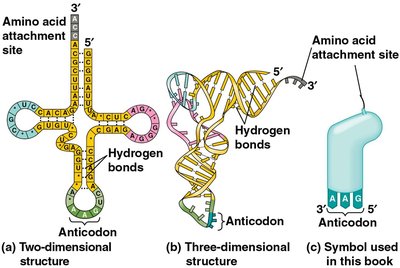
Structure and Function of tRNA
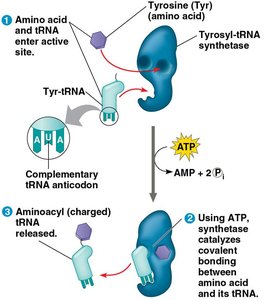
tRNAs are adapter molecules that match specific amino acids to codons in the mRNA. Each tRNA has an anticodon that base-pairs with a codon in mRNA and an amino acid attachment site. Aminoacyl-tRNA synthetases charge tRNAs with their corresponding amino acids using ATP.
Aminoacyl-tRNA: A tRNA linked to its amino acid.
Aminoacyl-tRNA synthetase: Enzyme that attaches the correct amino acid to its tRNA.
Ribosome Structure and Function
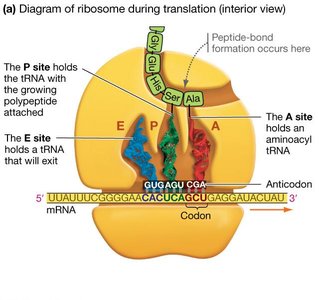
Ribosomes are large complexes of rRNA and proteins, composed of a small and a large subunit. They facilitate the binding of tRNAs to mRNA and catalyze peptide bond formation. Ribosomes have three binding sites for tRNA: the A (aminoacyl), P (peptidyl), and E (exit) sites.
A site: Holds the tRNA carrying the next amino acid.
P site: Holds the tRNA with the growing polypeptide chain.
E site: Site from which tRNAs exit the ribosome.
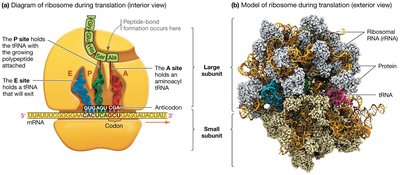
Phases of Translation
Translation occurs in three phases: initiation, elongation, and termination.
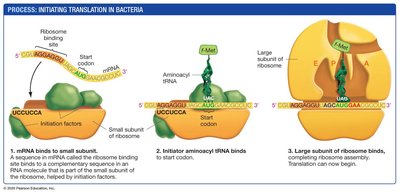
Initiation: The small ribosomal subunit binds to mRNA, and the initiator tRNA binds to the start codon. The large subunit then assembles, forming the complete initiation complex.
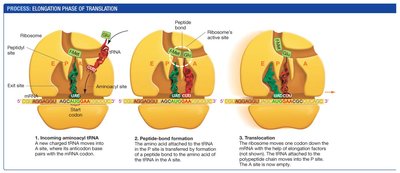
Elongation: Amino acids are added one by one to the growing polypeptide chain. Each cycle involves entry of an aminoacyl-tRNA into the A site, peptide bond formation, and translocation of the ribosome.
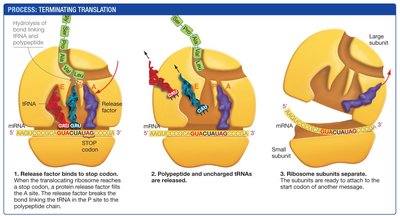
Termination: When a stop codon is encountered, a release factor binds, causing the polypeptide to be released and the ribosome to dissociate.
Post-Translational Modification
After translation, most proteins undergo post-translational modifications, such as folding, cleavage, or addition of chemical groups, to become fully functional. These modifications occur in various cellular locations and are essential for proper protein function.
Summary Table: Key Steps in Eukaryotic Gene Expression
Step | Location | Main Events |
|---|---|---|
Transcription | Nucleus | RNA polymerase synthesizes pre-mRNA from DNA template |
RNA Processing | Nucleus | Splicing, 5' capping, 3' polyadenylation to produce mature mRNA |
Translation | Cytoplasm | Ribosomes synthesize polypeptide from mRNA template |
Post-Translational Modification | Various | Folding, cleavage, chemical modification of polypeptide |
Key Terms and Concepts
Central Dogma: DNA → RNA → Protein
Transcription: Synthesis of RNA from DNA
RNA Processing: Modifications to pre-mRNA in eukaryotes
Translation: Synthesis of protein from mRNA
tRNA: Adapter molecule in translation
Ribosome: Molecular machine for protein synthesis
Polyribosome: Multiple ribosomes translating a single mRNA
Post-Translational Modification: Chemical changes to proteins after synthesis
