 Back
BackGene Function: Transcription, Translation, and the Genetic Code
Study Guide - Smart Notes

Gene Function and the Central Dogma
Overview of Genetic Information Flow
The central dogma of molecular biology describes the flow of genetic information within a biological system. It explains how DNA is transcribed into RNA, which is then translated into proteins, the functional molecules in cells.
DNA: Stores genetic information in the nucleus.
Transcription: The process by which an RNA copy (mRNA) is made from a DNA template.
Translation: The process by which the sequence of the mRNA is used to build a protein.

Example: The gene for hemoglobin is transcribed into mRNA, which is then translated to produce the hemoglobin protein in red blood cells.
The Genetic Code
Codons and the Triplet Code
The genetic code is a set of rules by which information encoded in mRNA sequences is translated into proteins. Each amino acid is specified by a sequence of three nucleotides, called a codon.
There are four RNA bases: adenine (A), cytosine (C), guanine (G), and uracil (U).
A 3-base codon system allows for 64 possible codons, which is more than enough to code for the 20 amino acids.
Some amino acids are specified by more than one codon (degeneracy of the code).
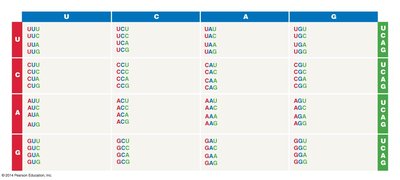
Example: The codon AUG codes for methionine and also serves as the start codon for translation.
Translating DNA to Protein
To predict the amino acid sequence from a DNA sequence, the DNA is first transcribed into mRNA, and then the mRNA codons are translated into amino acids using the genetic code.
Transcription replaces thymine (T) in DNA with uracil (U) in RNA.
Translation reads the mRNA in sets of three bases (codons).
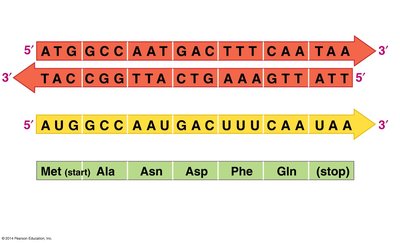
Example: The DNA sequence ATG codes for the mRNA codon AUG, which specifies methionine.
Transcription: DNA to RNA
Mechanism of Transcription
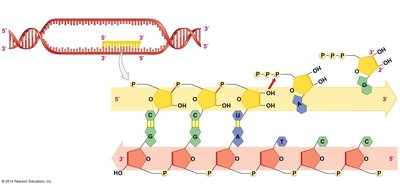
Transcription is the synthesis of an RNA molecule from a DNA template. This process is catalyzed by the enzyme RNA polymerase.
RNA polymerase binds to the DNA at a specific region called the promoter.
It synthesizes RNA in the 5' to 3' direction, using one strand of DNA as the template.
RNA polymerase does not require a primer to begin synthesis.
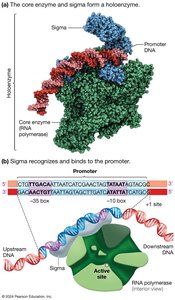
Example: In bacteria, the sigma factor helps RNA polymerase recognize the promoter region.
Initiation, Elongation, and Termination in Bacteria
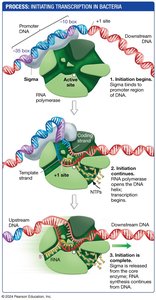
Transcription in bacteria involves three main stages: initiation, elongation, and termination.
Initiation: Sigma factor binds to RNA polymerase, forming a holoenzyme that recognizes the promoter.
Elongation: RNA polymerase moves along the DNA, synthesizing RNA.
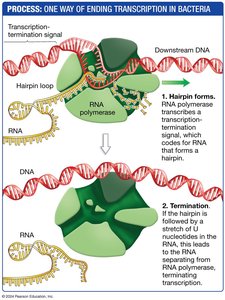
Termination: A hairpin loop forms in the RNA, causing the polymerase to dissociate from the DNA.
RNA Processing in Eukaryotes
Splicing and mRNA Modification
In eukaryotes, the primary RNA transcript (pre-mRNA) undergoes several modifications before becoming mature mRNA.
Splicing: Removal of non-coding sequences (introns) and joining of coding sequences (exons).
5' Cap: Addition of a modified guanine nucleotide to the 5' end for stability and ribosome recognition.
Poly(A) Tail: Addition of a string of adenine nucleotides to the 3' end for stability and export from the nucleus.
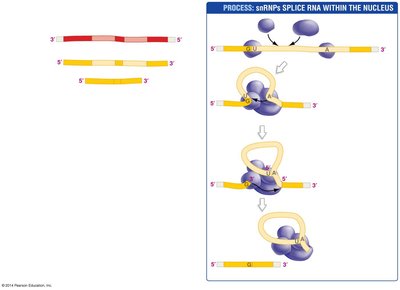
Example: The human beta-globin gene contains two introns that are removed during RNA processing.
Translation: RNA to Protein
Mechanism of Translation
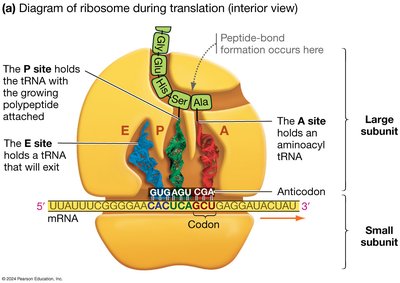
Translation is the process by which ribosomes synthesize proteins using the sequence of codons in mRNA as a template.
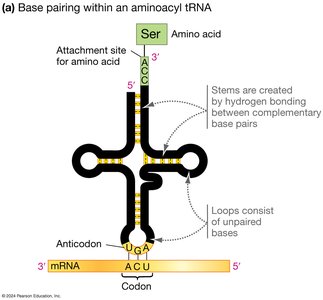
tRNA (transfer RNA) molecules serve as adapters, matching amino acids to codons in the mRNA via their anticodons.
Ribosomes facilitate the binding of tRNA to mRNA and catalyze peptide bond formation.
Translation occurs in three stages: initiation, elongation, and termination.
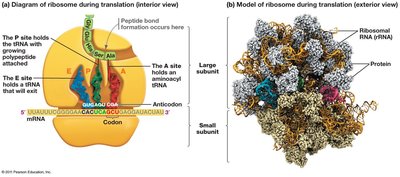
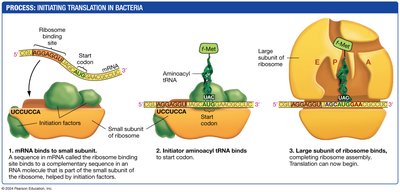
Translation Initiation, Elongation, and Termination
Translation begins with the assembly of the ribosome on the mRNA, continues as amino acids are added to the growing polypeptide, and ends when a stop codon is reached.
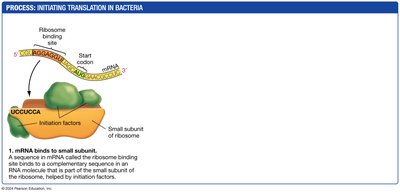
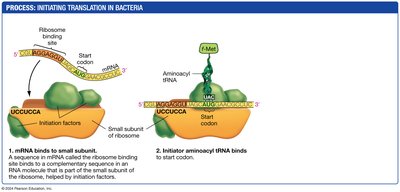
Initiation: The small ribosomal subunit binds to the mRNA, and the initiator tRNA binds to the start codon.
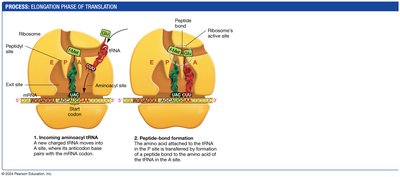
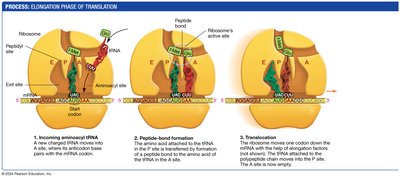
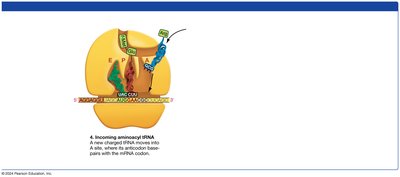
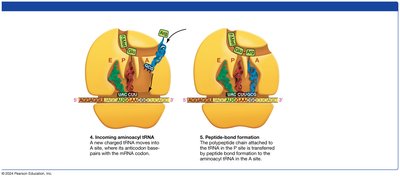
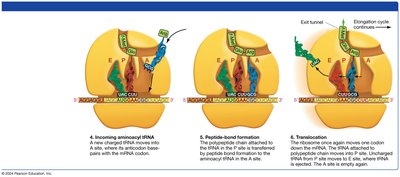
Elongation: tRNAs bring amino acids to the ribosome, where peptide bonds are formed.
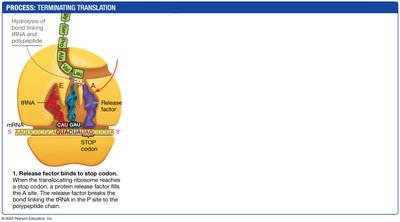
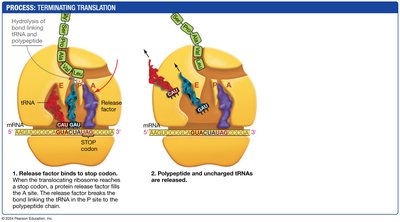
Termination: A release factor binds to the stop codon, releasing the completed polypeptide.

Summary Table: Key Steps in Gene Expression
Step | Location | Main Enzyme/Structure | Product |
|---|---|---|---|
Transcription | Nucleus (eukaryotes), cytoplasm (prokaryotes) | RNA polymerase | mRNA (primary transcript) |
RNA Processing | Nucleus (eukaryotes) | Spliceosome, capping and polyadenylation enzymes | Mature mRNA |
Translation | Cytoplasm (ribosomes) | Ribosome, tRNA | Polypeptide (protein) |
Additional info:
Post-translational modifications such as phosphorylation, glycosylation, and lipidation can further regulate protein function and localization.
In prokaryotes, transcription and translation are often coupled, occurring simultaneously in the cytoplasm.
